Abstract
Kynurenine 3-monooxygenase (KMO) is a therapeutically important target on the eukaryotic tryptophan catabolic pathway, where it converts L-kynurenine (Kyn) to 3-hydroxykynurenine (3-HK). We have cloned and expressed the human form of this membrane protein as a full-length GST-fusion in a recombinant baculovirus expression system. An enriched membrane preparation was used for a directed screen of approximately 78,000 compounds using a RapidFire mass spectrometry (RF-MS) assay. The RapidFire platform provides an automated solid-phase extraction system that gives a throughput of approximately 7 s per well to the mass spectrometer, where direct measurement of both the substrate and product allowed substrate conversion to be determined. The RF-MS methodology is insensitive to assay interference, other than where compounds have the same nominal mass as Kyn or 3-HK and produce the same mass transition on fragmentation. These instances could be identified by comparison with the product-only data. The screen ran with excellent performance (average Z′ value 0.8) and provided several tractable hit series for further investigation.
Keywords
Introduction
Kynurenine 3-monooxygenase (KMO; EC 1.14.13.9) is a β-nicotinamide adenine dinucleotide 2′-phosphate (NADPH)- dependent flavin hydroxylase that catalyzes the hydroxylation of L-kynurenine (Kyn) to form 3-hydroxykynurenine (3-HK) using molecular oxygen (scheme 1, supplemental material). This reaction occurs on the pathway of tryptophan catabolism and, in higher organisms, leads to the synthesis of the nicotinamide moiety of NAD/NADP, with quinolinic acid as an intermediate on the pathway. 1 Both 3-HK and quinolinic acid are neurotoxic,2,3 whereas kynurenic acid, which can also be formed from kynurenine by the action of kynurenine aminotransferase, is neuroprotective.4–6 The position of KMO at this branch point in the pathway makes it a potential therapeutic target for treating neurodegenerative disorders, such as Alzheimer disease 7 and Huntington disease. 8 Studies have already shown positive effects of KMO inhibitors in brain injury models, 9 although poor penetration of the blood-brain barrier is problematic. 7
Previously described assays for KMO, such as the measurement of absorbance of NADPH, 10 are typically insensitive and prone to interference by contaminating enzyme activities. A radiometric assay, based on the release of tritiated water, was described by Erickson et al. 11 and has been adapted to a 96-well plate format 12 but is unsuited to large-scale automation. Recently, the use of liquid chromatography–tandem mass spectrometry was reported as a sensitive, label-free, and direct method for monitoring the conversion of Kyn to 3-HK, which was successfully applied to the assay of KMO activity in tissue extracts as well as cellular assays with a throughput of approximately 90 s per sample. 13 We describe here a complementary application of mass spectrometry to the measurement of KMO activity, of approximately 10-fold higher throughput, using the RapidFire high-throughput mass spectrometry (RF-MS) system (Agilent Technologies, Lexington, MA). This enabling technology has previously been successfully used in lead discovery campaigns for challenging targets.14-17 It overcomes the problem of time-consuming sample separation by using an in-line solid-phase extraction (SPE) cartridge system for sample clean up and analyte concentration. This delivers the sample directly to an electrospray ionization (ESI) triple quadrupole mass spectrometer at a rate of approximately 7 s per well. Using RF-MS, a directed screen of 78,000 compounds was undertaken for human KMO using a recombinant source of full-length, membrane-bound protein. This screen resulted in several tractable hit series, examples of which were examined for their biochemical mode of action using the RF-MS assay.
Materials and Methods
Materials
L-kynurenine (Kyn), 3-hydroxy-DL-kynurenine (3-HK), NADPH, 4-(2-hydroxyethyl)piperazine-1-ethanesulfonic acid (Hepes), DL-dithiothreitol (DTT), ethylenediaminetetraacetic acid (EDTA), CHAPS, trifluoroacetic acid (TFA), D-glucose-6-phosphate disodium salt hydrate, and glucose-6-phosphate dehydrogenase from baker’s yeast (G4134) were purchased from Sigma-Aldrich Ltd. (Gillingham, Dorset, UK). High-performance liquid chromatography (HPLC)–grade acetonitrile and formic acid were supplied by Fisher Scientific (Loughborough, UK).
Cloning and Expression of Human KMO
Full-length human KMO was amplified by PCR from pcDNA5/FRT/V5-His-TOPO/hKMO (vector provided by the University of Edinburgh) and cloned into pGEX6P-1 (GE Healthcare) using BamH1 and Sal1 restriction sites. DNA encoding the N-terminal glutathione-S-transferase (GST) tag, followed by a PreScission protease cleavage site, and the full-length KMO was amplified by PCR from pGEX6P-1-KMO and cloned into pFastbac1 (Invitrogen, Carlsbad, CA) using Xbal and EcoR1 restriction sites. pFastbac1 GST-KMO was transposed into the baculovirus genome using the BAC-to-BAC technology (Invitrogen), and bacmid DNA was prepared and transfected into Spodoptera frugiperda (Sf9) cells using Cellfectin II (Invitrogen). Expression of a protein of the expected molecular weight (Mr 82,634) was seen by Western blot analysis using anti-GST-peroxidase conjugate.
