Abstract
Background
MicroRNAs (miRNAs) play a crucial role in the pathophysiology of sepsis, with aberrant expression documented in peripheral blood samples from sepsis patients. This study aims to investigate the causal relationship between circulating miRNAs and sepsis using Mendelian Randomization (MR) approaches.
Methods
We utilized miRNA expression quantitative trait loci (eQTL) data and genome-wide association study (GWAS) data on sepsis. MR analysis was conducted using Inverse Variance Weighted (IVW) and MR-Egger regression methods to identify miRNAs significantly associated with sepsis risk. Potential targets and related pathways of these miRNAs were also analyzed.
Results
IVW and MR-Egger regression analyses revealed that three circulating miRNAs (hsa-miR-125a-5p, hsa-miR-494-3p, and hsa-miR-885-5p) exhibited significant associations with reduced sepsis risk, indicating protective effects. Conversely, hsa-miR-196b-5p, hsa-miR-27b-3p, and hsa-miR-598-3p were linked to increased sepsis risk. The association between hsa-miR-27b-3p and increased sepsis risk was further validated in an independent cohort. Target identification and enrichment analysis indicated that the targets of these miRNAs are closely related to biological processes such as bacterial infection, apoptosis, and the FoxO signaling pathway.
Conclusion
Our study provides novel insights into the causal relationships between circulating miRNAs and sepsis through MR analysis, offering new perspectives for early prevention and treatment strategies for sepsis. These findings provide a foundation for further investigation into the potential application of miRNAs in the diagnosis and treatment of sepsis.
Introduction
Sepsis is a life-threatening organ dysfunction syndrome triggered by infection, characterized by a dysregulated host response leading to systemic inflammation. In China alone, there are over 9 million cases of severe sepsis annually, with an annual growth rate ranging from 1.5% to 8.0%, resulting in more than 800 000 deaths. 1 Despite significant advances in critical care and antibiotic therapy, sepsis remains one of the major public health concerns globally, associated with high morbidity and mortality rates. According to the World Health Organization, sepsis affects approximately 55 million people worldwide each year, causing 11 million deaths, imposing a substantial burden on global healthcare systems, particularly impacting children and low-income countries.2,3 The challenge of sepsis prevention and control is further exacerbated by the aging population and an increasing number of immunocompromised patients. 4 Therefore, investigating the pathogenesis of sepsis, exploring early diagnostic biomarkers, and developing novel therapeutic approaches are critically important.
MicroRNAs (miRNAs), endogenous non-coding RNA molecules approximately 21 nucleotides in length, have been recognized as key regulators of gene expression since their discovery in Caenorhabditis elegans in 1993. By binding to the 3′-untranslated regions of mRNAs, miRNAs can either degrade or inhibit protein translation. Recently, the pivotal roles of miRNAs in the pathophysiology of sepsis have garnered considerable attention. Studies have identified abnormal expressions of various miRNAs in the peripheral blood of sepsis patients, indicating their potential as biomarkers and their involvement in sepsis-related signaling pathways and cellular processes. 5 For instance, a composite marker comprising miR-150 and miR-4772-5p-iso can distinguish sepsis from systemic inflammatory response syndrome (SIRS) with high specificity and sensitivity. 6 Additionally, changes in specific miRNAs such as miR-146a 7 and miR-223 8 levels correlate with different stages of sepsis, suggesting their significance in predicting disease progression and patient outcomes. Understanding the roles of miRNAs in sepsis provides novel and viable research directions for developing new diagnostic, prognostic, and therapeutic strategies.
Recent studies have demonstrated causal relationships between plasma circulating miRNAs and various diseases, including COVID-19, 9 amyotrophic lateral sclerosis, 10 and Parkinson's Disease. 11 However, no studies have explored the relationship between plasma circulating miRNAs and sepsis. This study aims to investigate the causal relationship between miRNAs and sepsis using Mendelian Randomization (MR) methods. We will utilize existing miRNA expression quantitative trait loci (eQTL) data and genome-wide association study (GWAS) data on sepsis to identify miRNAs significantly associated with sepsis risk and analyze their potential targets and related pathways. Through this research, we aim to provide new insights and evidence for the early prevention and treatment of sepsis.
Materials and Methods
Acquisition of miRNA eQTL Data
In this study, miRNA eQTLs data were obtained from two distinct cohorts: 1) The dataset from Huan et al, 12 which included eQTL data for 280 high-quality miRNAs across 5329 individuals, was used as the training set. To minimize potential horizontal pleiotropy, we focused on cis-miRNA-eQTLs and excluded SNPs located within coding regions with synonymous or non-synonymous mutation consequences. Instrumental variables (IVs) were selected using a false discovery rate (FDR) adjusted by the Benjamini-Hochberg method, set at 0.1 (corresponding to an original P-value of 6.6 × 10−5). 2) The second cohort provided by Nikpay et al 13 included eQTL data for 2083 miRNAs in 710 individuals, serving as the validation set, with a P-value threshold of 1 × 10−5 for selecting relevant IVs. To harmonize miRNA identifiers across both datasets, we utilized the miRCarta v1.1 database. This approach not only effectively identified disease-related miRNAs but also ensured the accuracy and reliability of the results.
