Abstract
Background
Cardiovascular diseases (CVDs) are the leading cause of global morbidity and mortality. Long non-coding RNAs (lncRNAs) have emerged as potential diagnostic biomarkers for CVDs.
Aim
To systematically evaluate the diagnostic value of lncRNAs for CVDs through a systematic review and meta-analysis.
Methods
Studies focusing on lncRNAs as diagnostic markers for CVDs were searched from PubMed, Embase, and Web of Science databases. The meta-analysis was conducted according to the PRISMA 2020 guidelines. The diagnostic value of lncRNAs was calculated by pooled sensitivity, specificity, and area under the summary receiver operating characteristic (SROC) curve (AUC). Meta-regression and subgroup analyses were performed to explore heterogeneity sources. Publication bias and sensitivity analysis were also conducted.
Results
A total of 16 articles (18 lncRNAs, 1125 controls, and 1855 cases) were included. LncRNAs showed pooled sensitivity of 0.81 (95% CI = 0.76-0.85) and specificity of 0.86 (95% CI = 0.80-0.90) for CVDs diagnosis. The SROC analysis revealed an AUC of 0.90 (95% CI = 0.87-0.92), indicating high diagnostic accuracy. The positive diagnostic likelihood ratio (DLR) was 5.73 (95% CI =4.07-8.08), and the negative DLR was 0.22 (95% CI = 0.18-0.28). The diagnostic score was 3.25 (95% CI = 2.79-3.71), the diagnostic odds ratio was 25.77 (95% CI = 16.22-40.93). The expression trend of lncRNAs, control type, and disease type were the main sources of heterogeneity.
Conclusion
LncRNAs had high sensitivity and specificity for CVDs diagnosis, showing great potential as auxiliary diagnostic tools.
Keywords
Introduction
Cardiovascular diseases (CVDs), including myocardial infarction (MI), chronic heart failure (CHF), acute coronary syndrome (ACS), and coronary atherosclerotic heart disease (CAD), are the leading cause of global morbidity and mortality, posing a significant burden on public health.1,2 The pathogenesis of CVDs is intricate and multifaceted, involving a combination of genetic, lifestyle, metabolic, and inflammatory factors.3–5 Recent evidence highlights the critical roles of oxidative stress, inflammatory responses, apoptosis, and fibrosis in the development and progression of CVDs. 6 Timely and accurate diagnosis is crucial for effective treatment and prognosis of CVDs. CVDs are usually diagnosed by coronary angiography, electrocardiogram, or echocardiogram. 7 Common diagnostic biomarkers in clinical practice include high-sensitivity cardiac troponin (hs-cTn), creatine kinase-MB (CK-MB), and B-type natriuretic peptide (BNP). 8 However, these biomarkers can be affected by other diseases, reducing their diagnostic accuracy. 9 Thus, identifying new biomarkers with higher specificity and sensitivity is crucial for early CVD diagnosis and treatment. 10
Long non-coding RNAs (lncRNAs), noncoding RNAs longer than 200 nucleotides, have gained attention as potential biomarkers and therapeutic targets in various diseases, including CVDs.11,12 Liu et al found that lncRNA HOTAIR was downregulated in cardiomyocytes after ischemia/reperfusion (I/R), and could regulate apoptosis and cell cycle via fused in sarcoma. 13 LncRNA HCG15 is involved in cardiomyocyte apoptosis and inflammation by activating the NF-κB/p65 and p38 pathways. It was correlated with cTnT and could distinguish the MI patients from the healthy controls. 14 LncRNA PELATON is upregulated in ACS patients and has high diagnostic and prognostic value for it. 15 Additionally, the diagnostic value of lncRNAs in CVD has gained increasing attention. 16
Evidence suggests that lncRNAs can serve as diagnostic and prognostic biomarkers for CVDs. 17 Some studies show that individual lncRNAs have high sensitivity and specificity, while others suggest that combining them with traditional markers can improve diagnostic accuracy.18,19 Meta-analysis integrate data from multiple studies, quantifies sources of heterogeneity, and evaluates the overall diagnostic performance of lncRNAs, providing more reliable evidence for clinical use.
Given the increasing body of literature examines lncRNAs in CVDs, a systematic review and meta-analysis are essential to synthesize existing evidence and evaluate the diagnostic accuracy of lncRNAs as biomarkers. This study comprehensively assesses the diagnostic value of lncRNAs in CVDs, providing insights into their potential for clinical application and guiding future research directions.
