Abstract
The fluorescence correlation spectroscopy (FCS)–based competitive binding assay to screen for protein-protein interaction inhibitors is a highly sensitive method as compared with the fluorescent polarization assay used conventionally. However, the FCS assay identifies many false-positive compounds, which requires specifically designed orthogonal screenings. A two-colored application of the FCS-based screening was newly developed, and inhibitors of a protein-protein interaction, involving selective autophagy, were selected. We focused on the interaction of LC3 with the adaptor protein p62, because the interaction is crucial to degrade the specific target proteins recruited by p62. First, about 10,000 compounds were subjected to the FCS-based competitive assay using a TAMRA-labeled p62-derived probe, and 29 hit compounds were selected. Next, the obtained hits were evaluated by the second FCS assay, using an Alexa647-labeled p62-derived probe to remove the false-positive compounds, and six hit compounds inhibited the interaction. Finally, we tested all 29 compounds by surface plasmon resonance–based competitive binding assay to evaluate their inhibition of the LC3-p62 interaction and selected two inhibitors with IC50 values less than 2 µM. The two-colored FCS-based screening was shown to be effective to screen for protein-protein interaction inhibitors.
Introduction
Autophagy is an important intracellular bulk degradation system that is well conserved from yeast to mammals. 1 Cells use autophagy to degrade their cytosolic components, including damaged proteins and organelles, for homeostasis and viability. 2 To date, a number of autophagy-related proteins (Atg) and their functions have been investigated. 3 Autophagy was found to participate in various physiological functions, including intracellular clearance, development, differentiation, programmed cell death, antigen presentation, and removal of invasive pathogenic organisms.1–3
In the process of autophagy, membrane structures, so-called isolation membranes, emerge and elongate to sequester intracellular macromolecules by forming double membrane structures, termed autophagosomes. The autophagosomes then fuse with lysosomes to degrade the contents. 4 Two ubiquitin-like conjugating systems, the Atg8-conjugation system and the Atg12-conjugation system, function in the formation of autophagosomes.5,6
In the yeast Atg8-conjugation system, the C-terminal arginine of Atg8 is processed by the cysteine protease Atg4. 7 Subsequently, Atg8 is activated by Atg7, an E1-like enzyme, and then transferred to Atg3, an E2-like enzyme. Finally, the C-terminal glycine of Atg8 is covalently conjugated to the amino group of phosphatidylethanolamine (PE). 6 Atg8-PE is localized on the autophagosomal membrane and is required for autophagosome formation. 7 In mammalian cells, microtubule-associated protein1 light chain 3 (LC3), GABAA receptor-associated protein (GABARAP), and Golgi-associated ATPase enhancer of 16 kDa (GATE-16) have been identified as Atg8 homologs. They are conjugated to PE through the same ubiquitin-like conjugating systems and are also localized on the autophagosomal membrane.8,9
In mammalian selective autophagy, p62, an adaptor protein, captures abnormal proteins for selective degradation. Then LC3 protein on the isolation membranes in the process of autophagosome formation interacts with the p62 protein. Through the LC3-p62 interaction, the targeted proteins are incorporated into the autophagosome to be degraded by lysosomes.10–12 The interactions between human LC3 and p62 are considered to be important for the autophagy related to various diseases, such as hepatic injury and neurodegeneration.13–15 Thus, specific inhibitors of the interaction are desired to clarify the detailed mechanisms of autophagy.
We previously reported the fluorescence correlation spectroscopy (FCS)–based competitive binding assay, for efficient screening of protein-protein interaction inhibitors. 16 The FCS-based assay is a highly sensitive method as compared with the fluorescent polarization assay used conventionally, although the FCS assay identifies many false-positive compounds, which requires specifically designed orthogonal screenings.16,17 In this article, we screened about 10,000 compounds by a TAMRA FCS-based competitive binding assay, and then an Alexa647 FCS-based assay and an surface plasmon resonance (SPR)–based assay were used to characterize the hits. We now report newly obtained inhibitors of the human LC3-p62 interaction and the two-colored FCS-based competitive binding assay as a general screening method.
