Abstract
The inhibition or dysregulation of apoptosis plays an intimate role in the initiation and progression of cancer by confounding normal tissue homeostasis. We currently do not have a clinical method to assess apoptosis induced by cancer therapies. Phosphatidylserine (PS) is an attractive target for imaging apoptosis because it is on the exterior of the apoptotic cells and PS externalization is an early marker of apoptosis. PS-binding peptides are an attractive option for developing an imaging probe to detect apoptosis using positron emission tomography. In this study, four peptides were evaluated for PS-binding characteristics using a plate-based assay system, a liposome mimic of cell membrane PS presentation, and a cell assay of apoptosis. This work also describes two screening techniques to enable researchers to identify and optimize compounds that bind to PS. The results of our study indicate that all four peptides bind to PS and are specific to apoptotic cells. Two of the peptides in particular that have an additional cysteine residue are good potential candidates for development into imaging probes because they bind to PS with high affinity and specificity and they can be easily radiolabelled with 18F.
Introduction
Apoptosis is extensively studied because it is associated with cancer and several other diseases. The ability of tumor cells to evade apoptosis is one of the hallmarks of cancer. 1 Techniques for detection and assessment of apoptosis at the cellular level are well developed. Less well developed are molecular imaging modalities for assessing apoptosis at the clinical level, particularly in the context of apoptosis induced by cancer therapies. One of the goals of personalized medicine in cancer therapy is the selection and tailoring of therapies that achieve maximum tumor destruction with minimum side effects. Rapid assessment of tumor response to therapy is clinically challenging and would be greatly aided by a reliable molecular imaging modality that targeted apoptosis.
Changes observed at the cellular level during apoptosis include phosphatidylserine (PS) externalization, caspase activation, chromatin and nucleus condensation, reduction in the volume of the cytoplasm, and DNA degradation. A number of these changes, particularly caspase activation and PS externalization, provide selective targets for potential molecular imaging probes. PS is an attractive target for imaging apoptosis because it is on the exterior of the apoptotic cells, it represents a high-capacity target, and PS externalization is an early marker of apoptosis.2,3 PS targeting with fluorescent annexin V has been widely employed for in vitro imaging and in vivo probes, based on labeling annexin V with radionuclides, and has been developed to allow noninvasive imaging of apoptosis using nuclear medicine techniques.4–6 Radiolabeled annexin V, however, has failed to gain wide acceptance as a clinical imaging agent due to a variety of limitations, including less than optimal biodistribution. The difficulties encountered with various forms of radiolabeled annexin V are reviewed by Kapty et al., 4 and these issues illustrate the continued need to identify a robust and clinically acceptable molecular imaging probe for apoptosis.
In the search for alternative strategies for targeting PS externalization, a variety of approaches have been exploited, including peptide selection using phage libraries and phage display technology. Peptides have a number of potential advantages as radiotracers including small size, good tissue diffusion, rapid targeting, low antigenicity, easy synthesis, well-developed radiolabeling, fast blood clearance, and inexpensive production.7–10 Recently, Burtea et al. 11 isolated two peptides, LIKKPF and PGDLSR, which have high affinity and specificity for PS. LIKKPF was further investigated as a conjugate with gadolinium diethylenetriaminepentaacetic acid (Gd-DTPA) as a possible apoptosis imaging probe for use with magnetic resonance imaging (MRI). Preliminary MRI studies in an animal model of liver apoptosis and the ApoE-/- mouse model of atherosclerosis were conducted but were only moderately successful because of the low sensitivity of the peptide–Gd-DTPA imaging probe, which limits detection of apoptosis in areas with low PS concentration.
The preliminary report of these peptides in animal models of apoptosis prompted us to investigate their application as radiopharmaceutical probes for positron emission tomography (PET) imaging of apoptosis. We decided to incorporate an 18F label using established prosthetic groups. The prosthetic groups N-succinimidyl-4-[18F]fluorobenzoate ([18F]SFB) and N-[6-(4-[18F]fluorobenzylidene)aminooxyhexyl]maleimide ([18F]FBAM) were investigated for the ability to radiolabel the peptides with 18F. 12 Although [18F]SFB is reactive with amines and can therefore be directly attached at the N-terminus of peptides, we encountered a variety of problems with the radiochemistry using this approach and investigated the thiol-reactive [18F]FBAM, where we achieved superior yields and purity of products. The [18F]FBAM approach, however, required the modification of the 6-mer peptides 11 to the 7-mer peptides CLIKKPF and CPGDLSR by the addition of a thiol-containing cysteine residue at the N-terminus. The affinity and specificity of these modified peptides was previously unknown. We here describe the evaluation of two novel peptides and two previously isolated peptides in vitro using a plate-based assay system, using liposome mimics of cell membrane PS presentation and using a cell assay of apoptosis.