Materials and Methods
Literature Selection Strategy
Relative studies were selected from PubMed, Embase, and Web of Science databases by two authors, respectively. A search formula was designed using the following keywords (Supplementary Table 1): “long non-coding RNA” or “lncRNA”; and “cardiovascular disease” or “coronary artery disease” or “myocardial infarction”; and “diagnostic biomarker” or “sensitivity” or “specificity” or “ROC curve” or “detection” or “screening”. Articles were selected up to the start of this study (May 10, 2025). No language restrictions were applied. This study was conducted in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) 2020 statement. The completed PRISMA 2020 checklist is presented in Supplementary File 1.
Inclusion and Exclusion Criteria
After excluding duplicate literature, the remaining articles were independently screened by two researchers, with those not meeting the inclusion criteria excluded. Discrepancies were resolved through discussion.
Inclusion criteria included: (1) Patients diagnosed with heart failure, myocardial infarction, coronary atherosclerosis, or coronary syndrome. (2) Studies focused on diagnostic markers for cardiovascular disease. (3) Diagnostic markers were lncRNAs. (4) Studies provided sufficient data, including sample size, sensitivity, specificity, and AUC. (5) LncRNAs were derived from serum and detected by real-time quantitative polymerase chain reaction.
Studies with one of the following characteristics were excluded from this study: (1) Study type was meta-analysis, case report, or review. (2) Studies were animal experiments, cell/molecular mechanism studies. (3) Studies unrelate to cardiovascular diseases (heart failure, myocardial infarction, coronary atherosclerosis, or coronary syndrome). (4) Incomplete data.
Data Extraction and Quality Assessment
A standardized table was designed to extract the following information: first author, publication year, country, age, and sex of participants, sample size, control type, disease type, lncRNA name and expression trend, sensitivity, specificity, and AUC. The Quality Assessment of Diagnostic Accuracy Studies - 2 (QUADAS-2) tool was used to evaluate risk of bias in diagnostic studies.
Meta-Analysis
Data integration was performed using Stata 12.0 software. Pooled sensitivity, specificity, and area under the summary receiver operating characteristic (SROC) curve (AUC) were calculated to evaluate the diagnostic value of lncRNAs for cardiovascular diseases. Clinical applicability of lncRNAs was examined via positive diagnostic likelihood ratio (DLR) and negative DLR. Diagnostic score and diagnostic odds ratios were used to assess the diagnostic accuracy. Positive DLR is defined as the probability of a positive test result in diseased individuals versus non-diseased. Positive DLR >5 suggests moderate clinical utility. Negative DLR is the probability of a negative test result in diseased versus non-diseased. Negative DLR <0.3 suggests moderate exclusion capacity. Diagnostic odds ratio is the ratio of positive versus negative odds. When it more than 10 indicates strong diagnostic utility. Diagnostic score defined as the natural logarithm of the DOR, which linearizes the metric for meta-regression analysis. Forest plots for key subgroup was plotted by Review Manager 5.3. Heterogeneity was assessed using the I2 statistic, with I2 > 50% indicating significant heterogeneity. Covariates for bivariate random-effects meta-regression and subgroup analysis were pre-specified based on control type (classified as healthy individuals and others), age of the control group (continuous, centered at 60 years), disease type (categorized as AMI and others), age of the disease group (continuous, centered at 60 years), and expression trend of lncRNAs (categorized as up-regulated vs down-regulated per original studies). Between-study variance (τ2) were estimated using the REML estimator. τ2 was reported separately for sensitivity and specificity to quantify absolute heterogeneity on the logit scale. Publication bias was assessed using Deeks’ funnel plot. The stability of the results was verified by a sensitivity analysis. Statistical significance was set at P < 0.05.
Results
Literature Selection and Characteristics of Included Literature
A total of 1858 articles were retrieved from PubMed, Embase, and Web of Science databases. After removing the duplicated articles, 465 articles were screened by title or abstract. After deleting 64 reviews, 2 meta-analyses, 202 non-diagnostic studies, 80 studies not focused on CVDs, and 58 studies not focused on lncRNAs, 29 articles were included for the full text article selection. Finally, 16 articles15,18–32 (including 18 lncRNAs, 1125 controls, and 1855 cases) were enrolled for the meta-analysis (Figure 1, Table 1).
