Abstract
This report describes the implementation of an automated work cell with commercially available hardware and software, capable of handling up to 15 separate reagents for performing 96-well or 384-well assays but with a small footprint and only a single liquid dispenser and two plate washers. Extremely flexible software was used to enable this simple work cell to perform processes that would traditionally require a much larger, more expensive automation platform. With the development of the C-Myc assays for the targets DYRK, BMX, PERK, and FAK, the authors describe a software solution to multibatch assays to run simultaneously, reducing reagent dead volume and increasing the efficiency of running multiple assays such that the time to generate data across multiple targets was significantly shortened. Although a larger automated system with multiple robotic arms and extensive equipment would also be able to process multiple assays simultaneously, the work cell we have described represents an inexpensive and flexible, easily upgradable option suitable for a wider range of labs.
Keywords
Introduction
C
In high-throughput cell screening, a single assay may use most of the available equipment and resources for several days. Automation platforms for high-throughput cell assays often consist of multiple pieces of identical equipment, either forming an equipment pool to treat plates identically in parallel or dividing individual assay steps between different pieces of equipment. This way of working often requires a work cell with a large footprint and multiple robotic arms and can be expensive to implement. In secondary cell screening, maximizing throughput in an automated work cell is often a secondary consideration compared with increasing flexibility. In a centralized screening group consisting of a large number of scientists, the number of data points per assay per week is often far below the maximum throughput of each assay. However, as cell assays are often complex with multiple steps, availability of equipment can often become the limiting factor in a busy screening lab. Consequently, although automated work cells that can extend working hours can appear to be an attractive solution, they must be able to handle a large number of different assay formats and workflows with minimal user intervention.
In the Oncology Assay Sciences group in AstraZeneca, a wide range of cell assays are processed on a weekly basis. These use adherent and nonadherent cell lines, as well as cell lysates, and may simply require the addition of a single detection reagent before reading or may require more than four separate stages, each consisting of a plate wash, reagent addition, and incubation. In addition, the final detection steps include ArrayScan and Acumen endpoints, as well as whole-well fluorescence or absorbance endpoints using a range of plate readers. Although the compound dosing and incubation steps for each of these cell assays were already fully automated using an integrated Labcyte Echo work cell, the variation in assay formats and workflows across the screening group meant that the remainder of the assay process was always performed using standalone equipment.
To meet the demand for a flexible automation system that could be deployed to support a variety of project cell assays with high reliability requirements, AstraZeneca has worked with Process Analysis & Automation (PAA) to develop a small and inexpensive custom-built automated work cell termed the AutoAssay system that could successfully run the resource-intensive stages of assays in between compound incubation and final plate reading. This system was developed to automate any 96-well or 384-well format assay involving a liquid dispenser and/or plate washer with multiple steps and incubation periods. Design requirements for the system were that it should effectively integrate peripheral devices already proven and validated in ongoing cell assays in the screening group, offer a large number of different reagent additions despite having only a single liquid dispenser, be simple to use in a multiuser environment, and be easy and inexpensive to reconfigure when assay processes develop and projects change.
To batch assays on any automated system, however, there needs to be a standardized process in place with very few unique variables. This is because testing on any platform requires several adjustments to be made to the hardware and software procedures, and all of these variables need to be assessed at the same time. The transfer of manual enzyme-linked immunosorbent assays (ELISAs) to fully automated workstations can require various time-consuming steps and having a more standardized process of optimization in place can greatly reduce development timelines. 2 In one example of the use of this type of system, a generic process was designed that uses the same c-Myc tag on four different targets in an ELISA format. 3 This standardized approach meant that only a few adjustments to the automation processes were required once the original setup was complete. However, as the original software setup on the AutoAssay systems did not allow multiple assays to be run simultaneously, the generic c-Myc method could not be taken advantage of, and each 4-hr duration assay had to be run separately. The most beneficial use of an automated system, however, is its potential to batch several screening assays at once, which provides flexibility and efficiency. 4 Therefore, additional development work was undertaken on the software controlling the two systems, which allowed these multiple assays to be run in parallel.