Materials and Methods
General
Human recombinant annexin V-FITC was obtained from BD Biosciences (Franklin Lakes, NJ) and was used according to the manufacturer’s instructions. 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC), 1,2-dipalmitoyl-sn-glycero-3-[phospho-L-serine] (sodium salt) (DPPS), and cholesterol were obtained from Avanti Polar Lipids (Alabaster, AL). Peptides LIKKPF, PGDLSR, CLIKKPF, and CPGDLSR were purchased modified at the N-terminus with the fluorescent tag 5-carboxy fluorescein amidite (5-FAM) from LifeTein LLC (South Plainfield, NJ) with >95% purity. Binding buffer contains 10 mM HEPES (pH 7.4), 140 mM NaCl, and 2.5 mM CaCl2. All other reagents used were of analytical grade.
Fluorescence Plate Assay
The plate assay was used to test the affinity of peptides LIKKPF, PGDLSR, CLIKKPF, and CPGDLSR for PS. Annexin V was also used as a control. PS was immobilized on clear-bottom 96-well microplates (Corning Inc, Corning, NY) after solubilization in ethanol at a concentration of 1.0 mg/mL. 11 Wells were filled with 200 µL of this solution, and a film was formed after overnight evaporation of the ethanol at room temperature. The same procedure was used with phosphatidylcholine (PC) as control. PS- or PC-coated plates were incubated with 50 µL of peptide suspension for 1 h at 37 °C in the dark. The plates were washed three times with 50 µL binding buffer. Bound peptide was detected by a fluorescence plate reader FLUOstar OPTIMA (BMG Labtech GmbH, Offenburg, Germany), and PC software version V1.30 R4 (BMG LABTECH GmbH, Offenburg, Germany) was used for data collection.
Determination of the Dissociation Constant
The dissociation constant (Kd) of each peptide was determined by saturation experiments performed by the method described above. Serial peptide dilutions ranging from 10−4 M to 10−12 M (annexin V dilutions ranged from 10−7 M to 10−13 M) were prepared in binding buffer. The quantity of bound peptide obtained from the relative fluorescence unit values was plotted against the logarithmic concentration of total peptide input, and the curve was fitted according to a sigmoidal dose-response profile using GraphPad Prism version 5.04 for Windows (GraphPad Software, San Diego, CA). The peptide concentration at half-saturation corresponds to the Kd.
Preparation of Liposomes
For the PC liposome formulation (0 % PS), 1.2 mL of DPPC (20 mg/mL) and 1.2 mL cholesterol (4.89 mg/mL) were combined in a 250 mL round-bottom flask. For 10% PS liposomes, the solution was adjusted by the addition of 2.4 mL of DPPS (1.03 mg/mL). The solvent was removed under reduced pressure using a rotatory evaporator to form a thin film on the inside of the flask. 13 The dry lipid film was rehydrated in 2 mL of sodium acetate (1 M, pH 6.0). The solution was sonicated and extruded through polycarbonate membranes (pore size 2 µm, 25 mm; Nucleopore, Whatman, Piscataway, NJ).
Confocal Microscopy of Liposomes
Confocal microscopy analysis of liposomes was used to visualize the binding of peptides LIKKPF, PGDLSR, CLIKKPF, and CPGDLSR to 10% PS liposomes. The liposome formulation (10% PS) was washed twice with 500 µL PBS. The liposome formulation was then suspended in 500 µL binding buffer, and 200 µL of this suspension was transferred to an Eppendorf tube. Fluorescent peptides (0.1 nmol CLIKKPF and CPGDLSR, 0.01 nmol LIKKPF and PGDLSR) or annexin V (3 µL) was added to the tube followed by incubation in the dark for 15 min at 37 °C. After incubation, 300 µL binding buffer was added to the tube. Liposome samples were diluted for confocal microscopy; therefore, images of single liposomes were presented. Peptides samples were then analyzed using the Zeiss LSM 710 confocal microscope (Plan-Apochromat 10×/0.45 M27, Argon 488 laser) using 200 µL of sample in a dish plate (glass-bottom microwell dishes, 35 mm Petri dish, 14 mm microwell, 1.5 coverglass; MatTek Corporation, Ashland, MA). Annexin V stained liposomes were analyzed using the Zeiss LSM 510 confocal microscopy (40× 1.3 oil, Argon 488 laser) using 200 µL of samples in a dish plate.