Preparation of Membranes from Sf9 Cells Expressing Human GST-KMO
A P1 virus stock was generated from a single clone and used to infect 3 × 1.5 L cultures of Sf9 cells in 3 L Corning Fernbach flasks. The Sf9 cells were grown in Hyclone SFX media (Thermo Scientific, Waltham, MA) to about 3 × 10 6 cells/mL and were infected at a nominal multiplicity of infection of 3. Cells were harvested after 48 h and disrupted by blending in 50 mM Hepes, pH 7.4, 1 mM EDTA buffer containing protease inhibitors. A low-speed centrifugation (400 × g) was used to remove cell debris, followed by a high-speed centrifugation (75,000 × g) to pellet the membranes. The membranes were purified in a discontinuous sucrose density gradient by resuspending in 10% (w/v) sucrose and layering over 40% (w/v) sucrose, both in the above buffer. This was centrifuged at 150,000 × g, and the purified membranes were taken from the interface, collected by centrifugation at 100,000 × g, resuspended in buffer, and aliquoted for storage at −80 °C. A batch of 100 mg of KMO membranes (as determined by the Pierce BCA protein assay using bovine serum albumin as standard) was prepared and validated for use in the RF-MS assay.
Biochemical Assays
Assays were performed in 384-well V-base polypropylene plates (Greiner Bio-one, Stonehouse, UK) in a final volume of 10 µL of 50 mM Hepes, pH 7.5, 2 mM DTT, 1 mM EDTA, 0.1 mM CHAPS, 200 µM NADPH, 10 µM Kyn, and 8 µg/mL KMO membranes, unless otherwise stated. Assays were generally run for 2 h at ambient temperature (approximately 20 °C). When determining Km for each substrate, the substrate concentration was varied in the presence of 1 mM of the second substrate. A regeneration system, consisting of 3 mM D-glucose-6-phosphate and 1 U/mL glucose-6-phosphate dehydrogenase, was included in the NADPH Km determination to prevent substrate depletion at low-NADPH concentrations. Active site titration was performed by titrating 25 µg/mL KMO membranes with compound 1 ( Table 1 ) and terminating the subsequent reaction after 1 h. Determination of inhibitor competitivity with respect to kynurenine was performed in the presence of 200 µM NADPH by varying the concentration of both kynurenine and the inhibitor. Inhibitor reversibility experiments were performed by rapidly diluting a preformed complex between KMO and inhibitor into the assay components; 800 µg/mL KMO membranes were incubated with 1000×, 100×, or 10× the Ki of compound for 10 min before diluting 1:100 into assay mix. The time courses of the reactions were then followed by repeat sampling on the RapidFire system.
Inhibition of KMO by compounds 1 to 3. a
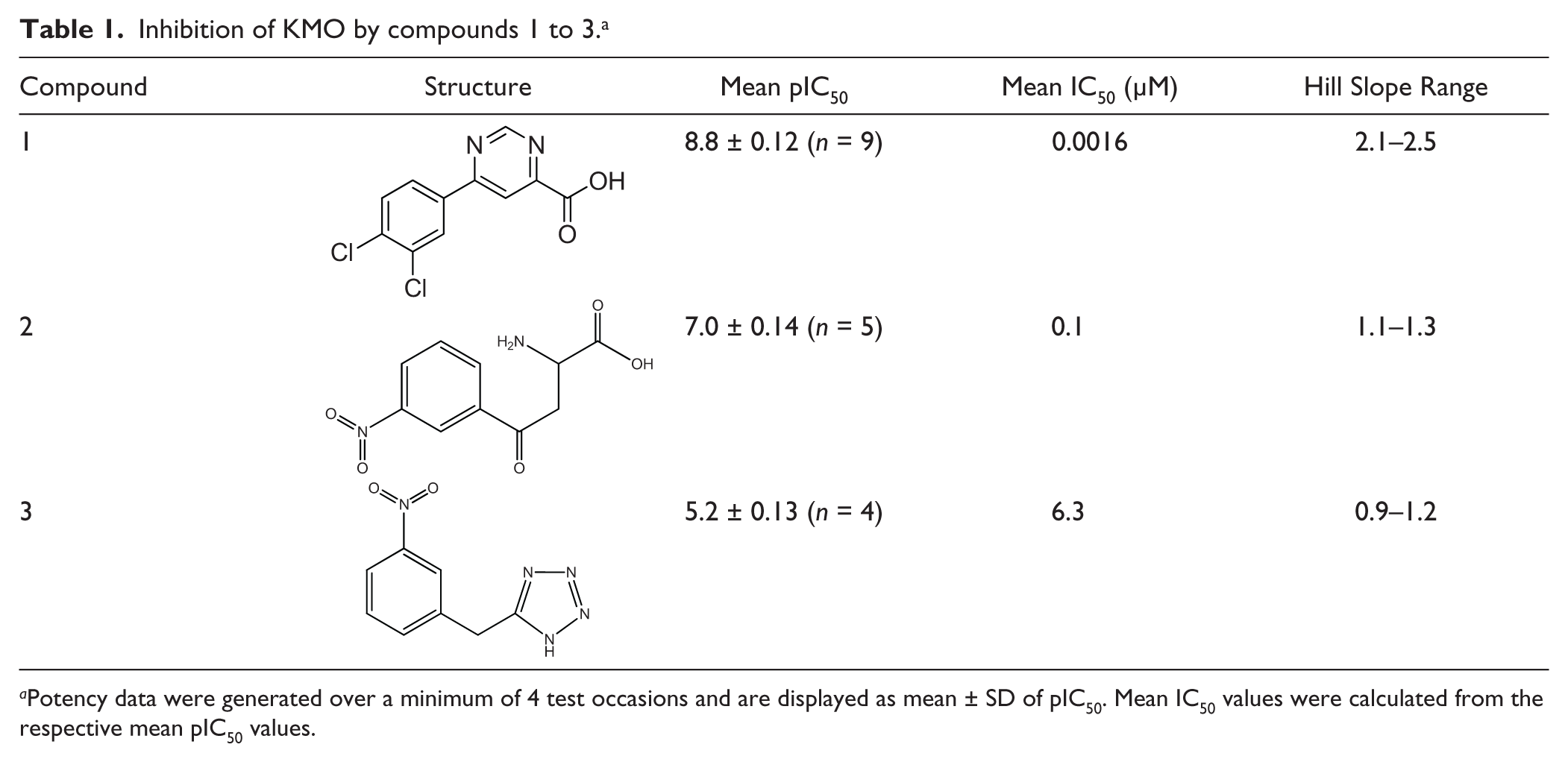
Potency data were generated over a minimum of 4 test occasions and are displayed as mean ± SD of pIC50. Mean IC50 values were calculated from the respective mean pIC50 values.