Sepsis GWAS Data Retrieval
Summary statistics from the sepsis GWAS cohort were obtained from the MRC-IEU OpenGWAS database (dataset ID: ieu-b-5088), based on resources from the UK Biobank. This dataset comprised 11 568 cases and 451 301 controls, totaling 462 869 European individuals. Additionally, the GWAS covered 12 243 540 SNPs.
Mendelian Randomization Analysis
TwoSampleMR v0.6.8 R package was employed for conducting two-sample Mendelian Randomization (MR) analysis 14 to investigate the association between miRNA expression levels and sepsis risk, following a previously described methodology.9–11 Initially, linkage disequilibrium analysis of IVs was performed within the training set using a clustering window of r² < 0.5 and 10 kb to select independent IVs. Corresponding SNPs were then extracted from the GWAS dataset, and normalization was applied to exposure and outcome data to ensure consistent effect allele usage. Palindromic SNPs were excluded. Two primary analytical strategies were adopted: Inverse Variance Weighted (IVW) and MR-Egger regression analyses. Criteria for identifying causally associated miRNAs included: (1) P-values < .05 in IVW tests and FDR < 0.1; (2) consistent direction of estimated effects in both IVW and MR-Egger models; (3) MR analysis based on at least three SNPs; and (4) significant results in validation studies to confirm high-confidence relationships. Horizontal pleiotropy was assessed using MR-Egger regression, and Leave-One-Out analysis was conducted to examine the influence of individual SNPs on overall causal inference. Heterogeneity among the causal estimates for each miRNA was assessed using Cochran's Q statistic. A P-value > .05 for Cochran's Q was considered indicative of a lack of significant heterogeneity, suggesting that the instrumental variables for a given miRNA provide consistent causal estimates. This methodology enhances the reliability of the findings and provides new perspectives on the role of miRNAs in disease.
Target Identification and Enrichment Analysis
To explore the functional impact of identified potential causal miRNAs, we utilized miRNet 2.0 and its online platform to identify their target genes. This tool is accessible via https://www.mirnet.ca/miRNet/home.xhtml (accessed on January 12, 2025). Only experimentally validated target genes sourced from the miRTarBase v9.0 database were considered. These miRNAs and their corresponding target genes were imported into Cytoscape software v3.10.2 to construct detailed miRNA-gene interaction network maps. For further analysis of biological processes involving these target genes, ClusterProfiler package was used for Gene Ontology (GO) annotation enrichment analysis, focusing particularly on Biological Process (BP) enrichment. An FDR threshold of less than 0.05 was set to adjust for multiple comparisons.
Results
Circulating miRNA eQTLs