Confocal Microscopy of Cells
Confocal microscopy analysis of cells was used to determine the binding specificity of the peptides CLIKKPF and CPGDLSR to apoptotic cells and to determine the cellular localization. Apoptosis was induced in Jurkat cells by seeding cells (2 × 105 cells) on 24-well plates (500 µL per well) and treatment with 2 µM camptothecin for 24 h. 11 Cells were pelleted by centrifugation at 500 × g for 5 min. Cells were washed with binding buffer. Each sample was resuspended in 20 µL binding buffer. Peptide (1.9 µM for CLIKKPF, 7.9 µM for CPGDLSR) or annexin V was added to cell suspension and incubated in the dark for 15 min at room temperature. Samples were spotted onto a glass slide for a minimum of 10 min to allow cells to adhere to the glass. Then 50 µL of 4% paraformaldehyde was added to the cell droplet and allowed to incubate for 5 min at room temperature. Slides were immersed in 1× PBS three times to wash samples. DAPI (20 µL of 0.3 µg/mL) was added to spotted samples for 3 min at room temperature. Slides were washed twice and allowed to dry before mounting. Slides were cured overnight to allow mounting media to set. Cells were then analyzed using the Leica SP5 confocal microscope (100×/1.44 oil objective, Argon laser 488 nm). The peptides were tested in apoptotic cells (camptothecin treatment) and healthy control cells (no camptothecin treatment).