RapidFire High-Throughput Mass Spectrometry Assay
Single-concentration screening was performed by dispensing 100 nL of 1 mM compound solutions in DMSO into 384-well V-base polypropylene plates using an Echo 555 acoustic dispenser (Labcyte, Sunnyvale, CA). This gave a final compound concentration of 10 µM in a 10 µL final assay volume (see below). A total of 100 nL DMSO was dispensed into columns 6 and 18 for high and low controls, respectively, with prior inactivation of the enzyme in column 18 by predispense of 30 µL of 0.5% (v/v) TFA. For concentration-response curves, 100 nL test compounds were dispensed as 11-point, 3-fold serial dilutions. The upper concentration in the assay was either 100 µM or 10 µM as appropriate.
Conditions for the high-throughput assay were the same as those described above. Assays were performed by initially dispensing 5 µL of a 2× enzyme solution (16 µg/mL KMO membranes in 50 mM Hepes, pH 7.5, 2 mM DTT, 2 mM EDTA, 0.2 mM CHAPS) into plates containing 100 nL compounds and incubating for 10 min at ambient temperature. Reactions were initiated by addition of 5 µL of 2× substrate solution (400 µM NADPH, 20 µM Kyn in 50 mM Hepes, pH 7.5, 2 mM DTT) and incubated for 2 h at room temperature before quenching the reaction with 30 µL of 0.5% (v/v) TFA. Plates were centrifuged at 2500 rpm for 10 min before analysis. All additions were made using a Multidrop Combi dispenser (Thermo Fisher Scientific). Quenched assay plates were transferred to a RapidFire200 integrated autosampler/SPE system (Agilent Technologies). Samples were aspirated from each well for 500 ms, and 10 µL was loaded directly onto a RapidFire micro-scale SPE C18 (type C) cartridge, which was washed for 3 s with HPLC-grade water containing 0.1% (v/v) formic acid to remove non-organic components. Analytes were then eluted into the mass spectrometer, in a 3 s elution cycle, using 80% (v/v) acetonitrile/water containing 0.1% (v/v) formic acid, and the cartridge was then equilibrated by washing with water containing 0.1% (v/v) formic acid for 500 ms. This gave a total cycle time of 7 s, enabling analysis of a 384-well plate in approximately 45 min.
Kyn and 3-HK were detected using a Sciex API4000 triple quadrupole mass spectrometer (Applied Biosystems, Concord, Ontario, Canada), equipped with an electrospray interface and operated in positive ion mode. Multiple reaction monitoring (MRM) was used to detect both Kyn and 3-HK using Q1/Q3 transitions at m/z 209.4 to 192.0 and m/z 225.3 to 208.2, respectively. The mass spectrometer used an ESI voltage of 5500 V and a source temperature of 600 °C, with a dwell time of 50 ms for each transition.
Data Analysis
Individual MRM transitions were saved as text files, and the extracted ion chromatograms were integrated and processed using the RapidFire peak integration software (version 3.6).
Using the integrated peak area data, the extent of conversion of Kyn to 3-HK within a sample could be expressed as a percentage using equation 1:
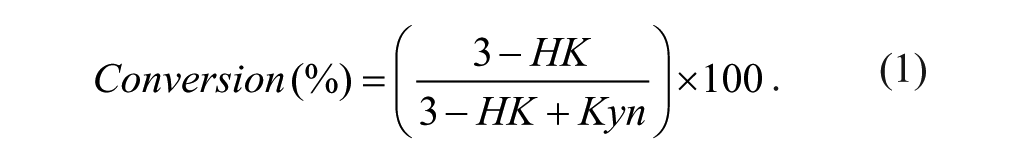
Screening data were further analyzed within ActivityBase (ID Business Solutions Ltd, Surrey, UK) to generate percentage inhibition data. For concentration-response screening, data were fitted to equation 2:
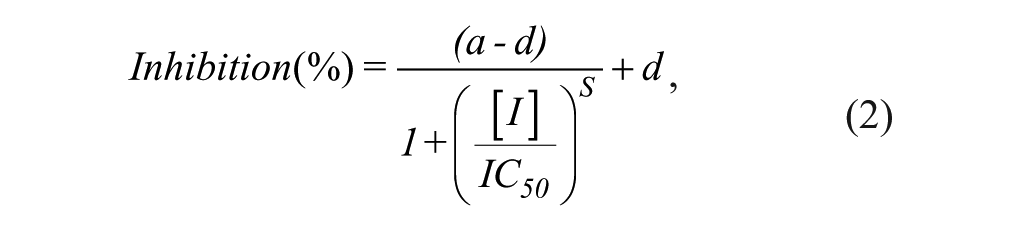
where a is the uninhibited response, d is the fully inhibited response, [I] is the inhibitor concentration, IC50 is [I] that gives 0.5 × (a-d), and S is the Hill slope.