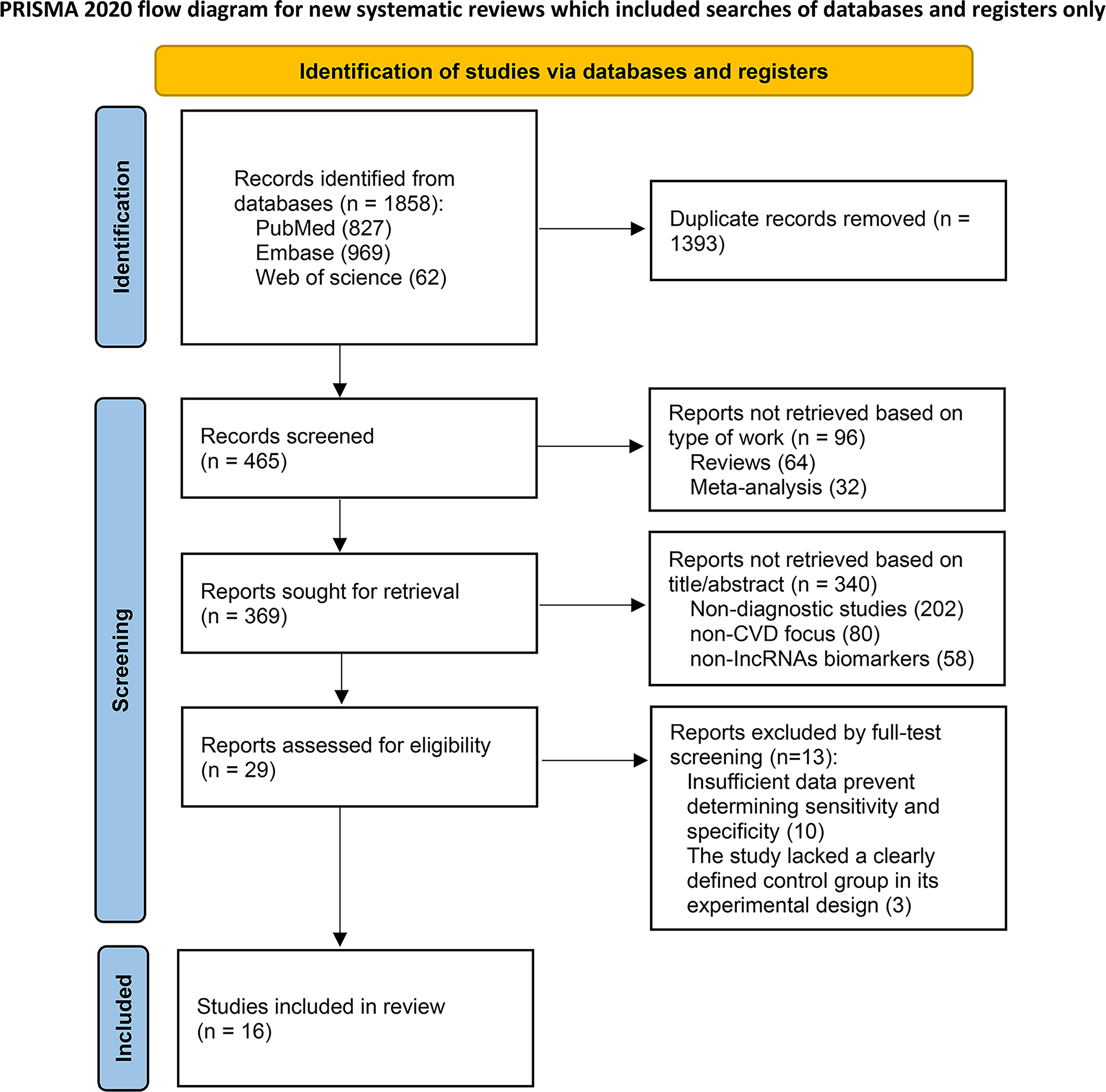
PRISMA 2020 flow diagram illustrating the study selection process from identification to inclusion.
Baseline Characteristics and Long non-Coding RNAs Expression Profiles of Studies Included in the meta-Analysis.

Notes. M, male; F, female; CVDs, cardiovascular diseases; AMI, acute myocardial infarction; UA, unstable angina; CHF, chronic heart failure; ACS, acute coronary syndrome; CAD, Coronary atherosclerotic heart disease; STEMI, ST-segment elevation myocardial infarction.
The control groups in these studies included healthy individuals (10 studies with 11 tests), chest pain (3 studies), and other individuals (3 studies). Subgroup analysis based on control group was performed between healthy individuals and others (chest pain + other individuals). Eight studies with 10 tests had the mean age of the control groups that were more than 60 years old; other studies were less than or equal to 60 years old. Disease in the case group were 6 studies (with 8 tests) for acute myocardial infarction (AMI), four studies for chronic heart failure (CHF), two studies for acute coronary syndrome (ACS), one study for coronary heart disease (CHD), one study for coronary atherosclerotic heart disease (CAD), and two studies for ST-segment elevation myocardial infraction (STEMI). Disease type in the subgroup analysis was conducted in the comparison between AMI and other diseases (CHF, ACS, CHD, CAD, and STEMI). Eleven studies (with 13 tests) had a mean age of patients of more than 60 years old. Based on original literature reports, 14 lncRNAs with the up-regulation trend, and another 4 were down-regulated (Table 1). Sensitivity, specificity, and AUC across studies were presented in Table 2.