Results
Fluorescence Plate Assay
The four fluorescently modified peptides were tested using a plate-based assay with annexin V- FITC as a positive control. The peptides all bound to PS, and the dissociation constant (Kd) for each peptide was determined ( Fig. 1 ). CLIKKPF (Kd = 1.9 µM) and CPGDLSR (Kd = 7.9 µM) have reduced affinity for PS compared with the original peptides LIKKPF (Kd = 0.69 µM) and PGDLSR (Kd = 0.40 µM), which do not contain an additional cysteine residue. Annexin V-FITC was found to have a dissociation constant of 6.6 nM in this assay and had a much higher affinity for PS when compared with the peptides ( Fig. 2 ).
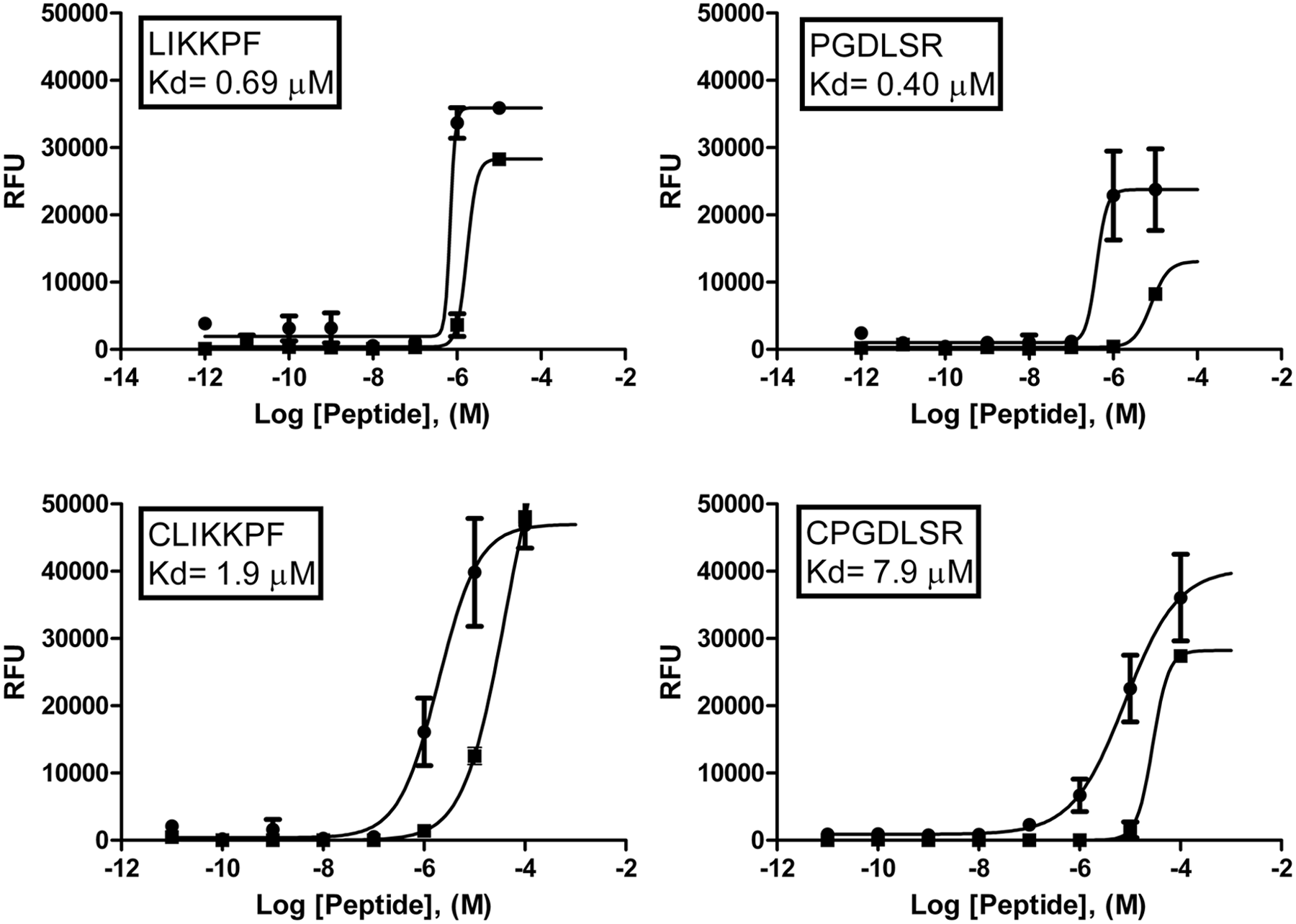
Binding curves for PS-binding peptides. Relative fluorescence units (RFU). PS: dark circles. PC: dark boxes. X-axis measurements in Log [Peptide], (M). The Kd in each box is for PS. The data presented are means ± SEM.
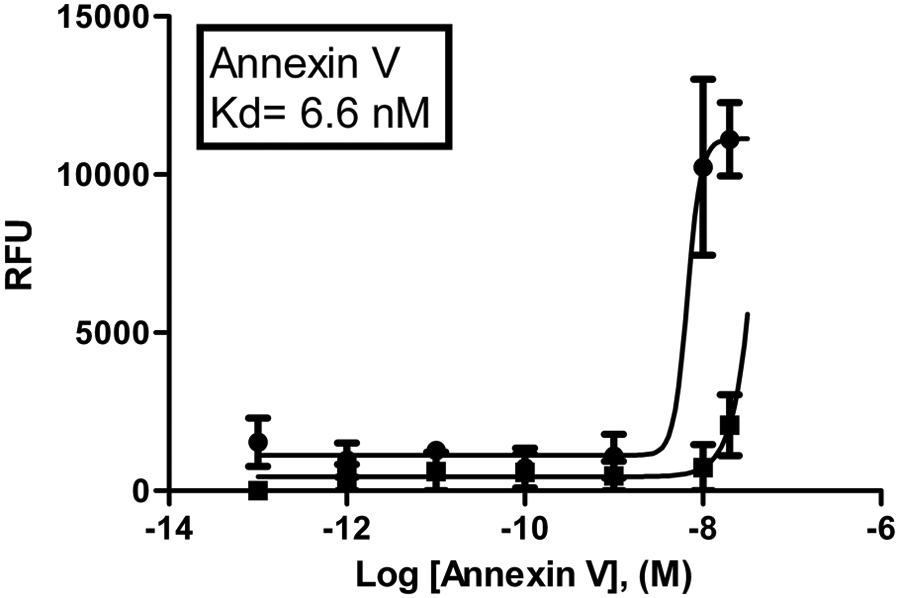
Binding curve for annexin V-FITC. Relative fluorescence units (RFU). PS: dark circles. PC: dark boxes. X-axis measurements in Log [annexin V], (M). Kd = 6.6 nM is for PS. The data presented are means ± SEM.