Steady-state enzyme kinetic data were fitted using nonlinear regression to the Michaelis-Menten equation 3 and the competitive inhibition equation 4 using GraFit version 7.0.3 (Erithacus Software Ltd, Surrey, UK).

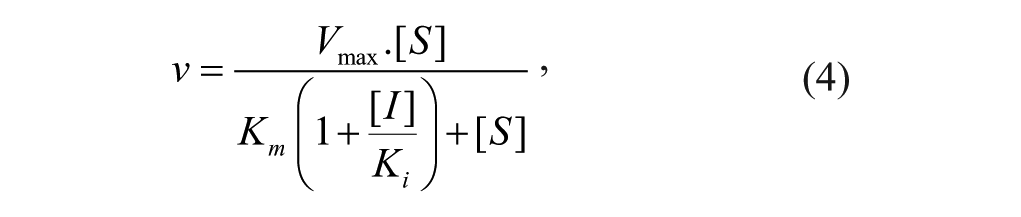
where ν is the observed rate at a substrate concentration [S], Vmax is the rate at saturating substrate concentration, Km is the Michaelis constant, [I] is the inhibitor concentration, and Ki is the dissociation constant of the inhibitor.
Morrison’s quadratic equation 5 was used to determine the active-site concentration of the enzyme using a tight-binding inhibitor. 18
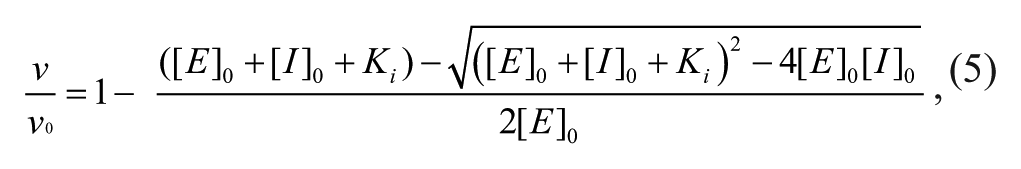
where ν0 is the rate in the absence of inhibitor, [E] 0 is the total active enzyme concentration, and [I]0 is the total concentration of inhibitor. When determining [E]0 with this equation, Ki was fixed at the value determined under the standard assay conditions.
Results and Discussion
Supply of Recombinant Human KMO
Human KMO is a membrane-bound protein that is anchored via a C-terminal region to the mitochondrial outer membrane. 19 In the recently published work of Winkler et al., 13 KMO activity was obtained by subcellular fractionation of liver tissue from various species, and relatively high concentrations of the resulting material were required for assay (~0.1 mg/mL total protein). To provide a consistent supply of KMO activity for an extended hit to lead campaign and avoid the need to obtain human tissue, recombinant expression was assessed. A soluble, bacterial form of KMO from Pseudomonas fluorescens has been expressed efficiently in Escherichia coli 20 ; however, our attempts at expressing the full-length human protein in this host were not successful. Recombinant human KMO has been expressed in mammalian cells by transient transfection,10,21 but the low levels of expression reported in these systems would have been prohibitive for an extended screening campaign.
At the outset of this work, an HEK293 stable cell line expressing full-length human KMO, previously generated for cell-based assay purposes, was evaluated. Membranes isolated from this cell line were active and could be used to configure a RapidFire assay; however, a concentration of approximately 0.5 mg/mL (based on total membrane protein concentration) was required to give sufficient turnover of Kyn (data not shown). As well as being challenging to supply at the required scale, the high concentrations of membrane protein were not well tolerated by the RapidFire SPE system, requiring frequent cartridge replacement. Baculovirus-mediated expression in Sf9 cells, which is often found to give high levels of heterologous gene expression compared with other eukaryotic expression systems, was evaluated as an alternative. An N-terminal GST fusion of full-length human KMO was found to express well in this system. KMO activity was associated with the membrane fraction only, and no KMO activity was detected in membranes prepared from uninfected Sf9 cells. This approach has subsequently been described by Amaral et al. 22 The level of activity was sufficient for a robust assay to be developed using only 8 µg/mL of Sf9 membranes, compared with 0.5 mg/mL of similarly prepared HEK293 membranes. Active site titration of the KMO activity in the 2 preparations of membranes was carried out with the potent inhibitor compound 1 ( Table 1 ) to determine whether expression levels or differences in posttranslational modification were responsible for the approximately 60-fold higher activity observed with the Sf9-derived membranes. This indicated that active KMO comprised approximately 2% of the total protein in the Sf9-derived membranes but only about 0.02% of the total protein in the HEK293-derived membranes. Thus, it seems likely that the differences in activity between the 2 preparations are due to higher expression levels in Sf9 cells.
The lower required concentration of Sf9-derived membranes was more compatible with the RapidFire system, with the life span of a cartridge generally exceeding 100 plates. A 4.5 L Sf9 cell culture provided 100 mg of KMO membranes, which was sufficient for testing >1 million assay wells. This represents a significant economy of materials compared with the equivalent requirement for HEK293-derived KMO membranes.