In this article, we describe the design and implementation of the AutoAssay work cells and the development and validation of a procedure to run multiple standardized assays concurrently for four different targets (FAK, PERK, DYRK, and BMX, as described by Macmillan et al. 3 ).
Materials and Methods
All materials, cell culture and transfections, ELISA plate preparation, and compound treatment of plated cells were performed as described by Macmillan et al. 3
AutoAssay system from PAA
The custom-built work cell (layout as shown in
OVERLORD2 software with event-driven scheduling was used by PAA to generate and operate the automated procedures, including all robotic plate-handling operations and controlling the programs and actions of all third-party equipment. This included the initial priming of the Multidrop Combi and PW384 plate washers and the automatic washing through of the Multidrop Combi head between each different reagent by switching reagent lines (via the Hamilton valves) to a reservoir containing distilled water and instructing the Multidrop Combi to conduct a series of priming and emptying steps. At the end of each assay run, the Brandel Waste Unit fed from the two PW384 plate washers was automatically emptied into the connected drain. Any errors on the system during a run were reported to the end user via an automatically generated e-mail.
Batch Manager software was custom designed by PAA for AstraZeneca to interface the end user with OVERLORD2 and was used to generate individual assay procedures composed of a number of assay stages (in these experiments, each stage involved a plate dispense and/or wash followed by a room temperature incubation), protocol names of the methods used on the reagent dispenser and/or plate washer during each stage, incubation times at the end of each stage, and prestage operations (e.g., additional priming or rinsing of the dispenser or plate washer and emptying of the waste trap during the assay). In addition, each assay procedure could also be supplied with estimated information detailing the volume of each reagent and wash solution required for a single plate at each stage, allowing the system to calculate required volumes for individual assay runs.
In addition to setting up standard operating procedures for each assay, Batch Manager provided the end user with a simple way to run the system on a day-to-day basis. New batches could be created with the following options: choice of assay procedure, number of assay plates, plates with or without lids, choice of reagent line for each addition, start and end locations for assay plates whether at room temperature or at 4 °C, and delayed start date and time if required. At assay setup, the end user was presented with an estimated required volume for each reagent and wash solution, based on the total number of assay plates and the estimated volumes per plate detailed in the procedure. Multiple batches could be added to the system in a queue, and the interface listed all queued, currently running, or completed batches. During the assay, the end user was provided with continuously updated details of the running stage or location of each assay plate, and at completion, all assay details and final timings were saved in an accessible database.
Generic ELISA procedure
The generic ELISA method described by Macmillan et al., 3 part I, was transferred into a Batch Manager procedure. In each assay, plates were washed three times with 400 µL of plate washer solution. For targets PERK, DYRK, and BMX, comparisons were made between running each assay manually (using standalone devices: a bench-top Matrix Wellmate for all reagent additions and a PW384 plate washer for all wash steps) and running each assay fully automated on the AutoAssay work cell. The plate washer programs were identical between the standalone equipment and the integrated equipment.
Analytical method and data analysis
The plates were analyzed using a TECAN Safire II plate reader with a fixed fluorescence setting at excitation wavelength 340 nm (10 nm bandwidth) and emission wavelength 465 nm (10 nm bandwidth). The method used an optimal gain calculated from the first plate in the stack.
There were several measures of data quality assessed for this study. The window between the mean maximum (Max) and mean minimum (Min) fluorescent signals was used to calculate a Z′ factor for every plate tested. 5 The coefficient of variation (CV) for the controls was also assessed for each plate of compounds tested. In addition, dose-response curves of several reference compounds were run in each assay to show a correlation with historical data. The IC50 for each compound tested was calculated by creating a response curve using the results from the 12-point dose range (30 µM to 0.00003).
Results
Implementation of the c-Myc ELISA on the AutoAssay system
Initial setup of the system indicated that the AutoAssay system could be used for running cellular screens that required multiple dispenser and/or washer steps.