Confocal Microscopy Analysis of Liposomes Stained with Fluorescent Peptides
The fluorescent peptides and annexin V-FITC were tested with 10% PS liposomes and 0% PS liposomes (PC liposomes;
Figs. 3
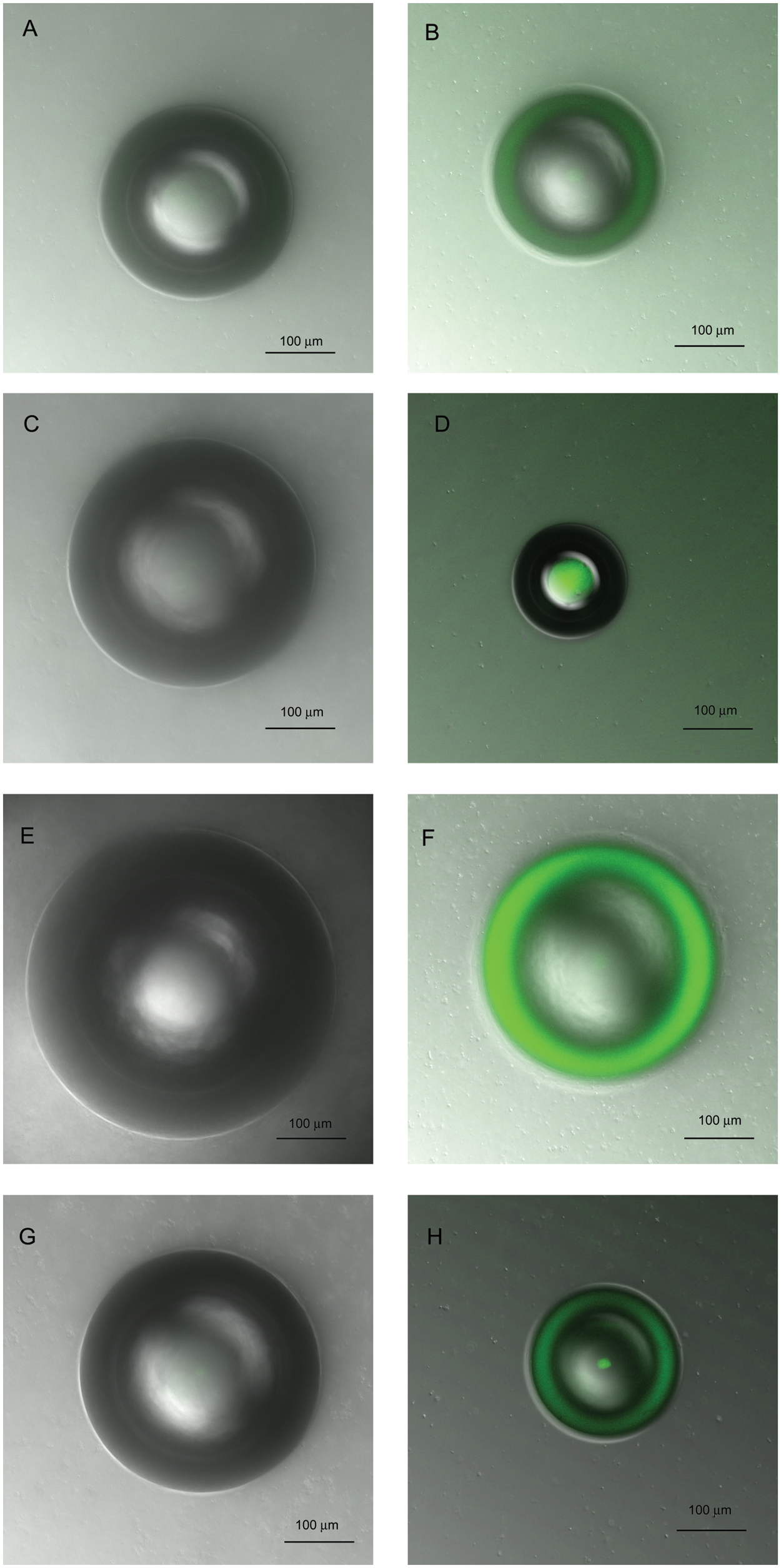
Confocal microscopy images of liposomes stained with fluorescent peptides. (
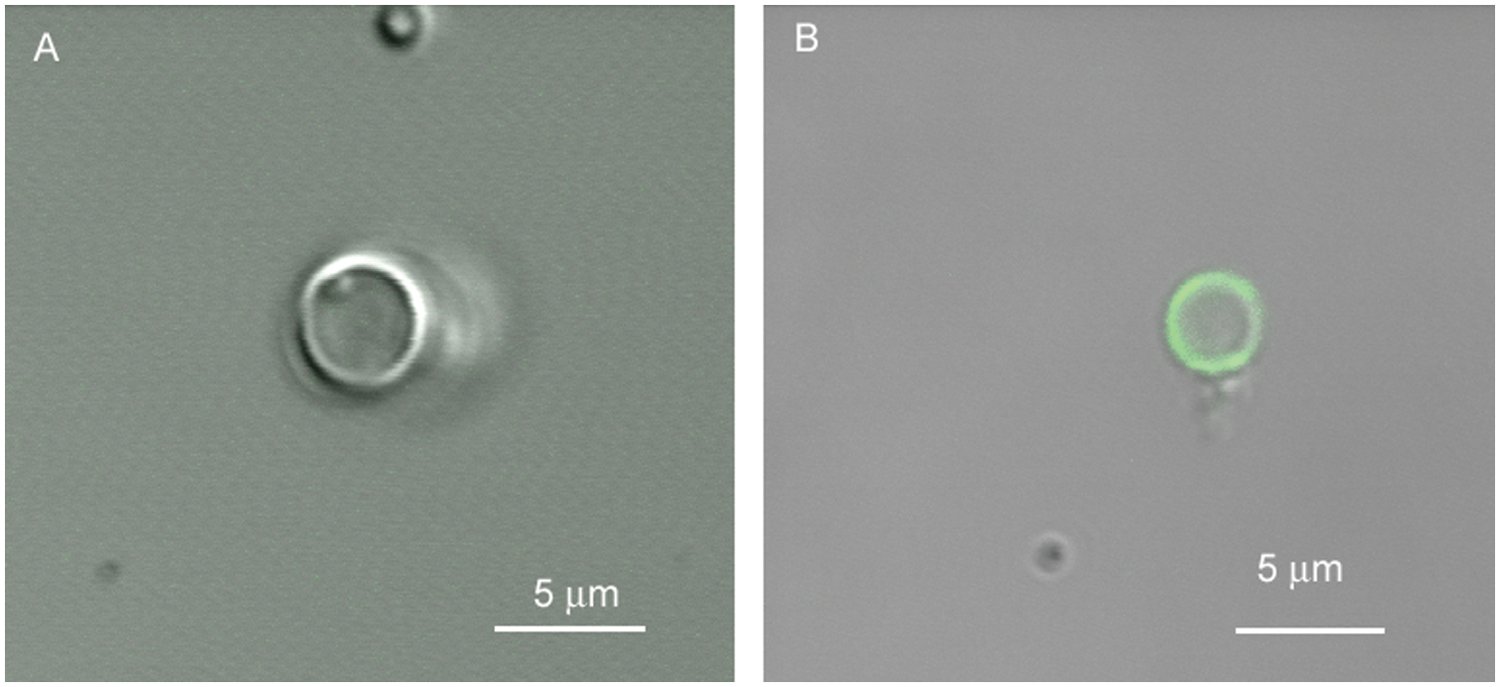

Confocal microscopy images of liposomes stained with annexin V-FITC. (
Confocal Microscopy Analysis of Cells Stained with Fluorescent Peptides
Cell studies with the peptides CLIKKPF, CPGDLSR, and with annexin V showed selective binding to apoptotic cells (camptothecin treated) and the absence of binding to healthy cells ( Fig. 5 ). Annexin V-FITC was observed at the cell membrane in the apoptotic cells. Although the peptides CLIKKPF and CPGDLSR showed generalized uptake in the cytosol and it is inferred the peptides also bound to the membrane, this was not proved definitely.
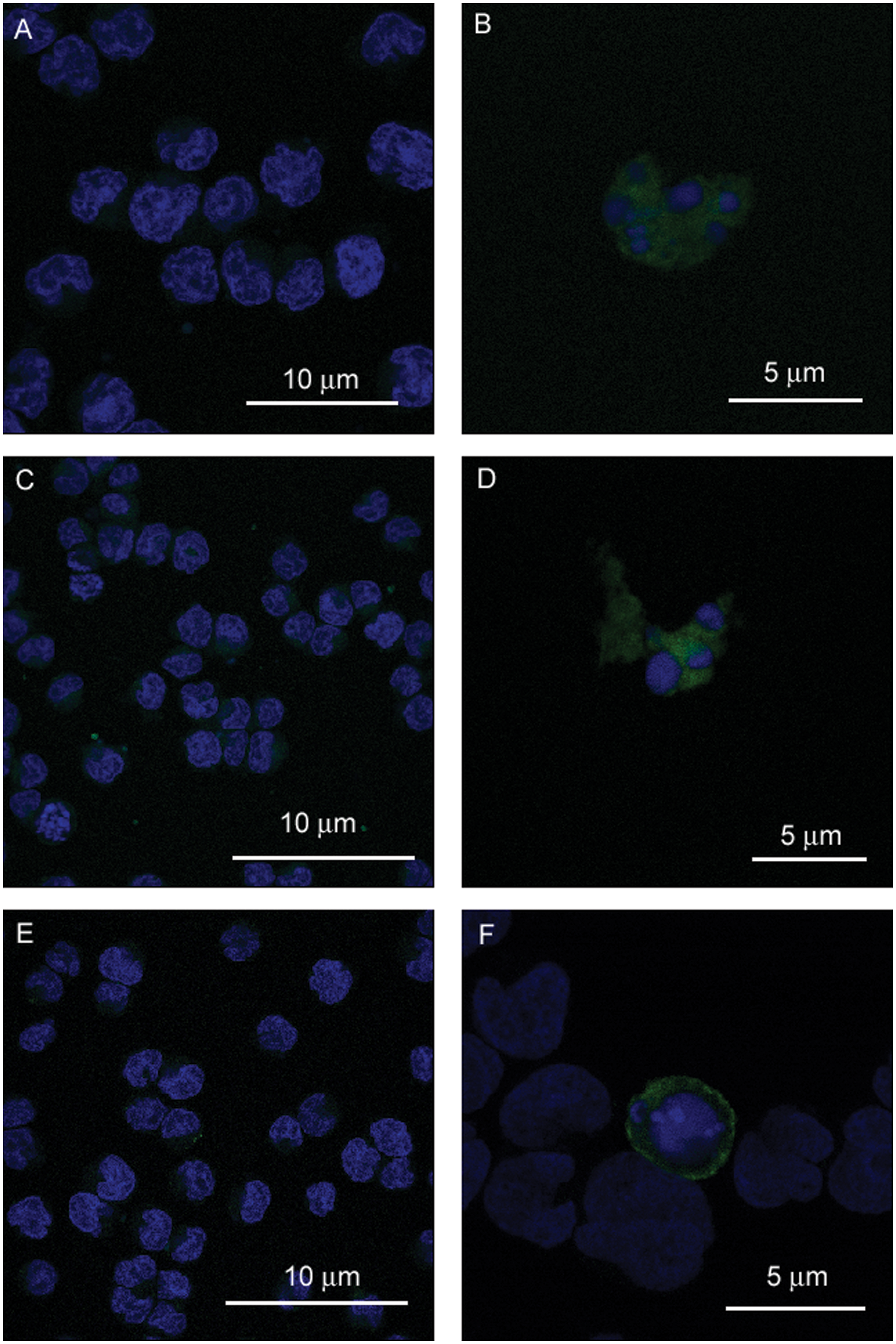
Localization of fluorescent peptides in Jurkat cells. Laser-scanning confocal microscope images were obtained to show staining of annexin V-FITC (green) or fluorescent peptides (green) in healthy and camptothecin-treated cells. Cells were also stained with DAPI (blue), which binds to DNA. (