Systematic Review of Long non-Coding RNAs Diagnostic Performance: Sensitivity, Specificity, and AUC Across Studies.
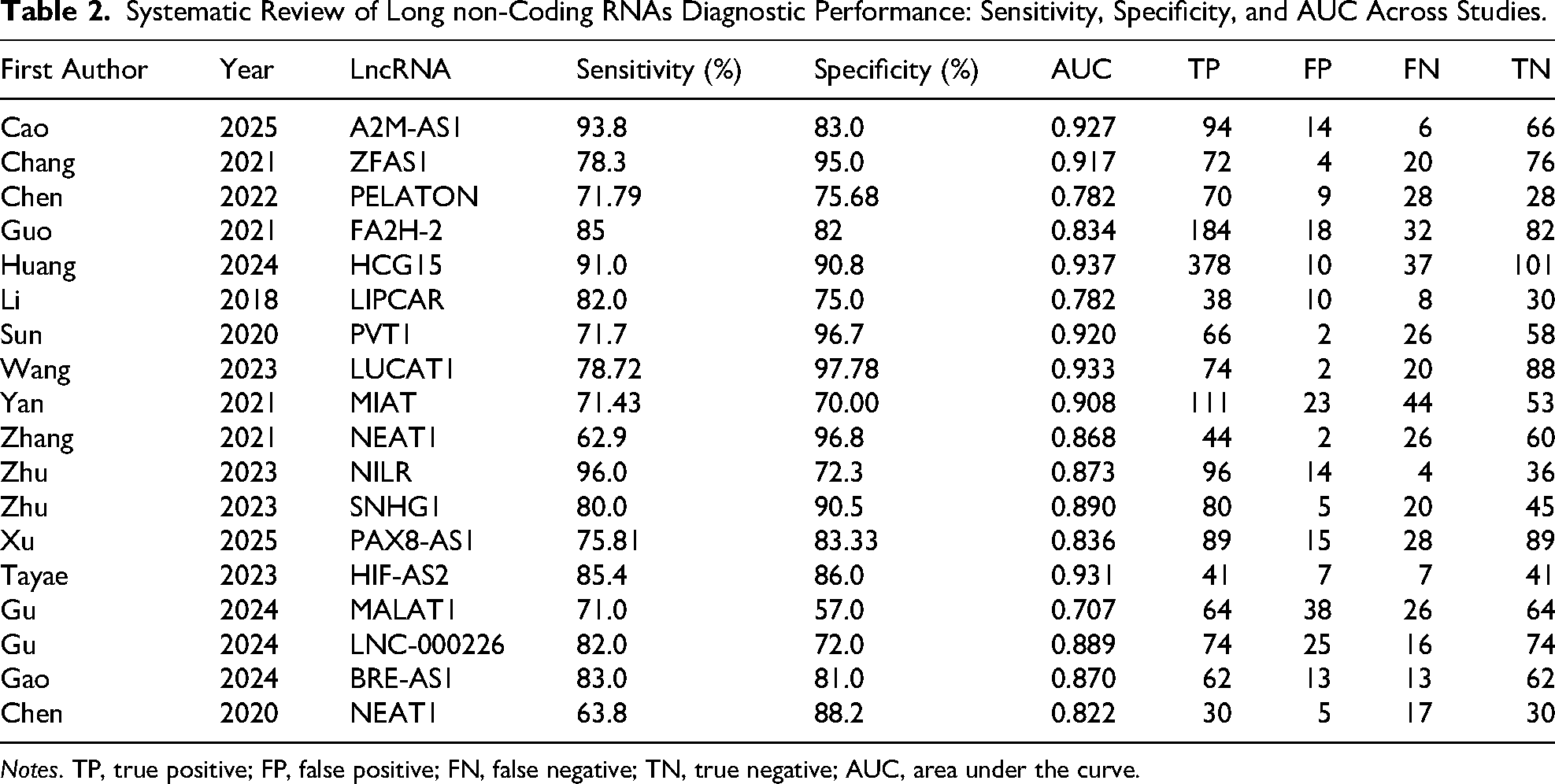
Notes. TP, true positive; FP, false positive; FN, false negative; TN, true negative; AUC, area under the curve.