The validation testing for the assays on the AutoAssay system showed that the Max and Min signals did show an increase in fluorescent signal compared with assays run on standalone equipment for some of the targets investigated, but neither the Z′ or the dose-response curves were significantly altered. Results are shown in
Table 1
and
Modular Method Using Standalone Equipment versus AutoAssay System for Targets PERK, DYRK, and BMX a
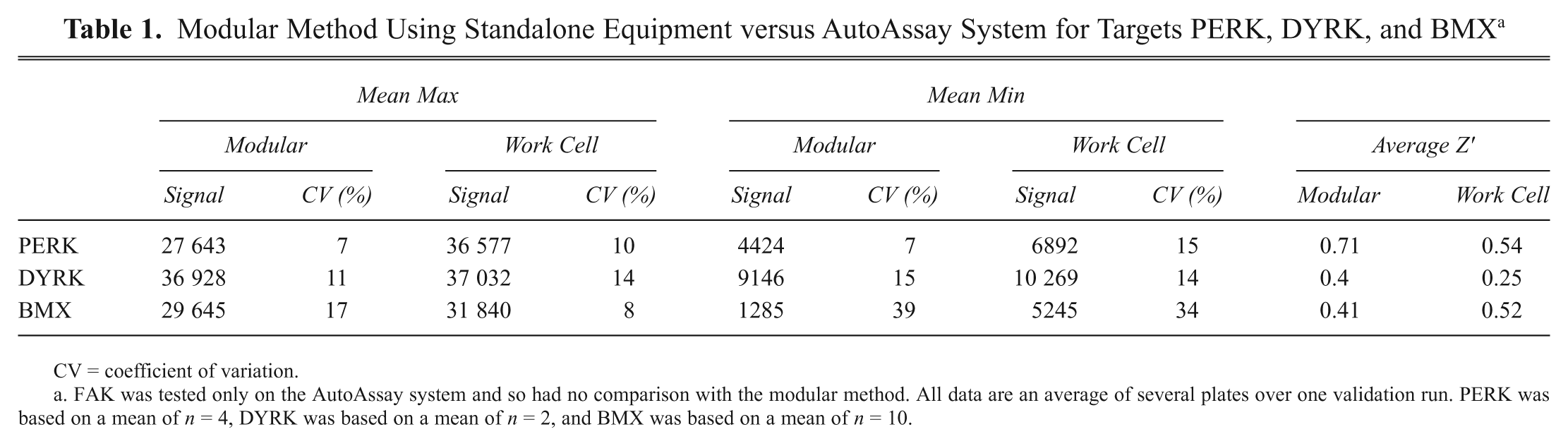
CV = coefficient of variation.
FAK was tested only on the AutoAssay system and so had no comparison with the modular method. All data are an average of several plates over one validation run. PERK was based on a mean of n = 4, DYRK was based on a mean of n = 2, and BMX was based on a mean of n = 10.
FAK was validated only on the automated work cell and not on standalone equipment as initial assay development occurred directly on the AutoAssay system. The results showed an 11-fold window between control signals (Max average = 34 407, Min average = 3175), and the mean Z′ for each run was 0.51.
PERK showed a 6.2-fold window for the controls using standalone equipment (mean Max = 27 643, mean Min = 4424). The mean Z′ for this assay was 0.71. The validation data set generated from using the AutoAssay system showed that the signal for both controls increased slightly (mean Max = 36 577, mean Min = 6892) but did not greatly reduce the window (5.3-fold). The Z′ calculated for the fully automated run was also slightly lower (0.54) but was acceptable for a cellular assay. The correlation between the PIC50s for the modular and AutoAssay system was very good (R2 = 0.95) and indicated no significant difference in data.
For DYRK, the manual method showed a 4-fold window between controls (mean Max = 36 928, mean Min = 9146), and the mean Z′ was 0.4. When moved to the AutoAssay system, there was an impact on results because the background increased slightly and the window was lowered to 3.6-fold (mean Max = 37 032, mean Min = 10 269), thus reducing the average Z′ to 0.25. The pIC50s showed a good correlation (R2 = 0.68), although poorer than other assays, and it was deemed that the move to full automation did have an impact on data quality.