RF-MS Assay Development
The direct measurement of the substrate and product of the KMO reaction by mass spectrometry is an ideal method for following KMO activity, being label-free, sensitive, and not subject to the interference associated with absorbance or fluorescence readouts. 15 Although this method has previously been used for routine assay of KMO activity,10,13 the use of the RF-MS system provided the increased throughput desirable for a lead discovery campaign. Mass spectrometer protocols were established for the simultaneous detection of Kyn and 3-HK, giving no signal cross-talk between the channels, thus allowing the percentage conversion to be calculated. Solid-phase cartridge of C4 (type A), C8 (type E), phenyl (type F), and C18 (type C) were evaluated, and C18 gave the best performance. The sampling and running parameters were optimized so as to give the best throughput without compromising sensitivity or reproducibility of detection. The sensitivity of detection of both Kyn and 3-HK was similar, and the response was linear, with concentration up to at least 40 µM, with the lower limit of robust detection for both analytes being approximately 100 nM ( Fig. 1 ).
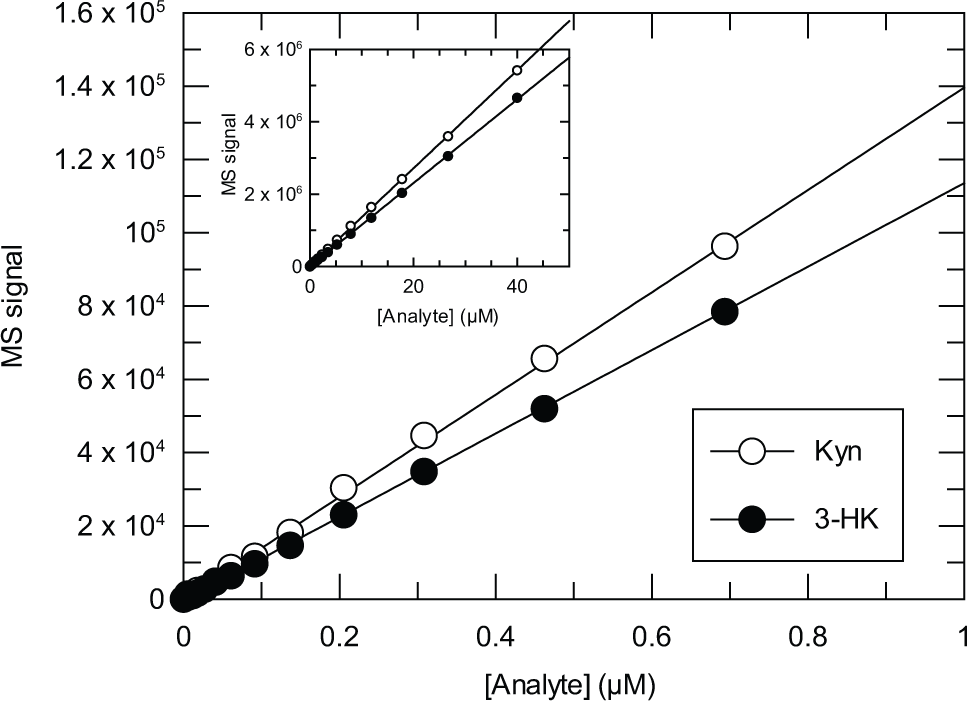
RapidFire mass spectrometry signal response to L-kynurenine (Kyn) and 3-hydroxykynurenine (3-HK).
The relationship between substrate turnover and the concentration of KMO membranes under the standard assay conditions (see the Materials and Methods section) is shown in Figure 2 . At high membrane concentration, the conversion of Kyn to 3-HK was quantitative. A concentration of 8 µg/mL KMO membranes was chosen for the screening assay, which gave between 40% and 50% turnover of Kyn under the standard assay conditions and excellent assay robustness. By using a feature of the RapidFire that allows repeat sampling from the same well, it was possible to follow the time course of the KMO reaction automatically at various KMO concentrations ( Fig. 3 ). This established that the reaction was linear over 2 h using 4 and 8 µg/mL KMO membranes. The pH optimum for the assay was found to be pH 7.5 (data not shown), in agreement with a previous report for human KMO. 10 A low concentration of Chaps detergent was included, as this was found to improve assay performance, possibly by ameliorating the adsorption of assay components to the plastic tubing of the Multidrop Combi dispenser. In addition, the inclusion of a subcritical micellar concentration of detergent has been shown to reduce the occurrence of nonspecific inhibition in diversity screening resulting from compound aggregation. 23 The incorporation of Chaps detergent was found not to affect the enzyme activity, the performance of the mass spectrometer, or the pIC50 values obtained for the 3 inhibitors shown in Table 1 . The assay was highly tolerant of DMSO, with negligible loss of activity up to 10% (data not shown).