Quality Assessment
Quality of enrolled studies was analyzed using QUADAS - 2 tool, in risk of bias and applicability concerns two aspects. The results showed that only one study had high risk in the index test bias (Figure 2 A and B). None of the included studies had low quality.

Quality assessment for enrolled studies. A, Risk of bias graph. B, Risk of bias summary.
Diagnostic Value of LncRNAs for CVDs
The summary analysis revealed high heterogeneity in both sensitivity (τ2 = 0.02, I2 = 84.22%) and specificity (τ2 = 0.02, I2 = 83.30%). LncRNAs had pooled sensitivity (0.81, 95% CI = 0.76-0.85) and specificity (0.86, 95% CI = 0.80-0.90) for CVDs diagnosis (Figure 3A). The SROC showed that the AUC was 0.90 (95% CI = 0.87-0.92) for lncRNAs (Figure 3B). Combined with the shape of the SROC curve, it indicated a high diagnostic value of lncRNAs for CVDs. The positive DLR was 5.73 (95% CI =4.07-8.08, Figure 4A), and the negative DLR was 0.22 (95% CI = 0.18-0.28, Figure 4B). It suggested the moderate clinical applicability of the present results. The diagnostic score was 3.25 (95% CI = 2.79-3.71, Figure 4C), and the diagnostic odds ratio was 25.77 (95% CI = 16.22-40.93, Figure 4D). It indicated the high diagnostic accuracy for lncRNAs.
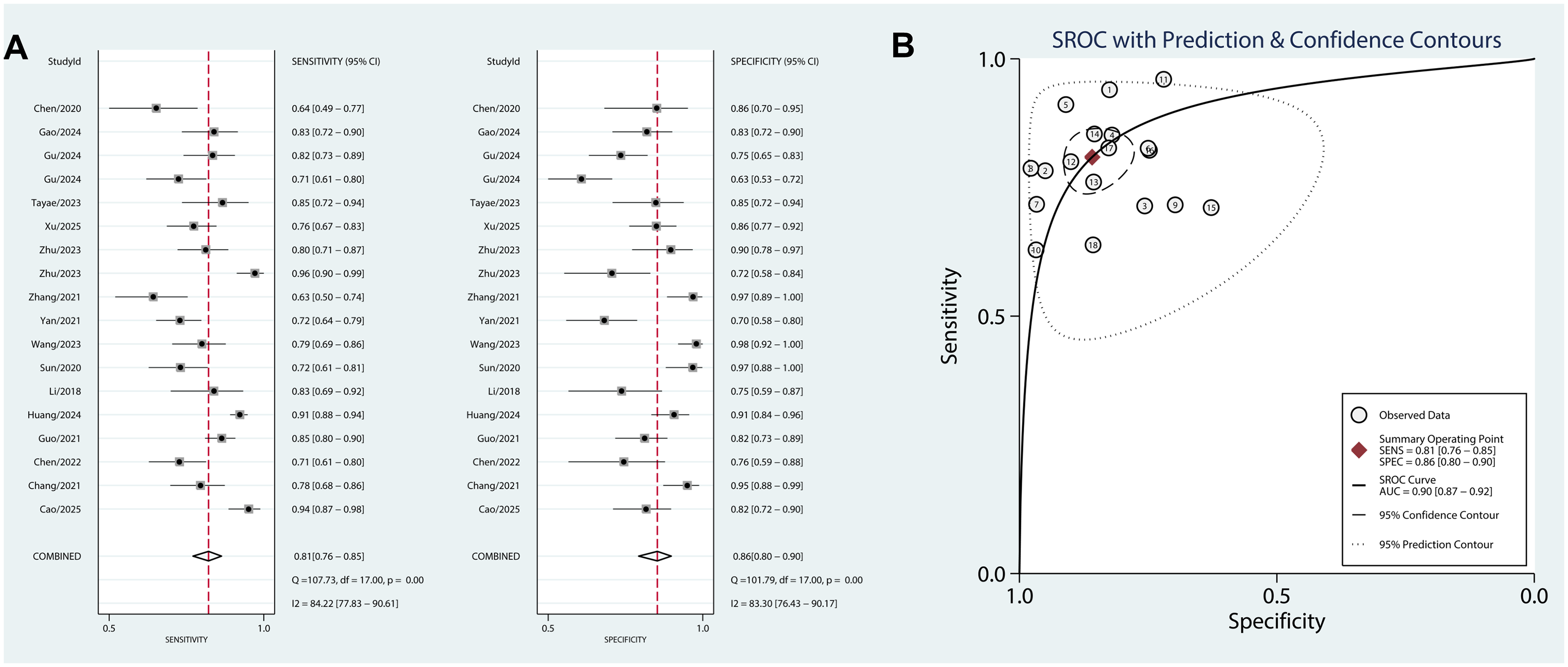
Diagnostic value of lncRNAs for CVDs. A, Pooled analyses of sensitivity (0.81 [0.76-0.85]) and specificity (0.86 [0.80-0.90]). B, SROC of lncRNAs for CVDs diagnosis (AUC = 0.90 [0.87-0.92]).