The results generated for BMX validation showed that the assay had a very good window of ˜23-fold between the controls (mean Max = 29 645, mean Min = 1285), and the average Z′ for the assay was 0.41. When validated on the AutoAssay system, there was an increase in signal seen for both controls (mean Max = 31 840, mean Min = 5245) and a reduction in the assay window (~6-fold). However, there was an increase in the average Z′ for the batches (0.52), and the CVs were lower (<15% on the AutoAssay system compared with 20% on standalone equipment). The correlation between the pIC50s generated for the two methods is very good for this target (R2 = 0.925), and there was no shift in potency for the IC50s.
Implementation of multibatching on the AutoAssay system
In Macmillan et al., 3 a standardized process was designed to ensure that all four c-Myc ELISAs used the same liquid dispenser and plate washer methods, all required the same Quanta-Blu reaction and stop solutions, and all had the same incubation periods. Only the primary and secondary antibodies differed between the assays. The OVERLORD2 runtime engine is a real-time, event-driven scheduler, ideal for applications in which every assay plate must be handled in exactly the same way and in which incubation times must be kept constant. However, in its original configuration, the AutoAssay system was unable to run the four c-Myc assays simultaneously (multibatching) to make the most efficient use of both time and reagent dead volumes.
To allow multibatching, a new OVERLORD2 procedure was written to allow the Multidrop Combi to change reagents during each of the two antibody addition stages rather than only between them. This used loops within the programming of each assay stage in which a software variable increased incrementally as each assay plate was processed. When the value of this variable reached a predetermined limit (i.e., after a certain number of assay plates had been processed), this would trigger a separate procedure to change the Multidrop Combi valve system from one antibody reagent stock to another, via a washing step with distilled water. At the point of batch creation, the end user set up the AutoAssay System in Batch Manager as a normal single assay and input the total number of assay plates to be run, including all plates in all assays. During the system initialization step, the end user was then presented with a series of questions to determine how the multibatching should run. The answers to each question provided the system with the predetermined variable limits that would trigger any reagent changes.
Multiplexed assay batching on the AutoAssay system had the outcome of larger plate numbers being processed through the system in a single run, rather than smaller plate numbers spread between separate runs. As reliable volume sensors for the waste trap were unavailable, the end user was initially required to manually empty the Brandel waste trap at the end of each run; however, the volume of wash solution used during these multibatch runs frequently exceeded the 25 L capacity of the trap container. To avoid this capacity limit, an additional prestage operation was included in the Batch Manager assay procedure to empty the waste trap prior to each new stage. A set time period for the waste pump to operate was used, automatically calculated during the assay as the number of plates in the entire run multiplied by the number of seconds required for the waste pump to empty the wash solution from a single assay plate.
Assay performance
Figure 1 shows the intra-assay and interassay variation for FAK, PERK, and BMX (DYRK was not transferred to the AutoAssay system). The data showed that the signals for the Max and Min controls remained consistent over a run and that the Z′ values varied little throughout a run. The Z′ values for FAK remained fairly constant and varied from 0.4 to 0.7 over eight runs and within a single run. PERK showed a slightly higher variability, ranging from 0.35 to 0.75 over nine runs; however, this is biased by one run that had a high Min signal of more than 10 000, and excluding this run gave a Z′ from 0.6 to 0.75 for the eight remaining runs. The intra-assay data for PERK indicated that there was very little difference between results as the data remained fairly constant at about 0.5 to 0.6. BMX was very tight over the six runs shown, and the Z′ remained between 0.5 and 0.7. Table 1 also shows that using the AutoAssay system could maintain low CVs (in most cases less than 15%) and for some targets improved the variability from the modular process (e.g., BMX max CV = 17% for modular and 8% for AutoAssay).