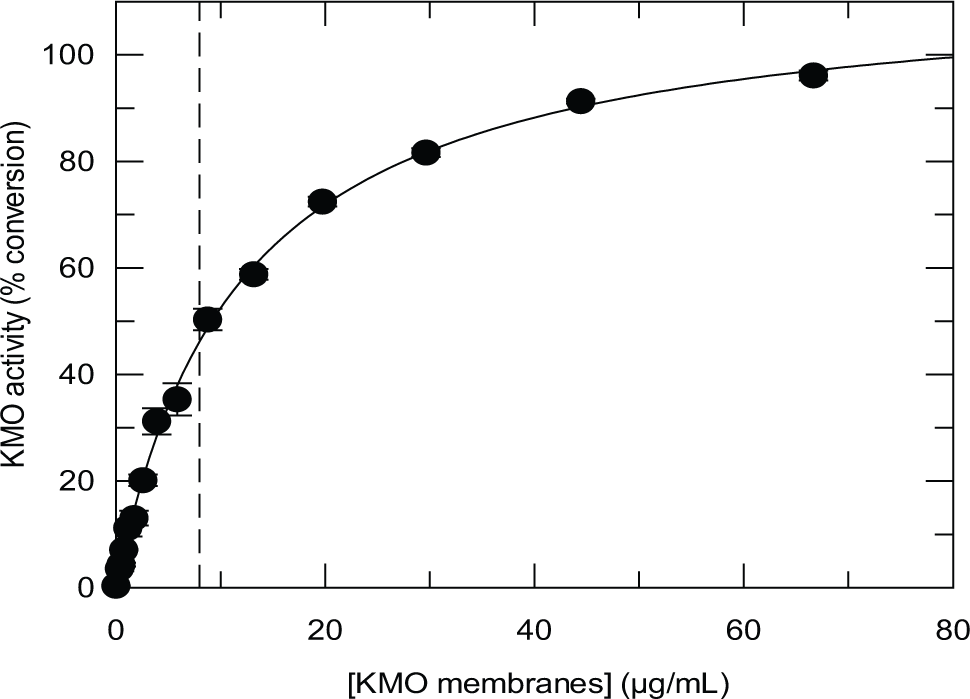
Relationship between L-kynurenine (Kyn) turnover and concentration of kynurenine 3-monooxygenase (KMO) membranes in a 2 h assay containing 10 µM Kyn and 200 µM NADPH. The concentration of KMO membranes used in the screen (8 µg/mL) is indicated by -----.
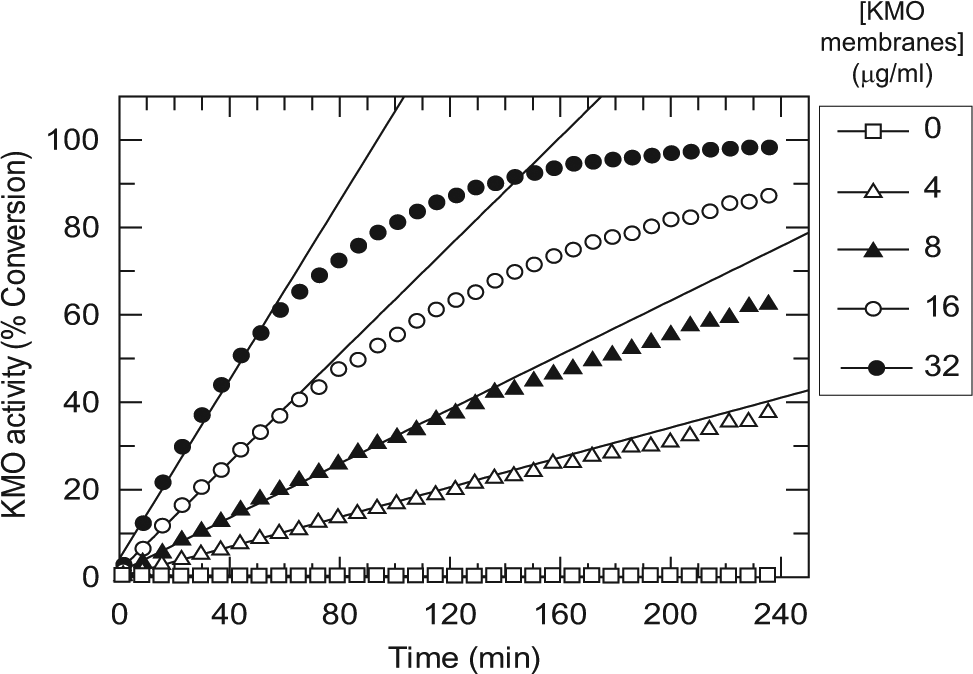
Time courses of the kynurenine 3-monooxygenase (KMO) RapidFire mass spectrometry (RF-MS) assay at increasing concentrations of KMO membranes. Sampling was performed by the RF-MS by repeat sipping from a deep 96-well plate. Product formation was linear with time up to 120 min at 4 and 8 µg/mL KMO membranes.
Km values of 7 µM for Kyn and 0.13 µM for NADPH were determined, using saturating concentrations of the unvaried substrate ( Fig. 4 ). Similar values were obtained for the HEK293-derived KMO membranes (data not shown). The Km value for Kyn is comparable to reported values of 13 µM for a crude extract of recombinant human KMO expressed in mammalian cells 21 and 14 µM for KMO obtained from human liver extracts. 13 Considerably higher Km values have been reported for both substrates against human KMO which had been expressed in mammalian cells and solubilized and purified in the presence of 0.2% Triton X-100. 10 Treatment of the Sf9-derived KMO with a membrane-solubilizing concentration of detergent (0.01% n-dodecyl-β-D-maltoside) also resulted in an elevation of the Km values for both substrates, to 42 µM for Kyn and 2 µM for NADPH, suggesting that the membrane environment is involved in substrate binding. The final conditions for the screening campaign were chosen to provide a robust, linear assay that was sensitive to detection of Kyn-competitive inhibitors. A saturating concentration of NADPH relative to its Km value was used to render the assay insensitive to NADPH competitive inhibitors.