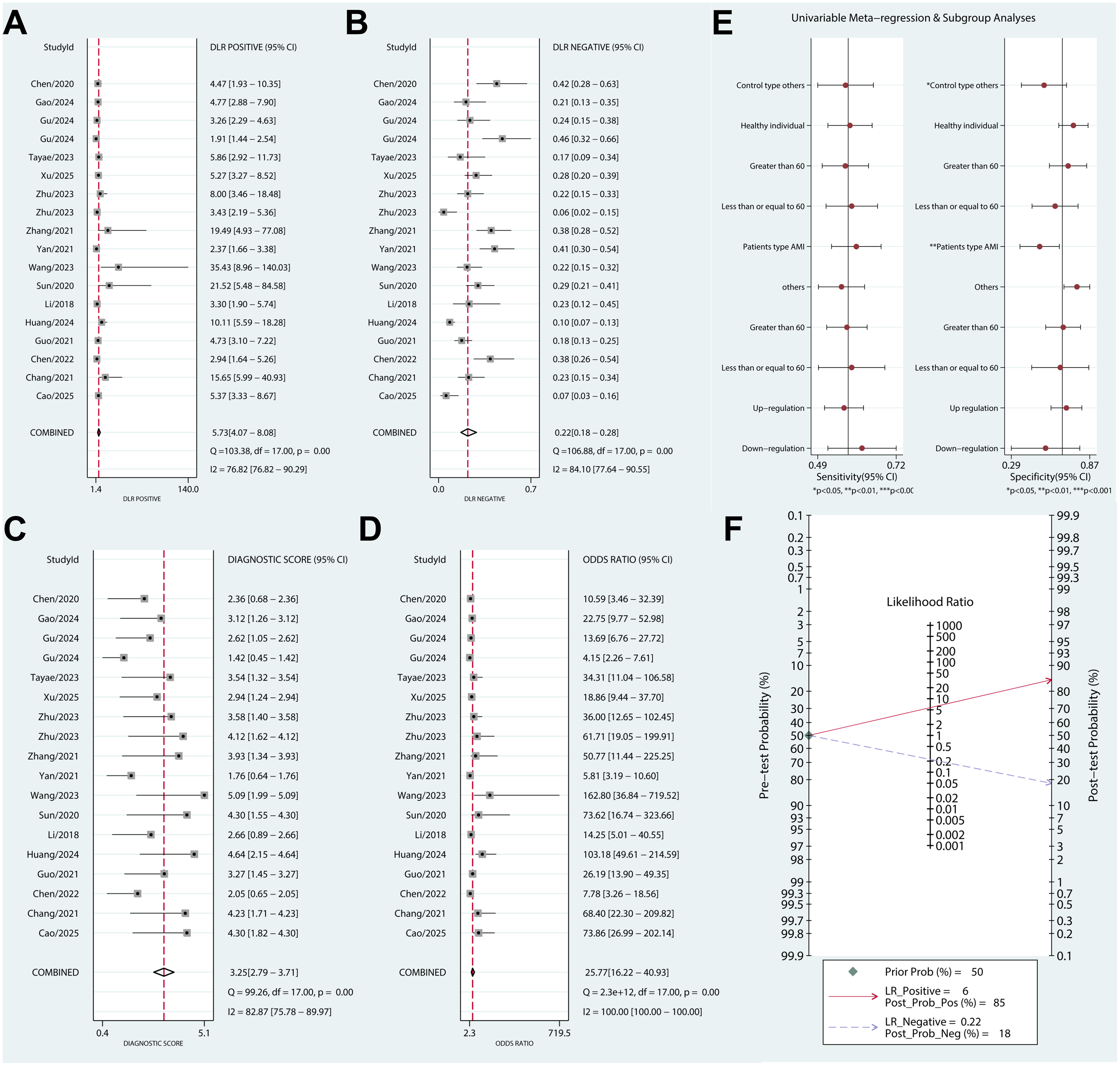
Diagnostic accuracy, clinical applicability, and subgroup analysis. A, Positive DLR (5.73 [4.07-8.08]). B, Negative DLR (0.22 [0.18-0.28]). C, Diagnostic score (3.25 [2.79-3.71]). D, Diagnostic odds ratio (25.77 [16.22-40.93]). E, Subgroup analysis. F, Fagan diagram.