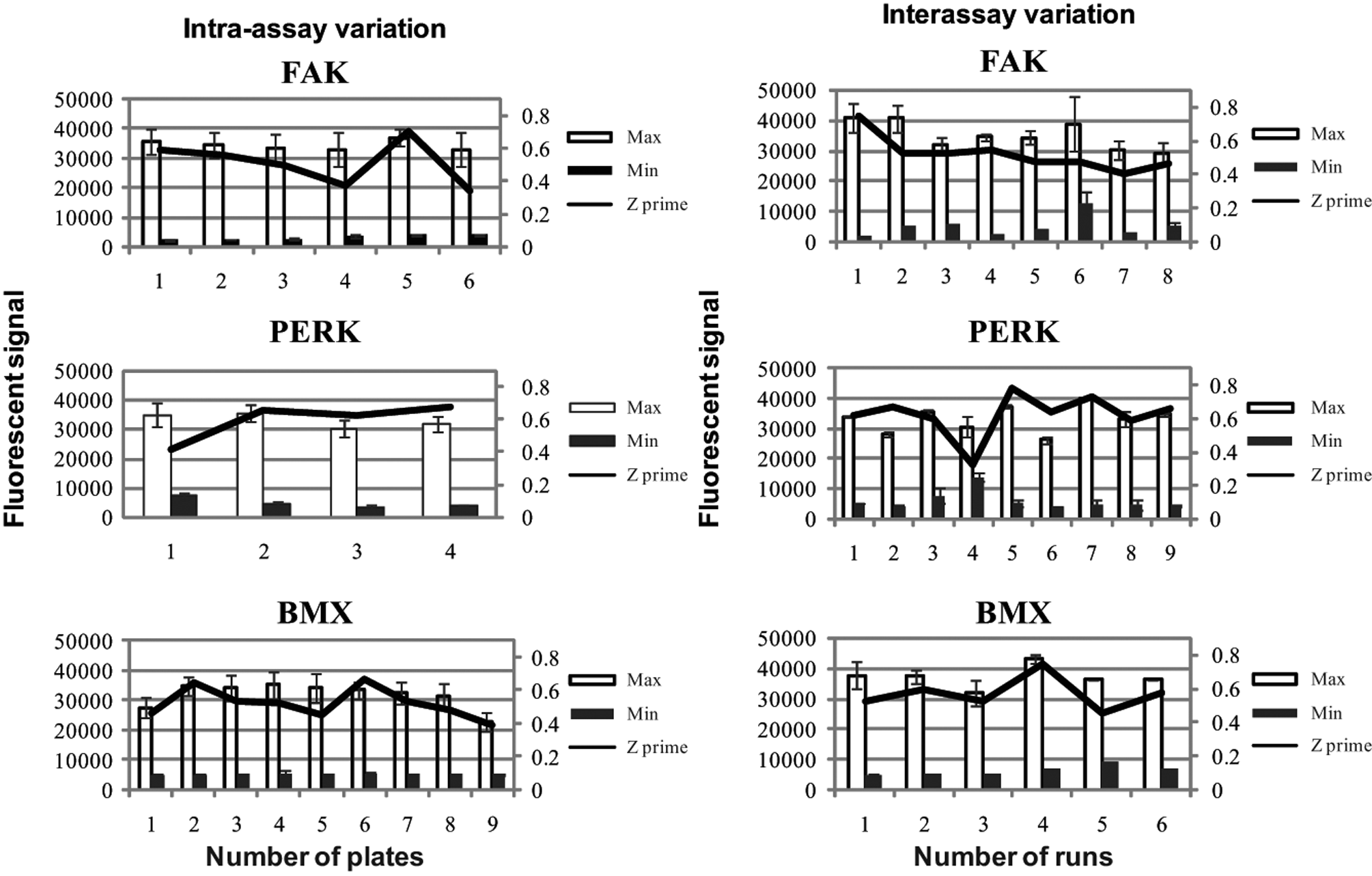
Intra-assay and interassay variation using the AutoAssay system for targets FAK, PERK, and BMX. It was deemed that DYRK could not be run using the work cell without further optimization. All standard deviations are based on n = 17 Max and Min controls run on each plate. For interassay variation, a minimum of n = 2 was carried out in each validation run for each of the targets investigated.
Discussion
Implementation of the c-Myc ELISA assays on the AutoAssay system
When implemented on the AutoAssay system, three of the assays showed data similar to the modular method and could be processed using the integrated system. The assay for DYRK, however, did not work well on the automated system. This might have been due to the fact that this particular target already had a small window between controls before testing on the system, and the ELISA procedure might have required further optimization before being moved to the work cell. The results for assay performance on the AutoAssay system implied that there was no significant impact on the data quality for the remaining three targets assessed and the system could improve the data quality and reproducibility from run to run (see Fig. 1 ). This may have been because the system is set up to treat each plate exactly the same, that is, each plate gets exactly 1 hr incubation and takes exactly the same time to process, thereby reducing some of the inherent viability seen with cellular assays that are processed manually.