Discussion
Despite intensive investigation, there is not yet a clinically acceptable molecular imaging agent targeting apoptosis. However, the recognition that apoptosis is an important process in human disease—whether caused by a pathological process such as a heart attack or whether induced through treatment such as chemotherapy—continues to encourage research in this area. The desired characteristics of an imaging agent for apoptosis include (1) rapid uptake and retention in apoptotic tissue, (2) rapid blood clearance of unbound tracer and rapid excretion, (3) selectivity for apoptotic cells and minimal uptake in healthy tissues, (4) generation of an on-target signal suitable for high-contrast molecular imaging, and (5) clinical acceptability in terms of availability, ease of preparation, cost-effectiveness, and biological compatibility. Peptides labeled with imaging radionuclides, and particularly PET-based probes, are an attractive option that can be potentially designed and selected to satisfy many of these requirements. The development of peptide-based imaging agents has been particularly fueled by the advent of phage display technology and its ability to identify binding sequences to a particular target from large libraries of random-sequence peptides. In this study, we investigated two PS-binding peptides described previously 11 (LIKKPF and PGDLSR) and two novel peptides produced by the addition of a cysteine residue (CLIKKPF and CPGDLSR). The addition of the cysteine residue provided a free thiol suitable for mild reaction with the 18F-containing prosthetic group [18F]FBAM. We have also evaluated these peptides to determine the binding characteristics using a plate assay and a liposomes assay to assess binding to PS.
Burtea et al. 11 used an enzyme-linked immunosorbent assay (ELISA) plate assay to test the binding affinity of phage peptides to PS. We have modified this method to allow affinity determination of our peptides by fluorescence measurements. Peptides can be readily modified to include a fluorescent tag for affinity studies with the advantage that these compounds can be used for further cell-based evaluation with confocal microscopy and flow cytometry. This method is applicable not only for peptides but also for proteins and oligonucleotides. Calcium is required for all four peptides and annexin V to bind to PS. Only background signal was observed when calcium was removed from the buffer.