Material and Methods
Preparation of the Human Atg8 Homolog, LC3 (1-120)
The DNA fragments encoding full-length human LC3 (1-120; Uniprot accession no. Q9GZQ8.3) and glutathione-S-transferase (GST; 1-220; pGEX-6P-1; GE Healthcare, Waukesha, WI) were cloned into the pET-28a (+) vector (Novagen, Madison, WI) as fusion fragments with an N-terminal hexahistidine tag and a C-terminal GST tag, and the proteins were expressed in Escherichia coli BL21 Star (DE3) (Invitrogen, San Diego, CA). The cells were grown at 37 °C, induced with 1 mM IPTG, and then further cultured for 4 h at 37 °C. The cells were suspended in phosphate-buffered saline buffer, pH 7.4, containing 0.1 mM PMSF, 0.2 mg/mL lysozyme, 5 µg/mL DNase, and 5 µg/mL RNase, and then were disrupted by sonication. The lysate was centrifuged, and after the supernatant was filtered through a 0.45-µm membrane (Millipore, Billerica, MA), it was applied to a GST Trap HP column (GE Healthcare), which was eluted with 50 mM Tris-HCl, pH 8.0, containing 10 mM reduced glutathione and 2 mM dithiothreitol. Subsequently, the eluted protein sample was applied to a His Trap HP column (GE Healthcare), which was eluted with a linear gradient of 50 to 500 mM imidazole in 20 mM Tris-HCl, pH 8.0, containing 100 mM NaCl. The eluted 6His-LC3-GST fractions were concentrated with an Amicon Ultra-15 filtration device (Millipore), and the buffer was exchanged to 20 mM Tris-HCl, pH 7.4, containing 100 mM NaCl. The total protein concentration was determined by a Bradford protein assay (Bio-Rad Laboratories, Hercules, CA).
Chemicals
For the screening, we used the compounds in the protein-protein interaction–focused library, stored in the Open Innovation Center for Drug Discovery ( http://www.ocdd.u-tokyo.ac.jp/) as 2 or 10 mM stock solutions dissolved in DMSO. The total set of 10,160 compounds has a mean molecular weight of 387.5 Da and an ALogP of 2.92.
FCS-Based Competitive Binding Assay
The FCS-based assay to detect protein-peptide interactions under solution conditions and to screen for compounds inhibiting the interaction was performed, as reported previously.16,17 Briefly, the diffusion time of a fluorescent peptide probe was determined by an FCS measurement, and then the autocorrelation function was calculated from the fluorescence intensity fluctuations caused by fluorescent molecules diffusing in and out of the detection volume. FCS measurements were performed on an MF20 fluorescence detection system (Olympus, Japan) using a 543 nm He/Ne laser line for illumination.
The p62-derived peptides (SGGDDDWTHLS: the p62-peptide, and TAMRA-SGGDDDWTHLS: the TAMRA-p62-probe) were purchased from Toray Research Center (Tokyo, Japan). The peptides were diluted to a final concentration of 1 nM in 10 mM HEPES-KOH buffer, pH 7.4, containing 150 mM NaCl and 0.005% (v/v) Tween 20, to suppress glass surface interactions. The probe peptide, the 6His-LC3-GST protein, and the compound were mixed and incubated at room temperature for 2 h before the FCS measurements. FCS measurements with an acquisition time of 10 s were repeated three times for each well. The percentage inhibition of each measurement was calculated by [(control diffusion time) – (measured diffusion time)]/[(control diffusion time) – (probe diffusion time)], where the control diffusion time was measured with a mixture containing 1µM LC3 and 1 nM probe molecule, and the probe diffusion time was measured with a mixture containing only 1 nM probe molecule. The χ2 value of each sample was determined as the mean value of the three single measurement data.
To assess the quality of the assay, the Z′ value was calculated as defined by the following equation: 1 – 3(SDcontrol + SDprobe)/(meancontrol – meanprobe), where SDcontrol is the standard deviation of the diffusion time obtained with the measurements of the control mixtures containing LC3 and the probe and SDprobe is that of the probe mixtures containing the probe without compound. 18
Secondary FCS-Based Competitive Binding Assay Using an Alexa647 Probe
For the second screening, we prepared an Alexa Fluor 647-labeled peptide (Alexa647-SGSG-SGGDDDWTHLS: the Alexa647-p62-probe). The CSGSG-p62-peptide was purchased from Toray Research Center and was reacted with Alexa647-maleimide (Invitrogen) according to the manufacturer’s instructions. The Alexa647-p62 probe was purified by high-performance liquid chromatography with an ODS-3 column (4.6 × 150 mm, 5 µm; GL Science, Japan), and the fluorescence of the eluted fractions was detected with a fluorescence detector (Ex 620 nm/Em 670 nm). FCS measurements using the Alexa647-p62-probe were performed using a 633 nm He/Ne laser line for illumination.