The valve system upstream of the liquid dispenser allows a large number of reagents to be used on the system at the same time. Although it was originally expected that stringent washing routines would be required between each reagent, the different reagents used in this study required only simple washing with distilled water between reagent line changes to prevent cross-contamination.
The AutoAssay system brought together off-the-shelf equipment with customized software and was designed to increase the efficiency and throughput of assays that had been previously processed manually. Although in this report optimization of each assay was originally done offline and assays were transferred onto the system only when optimization was complete, new assay builds can be optimized directly on the system, reducing validation timelines. However, more complicated factorial experimental design procedures can be achieved with other automated platforms and so-called smart robotics.6–8
Implementation of multibatching on the AutoAssay system
The programming solution for multibatching used in this study was limited to a maximum of four primary and four secondary antibody steps; however, introducing extra steps and options to allow more assays to be run concurrently would require only trivial changes to the program. The system can have a maximum of 15 reagent additions on any one run and can do an unlimited number of stages as long as a multidrop and/or a plate washer is used. For this work cell, incubation times are the limiting factor—a more advanced scheduling system would be required to allow multiple assays with different incubation times to be run concurrently. However, this would have a detrimental impact on reagent dead volumes, as the single liquid dispenser would require a wash procedure between every addition.
Cost and resource benefits of multibatching on the AutoAssay system
The implementation of the integrated AutoAssay system for use in cellular screening, and in particular its use to multibatch assays, has helped the running of cellular screens to become more efficient in the Assay Sciences group. Historically, each cellular assay was run by a single user, meaning that each assay had its own associated cost and requirement for automation and resource. Unless there was access to multiple pieces of automation, each assay had to be staggered to run at separate times or over consecutive days, meaning that it could take time to generate compound data across multiple targets. Each assay also required an excess volume for all the reagents to cover the dead volume associated with each piece of automation. This meant that there was significant wastage of common reagents across all the assays. The implementation of the AutoAssay system has enabled the running of multiple assays in a single procedure that can now be operated by a single user. In addition, although the work cell will not reduce the time taken to run the assay, it has meant that there is a resource saving because the initial setup of the system