Meta-Regression Analysis
Meta-regression and subgroup analyses were performed to explore the sources of heterogeneity in sensitivity and specificity. These analyses were conducted based on the control type (healthy individual and others), control age (>60 vs ≤ 60), disease type (AMI vs others), age of patients (>60 vs ≤ 60), and expression trend of lncRNAs (up-regulation vs down-regulation) (Figure 4E). Then, we found that the control and patient subject types were the sources of heterogeneity in specificity. Only the patient subject type contributes to the overall heterogeneity (τ2 = 0.03, P = 0.02, I2 = 75%). Subgroup forest plots for disease subtype have been added as Supplementary Figure 1.
Subgroup analysis demonstrated that lncRNAs had high specificity when the control group consisted of healthy individuals and the disease type was classified as non-AMI subgroups (Table 3).
Sample-Stratified Long non-Coding RNAs Diagnostic Performance in Cardiovascular Disease.

Clinical Applicability
The pre-test probability was set to 50%. Present pooled results increased the positive DLR to 85% and decreased the negative DLR to 18% (Figure 4F). These findings strongly support the use of lncRNAs for diagnosing CVDs and excluding non-CVDs.
Publication Bias and Sensitivity Analysis
Deeks’ funnel plot asymmetry test showed no significant publication bias (P = 0.38) in enrolled studies (Figure 5A). Most studies were located on a straight line at a 45-degree angle in the goodness of fit diagram (Figure 5B). It suggested the enrolled studies had high goodness of fit and met the normality assumption. Included studies were distributed near the 45-degree angle in the Bivariate normality (Figure 5C), suggesting the high goodness of fit for bivariate normality. Only one study exceeded the cook's distance (Figure 5D), indicating its significant influence on pooled results. The standardized residual diagram showed only one suspected outlier data point (the point has crossed the red line threshold of the diseased standardized residual, Figure 5E). Further examination of the validity and rationality of study 11 is required. These results suggested the robustness of the current results.

Publication bias and sensitivity analysis. A, Deeks’ funnel plot asymmetry test shows no significant asymmetry (Deeks’ test P = 0.38), suggesting minimal publication bias. B, Goodness of fit. C, bivariate normality. D, Influence analysis. E, Outlier detection.
Discussion
This study assess the diagnostic potential of lncRNAs in CVDs through a systematic review and meta-analysis. It aims to provide valuable insights for clinical practice and guide future research. Notably, this work is the first to comprehensively compile and evaluate evidence on lncRNAs as diagnostic indicators for CVDs, filling a critical gap in the existing literature.
To better understand the clinical relevance of lncRNAs as diagnostic biomarkers, the diagnostic performance metrics were carefully examined. Present results revealed that lncRNAs show high sensitivity and specificity for CVDs diagnosis. Positive DLR was 5.73, more than 5 indicated that a positive test result increases the post-test probability of CVDs by approximately 5.73-fold compared to the pre-test probability. While conventional thresholds classify positive DLR values more than 10 as “strong evidence” for disease confirmation, values between 5-10 remain clinically meaningful, particularly in high-prevalence settings. The negative DLR was 0.22, aligned with the “moderate exclusion” category (negative DLR from 0.2 to 0.3), though the failure to achieve a stricter threshold (negative DLR <0.1) suggests residual risk of missed diagnoses, necessitating caution in low-prevalence populations. 33 It was suggested to explore combinatorial algorithms integrating DLRs with clinical scores or novel biomarkers to enhance predictive accuracy. The DOR of 25.77 exceeded the conventional threshold of 10 for “strong diagnostic utility”, 34 indicating that lncRNAs exhibit exceptional accuracy in distinguishing CVD patients from controls. The high DOR supports lncRNAs as adjunctive tools to confirm CVD in ambiguous cases, complementing, but not replacing, established tools. Circulating lncRNAs may identify subclinical CVD stages, enabling preemptive interventions, such as atherosclerosis. 35 While established biomarkers like hs-cTn and BNP are cornerstone tools, they have inherent limitations, including delayed release kinetics, reduced specificity in comorbidities, and limited sensitivity for early-stage disease.9,10 LncRNAs address these gaps through earlier detection within 2 h of injury. Besides, a multiplex lncRNA panel can simultaneously capture inflammation, apoptosis, and fibrosis, distinguishing CVDs from other diseases. The cost-effectiveness of lncRNAs is substantially cheaper than cardiac MRI. By providing an earlier signal of cardiac injury, lncRNA could narrow the diagnostic window in acute care. Furthermore, specific lncRNA signatures may improve differential diagnosis and refine risk stratification beyond conventional markers. A multi-marker approach integrating lncRNAs with cTn or BNP could enhance overall diagnostic precision. However, clinical translation depends on developing standardized assays and validating these biomarkers in large, prospective cohorts. This may be because the expression changes of lncRNAs are sensitive to the occurrence and development of CVDs and are closely related to the pathological process of the disease. 36