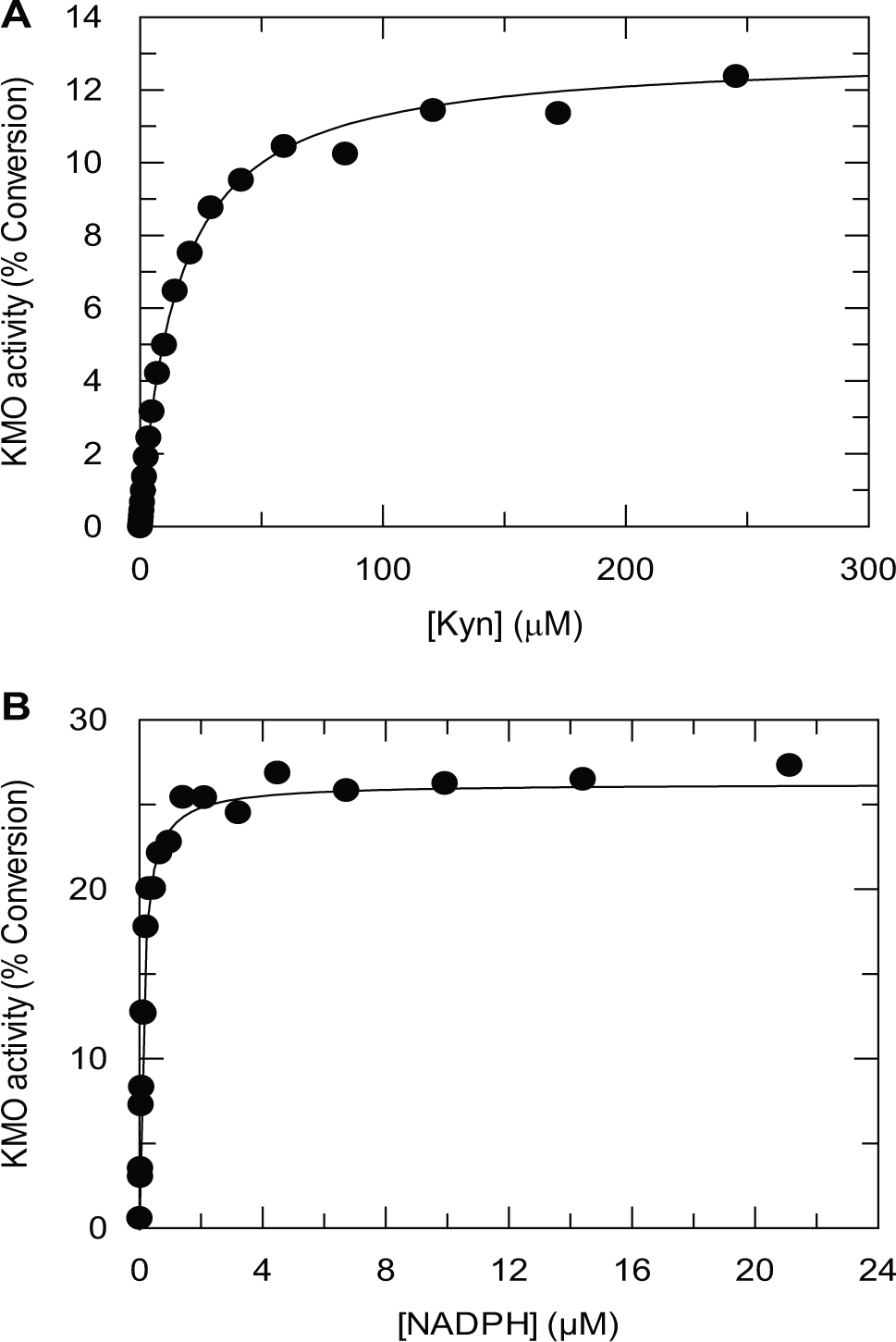
Determination of substrate Km values for kynurenine 3-monooxygenase (KMO) membranes. (
The assay was tested using some established KMO inhibitors ( Table 1 ). The highly potent compound 1 24 gave a mean pIC50 of 8.8 and displayed a high Hill slope, suggesting that this inhibitor was interacting with KMO under tight binding conditions. The observed potency is thus likely to be an underestimate of the true potency. This was confirmed by testing the inhibitor in a range of KMO membrane concentrations between 4 µg/mL and 32 µg/mL. The apparent potency was observed to decrease as the concentration of KMO membranes was increased. Compound 2 (m-nitrobenzoylalanine) gave a mean pIC50 value of 7.0, which compares favorably to a published pIC50 of 6.1 measured in the presence of 100 µM kynurenine. 4 The potency of this compound in the Rapidfire assay was unaffected by the concentration of KMO membranes in a range between 4 µg/mL and 32 µg/mL. The availability of the highly potent inhibitor (compound 1) allowed the concentration of active KMO in the membrane preparation to be determined by active site titration. Figure 5A shows the inhibition profile of compound 1 at an elevated assay concentration of KMO membranes (25 µg/mL). Analysis of these data using the Morrison equation showed that the binding-competent concentration of KMO was 6.5 nM at 25 µg/mL KMO membranes. Thus, the tight binding limit of the assay, run using 8 µg/mL KMO membranes, is ~pIC50 9.0.