SPR-Based Competitive Binding Assay16,17
Competitive binding experiments were performed at 25 °C, using a BIACORE 3000 (GE Healthcare) SPR instrument. The biotinylated p62-derived peptide (Biotin-SGGDDDWTHLS), purchased from Toray Research Center, was attached to a streptavidin-coated gold surface (Sensor Chip SA; GE Healthcare). The peptide was immobilized to levels of approximately 400 RU in each flow cell.
The LC3 protein (LC3-GST, final concentration of 50 nM) was incubated for 2 h with each of the inhibitor compounds, in 10 mM HEPES-KOH buffer, pH 7.4, 150 mM NaCl, 0.005% Tween 20, and 0.5% DMSO, and then injected into the instrument. The binding of LC3 to the immobilized p62 peptide on the Sensor Chip SA was monitored at a flow rate of 30 µL/min. In the absence of an inhibitor, the LC3 protein was captured at levels of approximately 3000 RU. The binding levels of LC3 were determined from the differences in the SPR signals from the flow cells immobilized with the p62 peptide and the reference cell. Additional instrumental contributions to the signal were removed by subtraction of the average signal of blank injections from the reference-subtracted signal. Data were fitted using a sigmoidal dose-response regression algorithm in GraphPad Prism (La Jolla, CA).
Results and Discussion
Primary FCS-Based Screen for LC3-p62 Interaction Inhibitors
Structural analyses of LC3-p62 complexes by X-ray crystallography and nuclear magnetic resonance studies revealed that the side chains of tryptophan and leucine in the AIM motif (WXXL) of p62 interact with the hydrophobic pockets of LC3, which are also conserved among the Atg8 homologs (PDB code: 2K6Q).19,20 Ichimura et al. 21 reported that the three consecutive N-terminal aspartate residues (DDD) form hydrophilic contacts with the basic surface of LC3 (PDB code: 2ZJD). The importance of the DDDWXXL motif in p62 for the interaction with LC3 was also confirmed by mutation analyses.11,19,21 On the basis of these data, we designed and prepared the p62-derived peptides (SGGDDDWTHLS: p62-peptide, and TAMRA-SGGDDDWTHLS: TAMRA-p62-probe) to evaluate the LC3-p62 interaction. The binding of the TAMRA-p62 probe to the 6His-LC3-GST was first tested by the FCS measurements. The protein-peptide interactions were detected as the increased diffusion time of the TAMRA-p62 probe. As a result, the KD value of LC3 binding to the TAMRA-p62 probe was determined to be 500 nM ( Fig. 1 ).
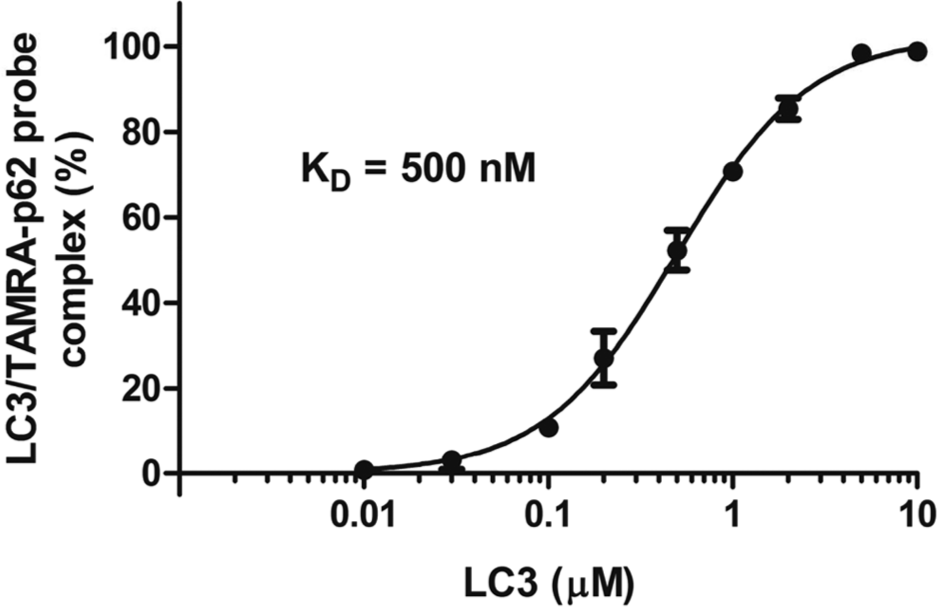
The LC3-p62 probe interaction, determined by the TAMRA fluorescence correlation spectroscopy assay. The TAMRA-p62 probe was incubated with the indicated concentrations of the LC3 (6His-LC3-GST) protein, and the diffusion times were measured and analyzed. The KD value for the TAMRA-p62 probe interaction with LC3 was determined to be 500 nM. The error bars represent means ± SD.