takes only 15 min. This approach of batching multiple assays together has made significant efficiency savings through reduced resource requirement, reduced reagent wastage, and reduced requirement for multiple pieces of automation equipment. Compound data across multiple targets can now be generated more quickly and within a single run. As an example, prior to implementation of multibatching, four separate target assays would have been run either by four separate users over several days or a couple of users over a week. After introduction of the AutoAssay system, one user can run four separate target assays in 2 days ( Fig. 2 ).
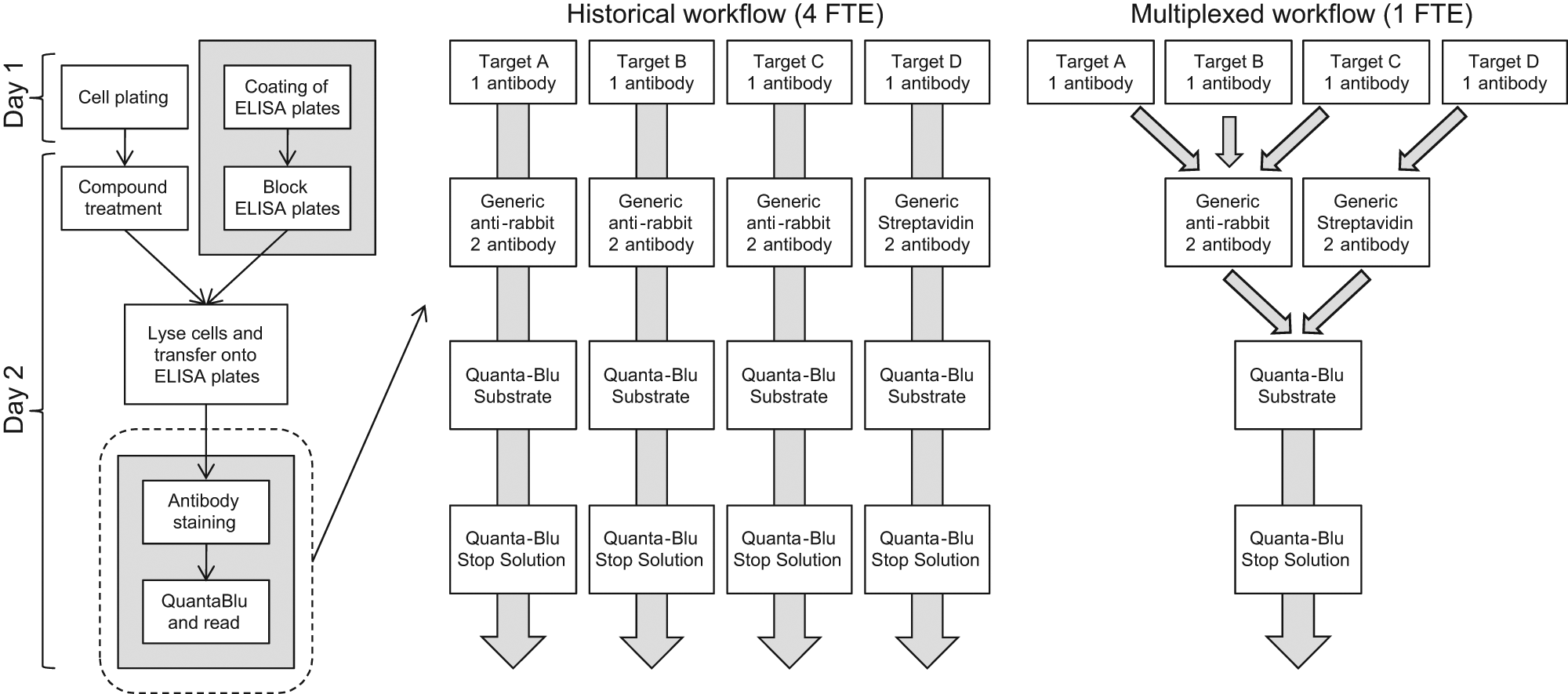
Schematic of the generic layout of the enzyme-linked immunosorbent assay procedure and the effects the process changes have on the efficiency of running several assays and the impact on resource. The sections in the gray boxes are those currently run using the AutoAssay system. The right-hand figure shows how the workflow has improved from the previously run process to allow a single scientist to run all four targets as one assay, rather than four different assays run by four different scientists (with associated time and equipment conflicts).
Further uses and improvements
Although this article describes the automation of c-Myc ELISAs, this system can also be used for multibatching other types of cellular assays. For example, we now use this system to run multiple imaging assays from the fix stage through measurement on high-content readers (Acumen or ArrayScan) with high success. This is possible because the system’s air filters allow hazardous chemicals such as formaldehyde to be used within the work cell. The implementation of this system across a range of assay formats within AstraZeneca’s Oncology Cell Assay Sciences group has led to significant increases in efficiency and learning for future cellular assay delivery.
An external shelf was installed on the AutoAssay system (shown in
The AutoAssay system contains both a 96-pin head and a 384-pin head plate washer to allow the processing of plates in either format. However, as more cell assays become validated in 384-well assay plates, a simple upgrade to the system would be to have two separate 384-pin head plate washers. As the washing step can be the rate-limiting factor in terms of the number of plates that can be processed per hour, the ability of the system to automatically alternate between two identical plate washers would significantly reduce the time between individual plates and increase the throughput of the system.
Footnotes
Acknowledgements
We thank Eric Tang and Jonathan Wrigley at AstraZeneca and Rob Harkness, Jon Newman-Smith, Andy Mash, and Ian Pye at PAA for their contributions. The involvement of many staff at AstraZeneca in c-Myc cellular assays has been invaluable. Marie Macmillan was contracted to AstraZeneca via Morson International, Adamson House, Centenary Way, Salford, Manchester, M50 1RD, England.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