To explore heterogeneity in sensitivity and specificity, meta-regression and subgroup analyses were conducted. The expression trend of lncRNAs, control type, and disease type were identified as the main sources of heterogeneity. The expression levels of lncRNAs can be influenced by various factors like disease progression, tissue origin, and individual genetic differences. These variations in expression may directly impact the diagnostic accuracy of lncRNAs. Changes in their expression levels can affect their reliability as biomarkers. When using healthy individuals as controls versus patients with other cardiovascular conditions, differences in specificity can arise. The heterogeneity in disease types also matters. Each cardiovascular disease has unique pathophysiological mechanisms that lead to differential lncRNA expression and variations in diagnostic performance. Controlling for these factors in study design could improve the comparability and reliability of results.
The 16-study analysis included 1855 cases and 1125 controls. However, significant heterogeneity persisted, primarily in lncRNA expression trends and control group types, which may affect result stability. Another significant limitation is geographic bias. Most eligible studies originated from China. This pronounced geographic bias potentially compromises the external validity of our conclusions, particularly their applicability to non-Asian populations and disparate healthcare systems. Therefore, prospective, multicenter validation studies involving diverse geographic and ethnic populations are warranted to certify these findings. While the diagnostic value of lncRNAs for CVDs was preliminarily confirmed, their specific biological mechanisms in disease development remain largely unexplored, limiting a deeper understanding of their diagnostic principles. Additionally, the feasibility and cost - cost-effectiveness of lncRNAs’ clinical application require further clinical validation.
Given these considerations, future research should focus on a larger sample size, multi-center studies to verify lncRNAs’ diagnostic performance, and enhancing the extrapolation and clinical value of results. The role of lncRNAs in CVDs, such as gene expression regulation and cell function impact, warrants in-depth exploration to underpin lncRNA-based therapies. Third, investigating the diagnostic value of combining lncRNAs with traditional or novel biomarkers could improve both diagnostic accuracy and applicability. Finally, prospective cohort studies are essential to assess how lncRNAs change across different CVD stages and their prognostic value, offering a more complete picture of their clinical potential.
Conclusion
This study demonstrates that lncRNAs have high sensitivity and specificity for diagnosing CVDs, indicating their potential as auxiliary diagnostic tools. Meanwhile, the heterogeneity of results indicated the need for caution in practical applications. Future research should focus on increasing sample sizes, delving into biological mechanisms, merging biomarkers, and carrying out prospective studies. These approaches are expected to enhance the diagnostic application of lncRNAs in CVDs.
Supplemental Material
sj-docx-1-cat-10.1177_10760296251410926 - Supplemental material for LncRNAs as Emerging Diagnostic Biomarkers for Cardiovascular Diseases: Evidence Synthesis Through Systematic Review and Meta-Analysis
Supplemental material, sj-docx-1-cat-10.1177_10760296251410926 for LncRNAs as Emerging Diagnostic Biomarkers for Cardiovascular Diseases: Evidence Synthesis Through Systematic Review and Meta-Analysis by Baocheng Zhu and Yong Liang in Clinical and Applied Thrombosis/Hemostasis
Supplemental Material
sj-docx-2-cat-10.1177_10760296251410926 - Supplemental material for LncRNAs as Emerging Diagnostic Biomarkers for Cardiovascular Diseases: Evidence Synthesis Through Systematic Review and Meta-Analysis
Supplemental material, sj-docx-2-cat-10.1177_10760296251410926 for LncRNAs as Emerging Diagnostic Biomarkers for Cardiovascular Diseases: Evidence Synthesis Through Systematic Review and Meta-Analysis by Baocheng Zhu and Yong Liang in Clinical and Applied Thrombosis/Hemostasis
Footnotes
Competing Interests
The authors declare no competing interests.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