To screen the 10,160 compounds for LC3-p62 interaction inhibitors, we performed the FCS-based competitive binding assay 17 using the TAMRA-p62 probe. The assay mixture contained 1 µM 6His-LC3-GST, 1 nM TAMRA-p62 probe, and 10 µM of each compound in 10 mM HEPES-KOH buffer, pH 7.4, containing 150 mM NaCl, 0.5% DMSO, and 0.005% Tween 20. The binding activity of the TAMRA-p62 probe to the 6His-LC3-GST with the compounds was calculated as the percentage ratio of the diffusion time, in comparison to that without a compound ( Fig. 2 ). As the positive control for the screen, 20 µM of the nonlabeled p62 peptide was used, and the assay results are indicated by the open circles in Figure 2 . We selected positive compounds that decreased the binding activity of the TAMRA-p62 probe to LC3 and thus decreased the measured diffusion times. The mean Z′ value obtained per 384-well plate in this screen was 0.63 ± 0.076. Thus, the assay was considered to exhibit good quality for use in the screening. 18
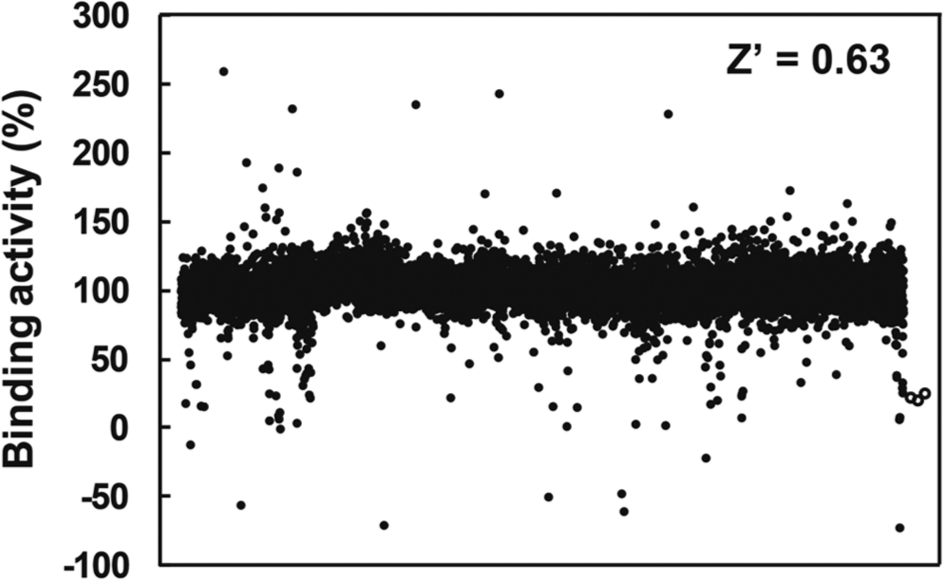
Primary screening of 10,160 compounds. The fluorescence correlation spectroscopy–based competitive assays were performed using the TAMRA-p62 probe. Open circles indicate the inhibition values of nonlabeled p62 peptide, used as a positive control. The mean Z′ value was 0.63 ± 0.076, calculated using the diffusion times measured with control and probe mixtures.
The χ2 value indicates the difference between fluorescent intensity distribution obtained by FCS measurement and that calculated by a fitting model. 16 The mean χ2 value of the measurements of control samples without compounds was 170.1, and the standard deviation was 44.8 in this assay. Thus, we omitted the results of 128 compounds with χ2 values that were more than the mean value plus 3 times its standard deviation (304.5) as void data, because the measurements with mixtures containing these compounds showed the hindered autocorrelation fitting and the results were not guaranteed to be accurate. Therefore, the screening of 10,032 compounds revealed 29 compounds that exhibited more than 70% inhibition of the LC3-p62 interaction at 10 µM.