The plate assay indicates that all four peptides bind to PS. The binding affinity of CLIKKPF and CPGDLSR is lower than the original peptides (LIKKPF and PGDLSR). This is likely due to the addition of the cysteine residue at the N-terminus, which could modify the PS-binding site on the small peptide. According to the methods used by Burtea et al., 11 the phage displayed LIKKPF has a Kd = 2 nM and phage displayed PGDLSR has a Kd = 1 nM. This is much higher affinity for PS than we determined for the peptides LIKKPF (Kd = 0.69 µM) and PGDLSR (Kd = 0.40 µM). However, the assay used by Burtea et al. 11 is an absorbance-based ELISA technique using phage peptides (unmodified) identified by anti-M13 monoclonal antibody conjugated to horseradish peroxidase. It has been previously reported that the binding affinity of the peptide is often lower than the original phage-displayed peptide. 14 This can be due to two reasons: First, a phage carries five copies of the hexapeptide that are able to interact with PS on the plate compared with a single peptide sequence in which only one copy interacts with PS. The second possible reason for increased affinity of the phage is that the phage coat protein may induce a conformational change in the peptide that results in a higher affinity for PS than compared with the conformation of the free peptide. In our study, we tested the purified free peptide, and Burtea et al. 11 reported the dissociation constant values for the phage-displayed peptides. The distinction between the phage and the peptide is important because the use of phages in vivo is highly unlikely because they are immunogenic.15,16
According to previous reports, human recombinant annexin V has a Kd = 0.5 nM. 17 Our determined value of annexin V-FITC in this plate assay was 6.6 nM. There are a few possibilities to explain this difference from the previous literature. It is known that the addition of a fluorescein modification alters the binding affinity of annexin V. 18 The affinity of annexin V to PS decreases as more amino groups are modified by the addition of an FITC molecule. 17 According to the supplier, there are two fluorescein labels per annexin protein, which may account for the reduced affinity of annexin V-FITC compared with human recombinant annexin V.
The liposome construct is designed to be similar to apoptotic cells with respect to PS externalization; in this regard, it can be considered a mimic of apoptotic cells with externalized PS. In cells, PS constitutes approximately 10% of total phospholipids in the lipid membrane,19–21 and during apoptosis, PS is externalized to the outer membrane layer of the cell. The results of the liposome assay indicate that all four peptides and annexin V bind to 10% PS liposomes but do not bind to PC liposomes (0% PS) according to confocal microscopy analysis. These results are in agreement with previous literature, 11 which show that LIKKPF and PGDLSR bind to plate immobilized PS. The results of the liposome assay support the data from the fluorescence plate assay in that all of the peptides bind to PS but not to PC at the concentrations tested.
The liposome assay presents a readily prepared formulation that provides a population that is similar to apoptotic cells with respect to the membrane presentation of PS. This assay has a number of advantages over conventional cell-based assays, including the ease of preparation, the ability to modify PS concentration, and the elimination of the requirement to induce and track the progress of apoptosis in cell studies. Liposomes can be analyzed by confocal microscopy or flow cytometry because they are sufficient size for these analytical methods. The designed composition of the liposomes provides a reproducible test system for comparing and contrasting results between labs working on PS-binding constructs.
The cell assay indicates that CLIKKPF and CPGDLSR bind specifically to apoptotic cells and do not bind to healthy cells. Microscopy imaging indicates that CLIKKPF and CPGDLSR are present in the cytosol, and it is inferred that they also bind to the membrane bilayer. This is possibly because of the small size of the peptides, relative to annexin V, which may allow them to cross the cell membrane, but the exact nature of the uptake and retention of the probes is unknown. Notably, fluorescence from the PS-targeted peptides was observed only in the apoptotic cells and not at the membrane or in the cytosol of healthy cells. Annexin V localized as expected in our studies at the cell membrane of apoptotic cells and was not evident in the cytosol. The cell-based studies serve to support the results of the plate and liposome assay, providing strong evidence that CLIKKPF and CPGDLSR are PS-binding peptides that have a high specificity for apoptotic cells. The cell data also provide a validation of the simpler assay techniques provided by plate immobilized PS and by PS liposomes. The peptides LIKKPF and PGDLSR were not tested to determine if they bind to apoptotic cells because this was previously reported. 11
Both of the noncell assays can be easily reproduced, can be used to test a wide variety of concentrations of a compound, and do not require extensive lab material and equipment required for cell culture techniques. These assays can also be applied to other classes of compounds such as small molecules, proteins, and oligonucleotides. The plate assay has been used previously by our group to screen several nucleotide sequences derived by computational modeling, which were designed to bind to PS. 22 These assays have allowed us to rapidly test modifications made to our lead compounds to determine if binding affinity is improved or impaired. Both of these assays may prove to be valuable tools in the rapid in vitro assessment of potential apoptosis targeting probes that bind to PS and could help to accelerate discovery of compounds that can be used for imaging apoptosis. The two novel PS-binding peptides described in this study are presently being evaluated in a murine tumor model of apoptosis as candidates for molecular imaging as their 18F-labeled analogs.
Footnotes
Acknowledgements
We gratefully acknowledge the technical help of Gerry Baron from Imaging Facility, Department of Oncology. We would like to thank the Alberta Cancer Research Institute (ACRI) for financial support for this research.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received funding from the Canadian Institutes of Health Research. JK received an Alberta Cancer Research Institute Graduate Studentship award.