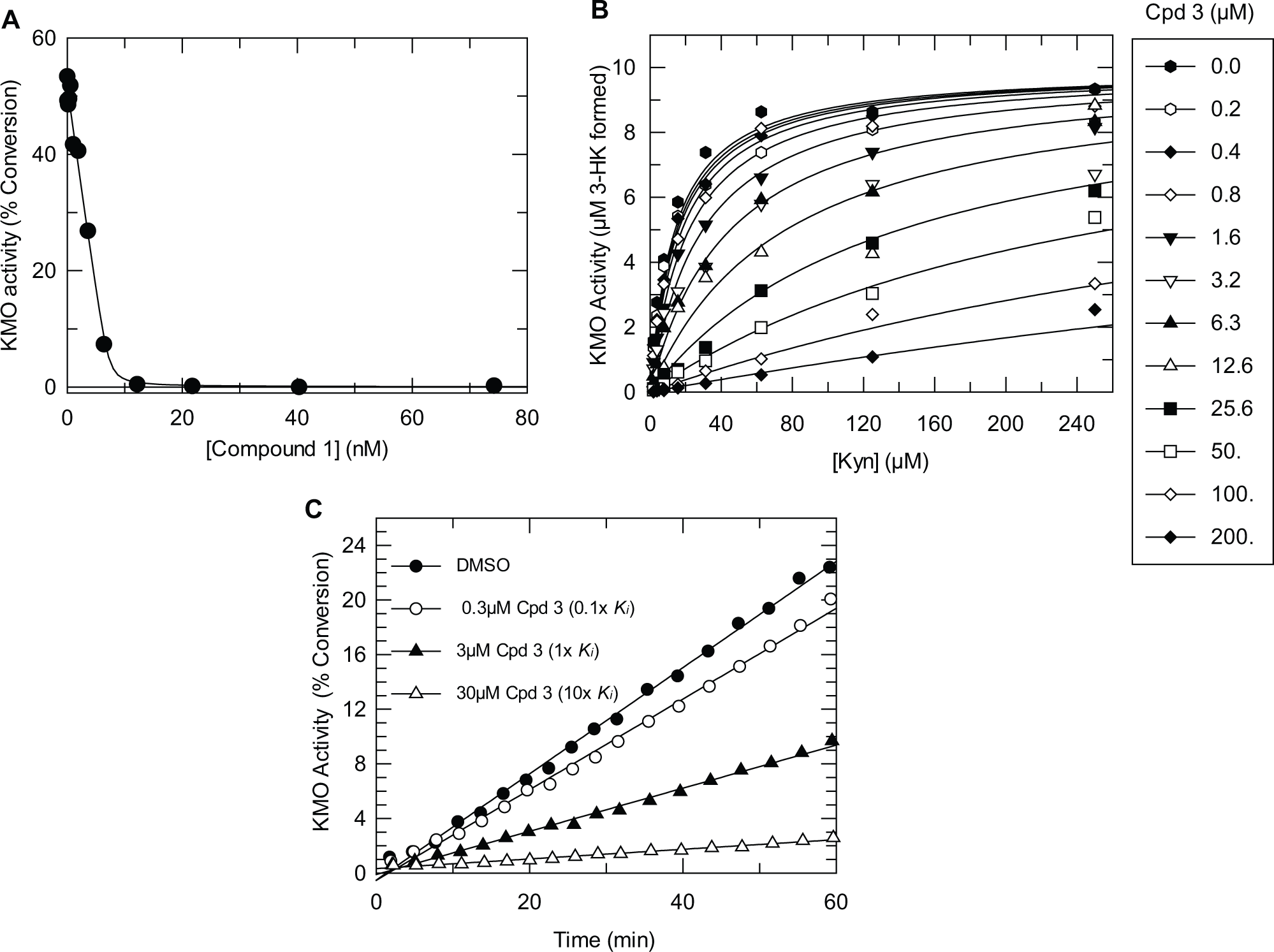
(
Screening Campaign
The RapidFire assay was used for a directed screen of approximately 78,000 compounds of low molecular weight (<300 kDa) and low lipophilicity (logP <3). Single-concentration screening was performed at 10 µM compound concentration. Performance was good, with an average Z′ of 0.8. The hit rate was 1.9% at a robust cutoff of 22.5%. A total of 247 compounds showed >50% inhibition, and 608 compounds were selected for concentration response testing in duplicate from a top concentration of 100 µM. A total of 264 compounds were flagged as active, with pIC50 >4.0, with 15 compounds displaying a pIC50 >6.0. Of these, a subset of 13 compounds was found to be falsely identified as hits due to their having an identical mass and product ion fragment as the substrate Kyn, resulting in an MRM signal from the compound appearing in the substrate channel. This gave an apparent increase in the denominator of equation 1 with increasing compound concentration, giving a false-positive result. Such compounds could be readily identified by analyzing the 3-HK product data only, which was subsequently adopted as the procedure for routine screening, with particular scrutiny of any compound with molecular mass ±1 Da of Kyn or 3-HK.
A steady-state kinetic analysis was performed on selected hits to establish whether the mode of inhibition was competitive with respect to Kyn, and a rapid-dilution experiment was performed to confirm reversibility. One example of a tractable hit identified by the screen is given in
Table 1
(compound 3), and the associated kinetic data are shown in
Figure 5B
In summary, an RF-MS assay has been successfully applied to a directed screen of human KMO using an isolated membrane preparation of recombinant origin. The screen delivered tractable low-molecular-weight hits, with acceptable biochemical mode of action, examples of which are currently being progressed to further in vitro and in vivo studies.
Footnotes
Acknowledgements
D.J.M. gratefully acknowledges the academic mentorship of Professor John Iredale and Professor O. James Garden, and the research support of Dr Xiaozhong Zheng, University of Edinburgh. D.J.M. and S.P.W. acknowledge the invaluable assistance of Dr Diane Harbison, University of Edinburgh/Edinburgh Bioquarter.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors declared receipt of the following financial support for the research, authorship, and/or publication of this article: D.J.M. is funded through a Health Foundation/Academy of Medical Sciences Clinician Scientist Fellowship and the MRC Developmental Pathway Funding Stream portfolio award to the University of Edinburgh.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