Secondary FCS-Based Screen Using an Alexa647 Probe
To omit the false-positive compounds, the second FCS-based competitive binding assay was performed using the Alexa647-p62 probe. Because the Alexa647 probe has higher excitation/emission wavelengths than those of the TAMRA probe, the fluorescent measurements with the Alexa647 probe may be less susceptible to hindrance by autofluorescence of the compounds. In addition to the higher fluorescent wavelength, the fluorescence intensity per molecule of the Alexa647 probe is stronger than that of TAMRA, thus enabling more accurate autocorrelation analyses by the FCS instrument.
The binding of the Alexa647-p62 probe to the 6His-LC3-GST was tested by FCS, under the same conditions as those used with the TAMRA-p62 probe. The KD value for the interaction of LC3 with the Alexa647-p62 probe was 2.0 µM. We performed the FCS assays of the 29 hit candidate compounds, using an assay mixture containing 1 nM of the Alexa647-p62 probe and 4 µM LC3. The concentration of LC3 added to the mixture was higher than that used in the TAMRA probe assay. Therefore, the percentage inhibition by the p62 peptide (the positive control) was 35.4%, and weaker than that determined by the TAMRA probe assay (72.9%). The mean χ2 value of the control measurements without compounds was 98.4, and the standard deviation was 18.7, indicating the high reliability of the measurements with the Alexa647-p62 probe. A summary of the inhibition by each compound, measured by the FCS assay at 10 µM, is provided in Figure 3 . The mean Z′ value obtained per 384-well plate in this screen was 0.67. Thus, the assay was performed in good quality.
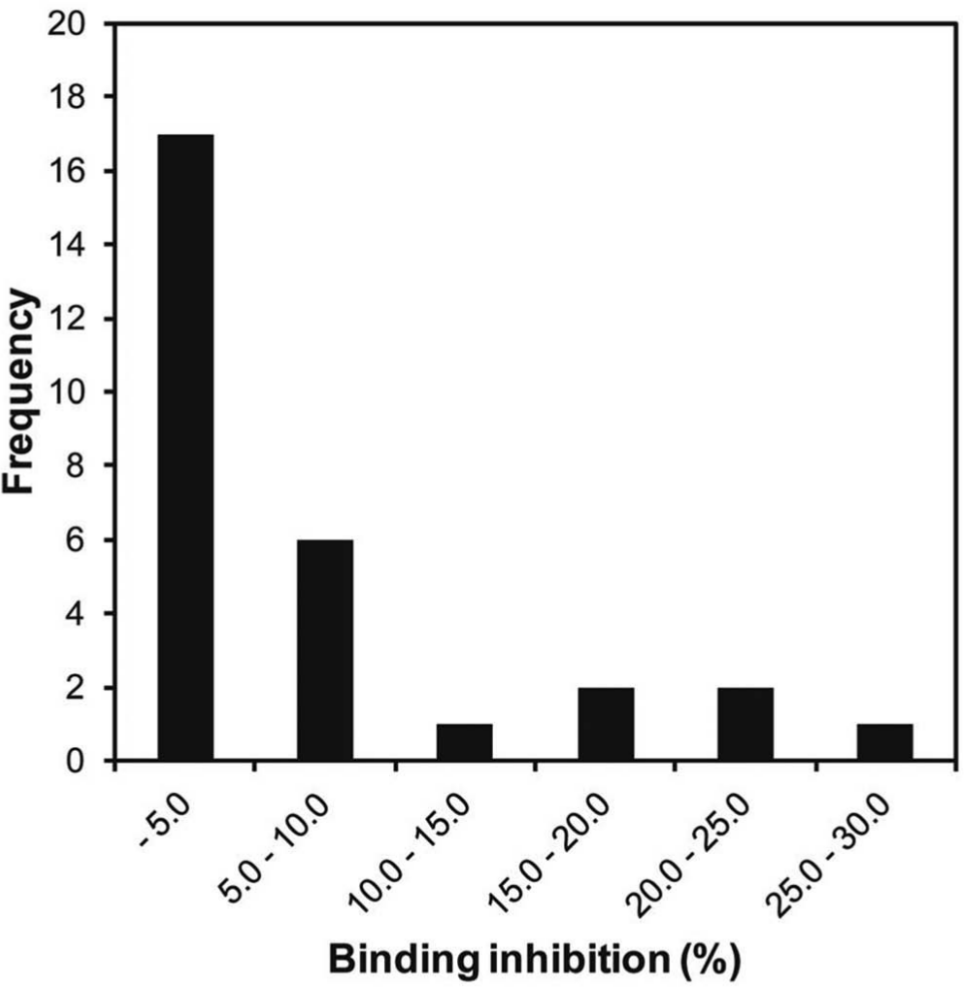
Alexa647 fluorescence correlation spectroscopy (FCS)–based assay of the 29 hit candidate compounds. The inhibition values were determined by FCS with the Alexa647-p62 probe. The numbers of compounds that exhibited the indicated inhibition values were counted and are shown as the frequency.
The calculated variability of diffusion time of control mixture, containing LC3 and the probe, existed 519.7 to 628.2 µs, which was calculated as follows: meancontrol (574.0 µs) ± 3 × SDcontrol (18.1 µs), where meancontrol and SDcontrol are the mean and standard deviation of the diffusion times obtained with the measurements of the control mixtures. It means that percentage inhibition values have a margin of error within ±9.4%. Considering variability of measured diffusion times, inhibition data of 19 compounds (<9.4%) were too weak to put confidence and those of the other four compounds (<–9.4%) were omitted as insufficient results. The remaining six compounds displayed more than 9.4% inhibition of the LC3-p62 interaction and are considered as candidate hit compounds. The false positives detected by the TAMRA-labeled probe assay may include compounds that interacted with TAMRA moiety and hindered the binding of the TAMRA probe to LC3 protein in the screening.
SPR-Based Competitive Binding Assay for True Hit Evaluation
To validate all 29 candidate hit compounds, we performed the SPR-based competitive binding assay with the Biacore 3000 instrument. The biotinylated p62-derived peptide was immobilized on a sensor chip, and assay mixtures containing 50 nM LC3 and each 10 µM compound were injected after a 2 h incubation. When 50 nM LC3 was injected, the amount of LC3 binding to the immobilized peptide was the half saturation level with a margin of error within ±7.6%. Thus, results of two compounds (<–7.6%) were considered as insufficient data. Figures 4a and b indicate the correlation of the percentage inhibition values of each compound obtained by the TAMRA (a) and Alexa647 (b) FCS assays and the SPR assay at 10 µM.
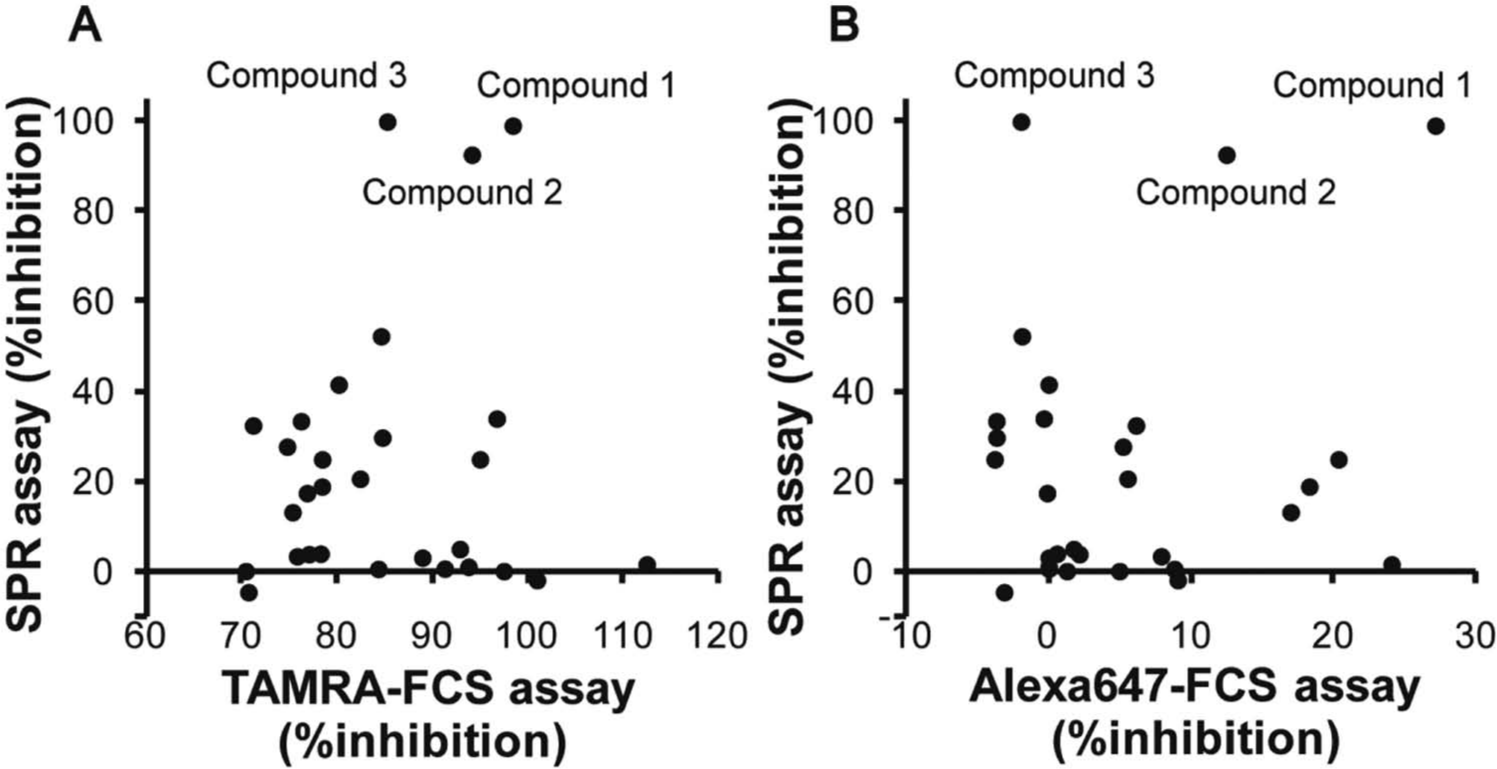
Characteristics of the 29 hit candidate compounds. The correlations of the percentage inhibition determined by the TAMRA fluorescence correlation spectroscopy (FCS) assay versus that by the surface plasmon resonance (SPR) assay (
Only three compounds (compounds 1, 2, and 3) showed inhibition of the interaction of LC3 with the immobilized p62 peptide. One of the reasons for the false positives may be that the hydrophobic compounds have affinities to fluorescent moieties of probes used in the FCS-based assays, bind the fluorescent probes, and then inhibit their interaction with the LC3 protein. Some other compounds may bind the LC3 binding pocket; however, they inhibit the interaction of LC3 with the fluorescent p62 probes only and not with the original p62 probe.
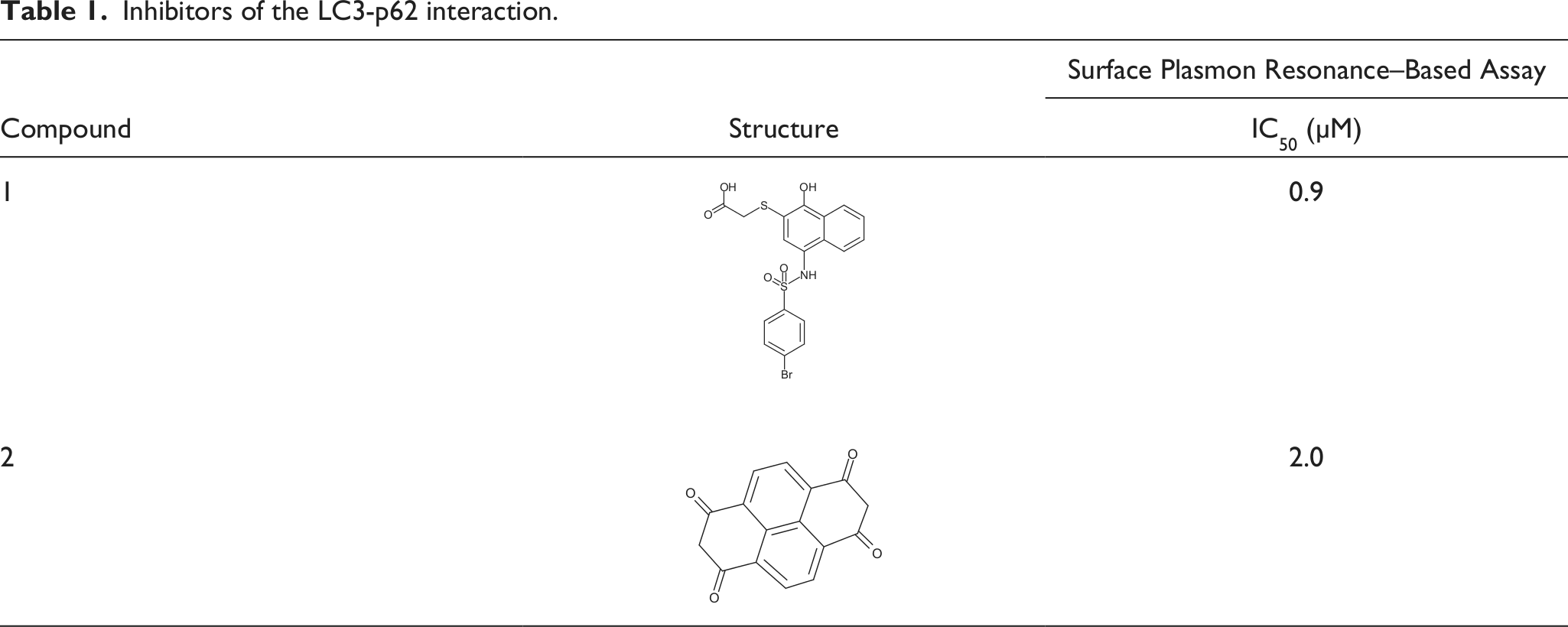
The inhibition dose-response curves for the compounds were determined ( Fig. 5 ). The IC50 value determined for compounds 1, 2, and 3 were 0.9, 2.0, and 3.5 µM, respectively. The dose-response curve for compound 3 was much steeper than the standard curve, so we selected compounds 1 and 2 as hit compounds, ( Table 1 ). We would like to test additional compounds with a similar structure to compound 1 to search for better inhibitors.
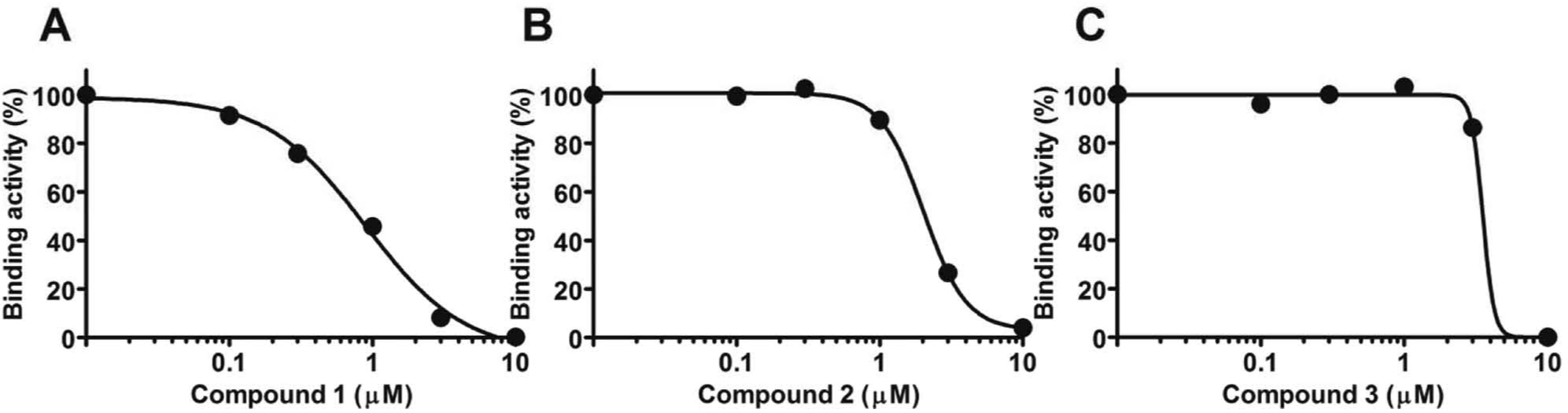
Binding inhibition dose-response curves for compounds 1, 2, and 3. Various doses of inhibitors or DMSO vehicle were added to the assay mixtures and incubated for 2 h before being injected into the BIACORE 3000 instrument.
Inhibitors of the LC3-p62 interaction.
As the result, we concluded that the two-colored FCS assay to screen for protein-protein interaction inhibitors is a useful general method. Considering data acquisition speed of the FCS measurement, the assay is better suited for screening less than tens of thousands compounds. We recommend the fluorescent-based high-throughput screening assay 17 to screen a large-scale library. The assay evaluated the protein-peptide interaction by detecting the quenching of fluorescence of GFP fused to a protein, after its interaction with a target peptide labeled with a fluorescence quencher.
Recently, it has been demonstrated that the targets degraded through selective autophagy in mammalian cells are involved in cellular regulation, oxidative stress responses, and diseases, including tumors and neurodegenerative disorders.22–24 The identification of LC3-p62 interaction inhibitors might lead to the development of valuable research tools for more detailed analyses of autophagic processes.
Footnotes
Acknowledgements
We thank the Open Innovation Center for Drug Discovery (The University of Tokyo) for providing chemical samples.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by Targeted Proteins Research Program, from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