From the cohort provided by Huan et al, we initially selected 9612 single nucleotide polymorphisms (SNPs) as instrumental variables. After filtering, 9517 SNPs not directly associated with coding gene regions were identified for further analysis. Additionally, 17 SNPs linked to mir-213 were excluded due to ineffective mapping to known miRNA identifiers. Ultimately, this study focused on 9500 SNPs significantly associated with 75 circulating mature human miRNAs for in-depth investigation (Table 1, Supplemental Materials: Table S1). Figure 1 illustrates the workflow diagram of this MR analysis.
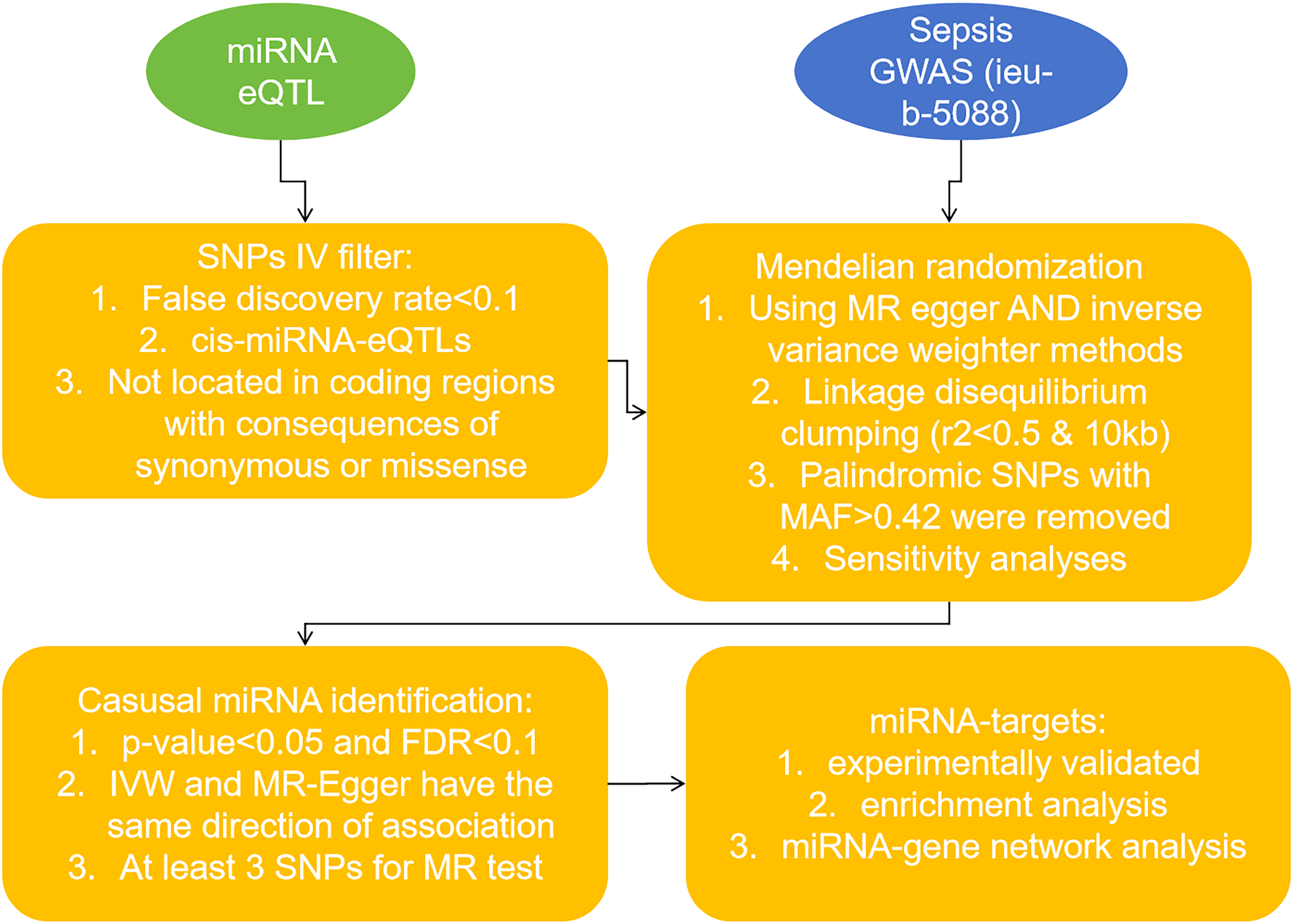
The Workflow Diagram of this MR Analysis.
The List of Top SNPs Per miRNA in the Huan Et al Cohort.

Sepsis-Associated Circulating miRNAs
Using IVW and MR-Egger regression, we identified all miRNAs significantly associated with sepsis risk (Figure 2, Supplemental Materials: Figure S1), without observing significant horizontal pleiotropy (Table 2). After Bonferroni correction for multiple testing, three blood miRNA markers were significantly associated with reduced IDD risk: hsa-miR-125a-5p (IVW, OR: 0.971, 95% CI: 0.952-0.991, P-value: .004), hsa-miR-494-3p (IVW, OR: 0.976, 95% CI: 0.963-0.989, P-value: .001), and hsa-miR-885-5p (IVW, OR: 0.954, 95% CI: 0.924-0.984, P-value: .003). Conversely, three blood miRNA markers were significantly associated with increased sepsis risk: hsa-miR-196b-5p (IVW, OR: 1.069, 95% CI: 1.028-1.111, P-value: .001), hsa-miR-27b-3p (IVW, OR: 1.097, 95% CI: 1.035-1.162, P-value: .002), and hsa-miR-598-3p (IVW, OR: 1.066, 95% CI: 1.031-1.103, P-value < .001). Leave-One-Out results for miRNAs are shown in Figures S2-S7, indicating good stability and reliability of the findings. Cochran's Q test indicated no significant heterogeneity for the instrumental variables of these miRNAs (all P > .05, Table S2), supporting the consistency of the effect estimates.

Associations of Serum-Circulating miRNAs with Protective and Harmful Effects on Sepsis in the Discovery Dataset. nSNP: Number of Single Nucleotide Polymorphism; CI: Confidence Interval; MR: Mendelian Randomization.
Pleiotropy Tests of Instrument Effects in the Discovery Dataset.

Mechanisms of miRNA Regulation in Sepsis Risk
To further explore the potential mechanisms by which these causal miRNAs might influence sepsis, we predicted 1047 experimentally validated targets using miRNet. Among these, hsa-miR-27b-3p had the most targets (439), followed by hsa-miR-125a-5p (280 targets), hsa-miR-196b-5p (162 targets), hsa-miR-494-3p (142 targets), hsa-miR-885-5p (73 targets), and hsa-miR-598-3p (26 targets) (Figure 3). KEGG pathway enrichment analysis revealed that these targets are involved in pathways related to bacterial infections, proteoglycans in cancer, cellular senescence, FoxO signaling, relaxin signaling, and various cancer pathways (Figure 4). GO annotation enrichment indicated significant enrichment in biological processes such as myeloid cell differentiation, response to oxygen levels, cell growth, and apoptosis (Figure 5). These analyses provide plausible biological hypotheses for future functional studies to validate.
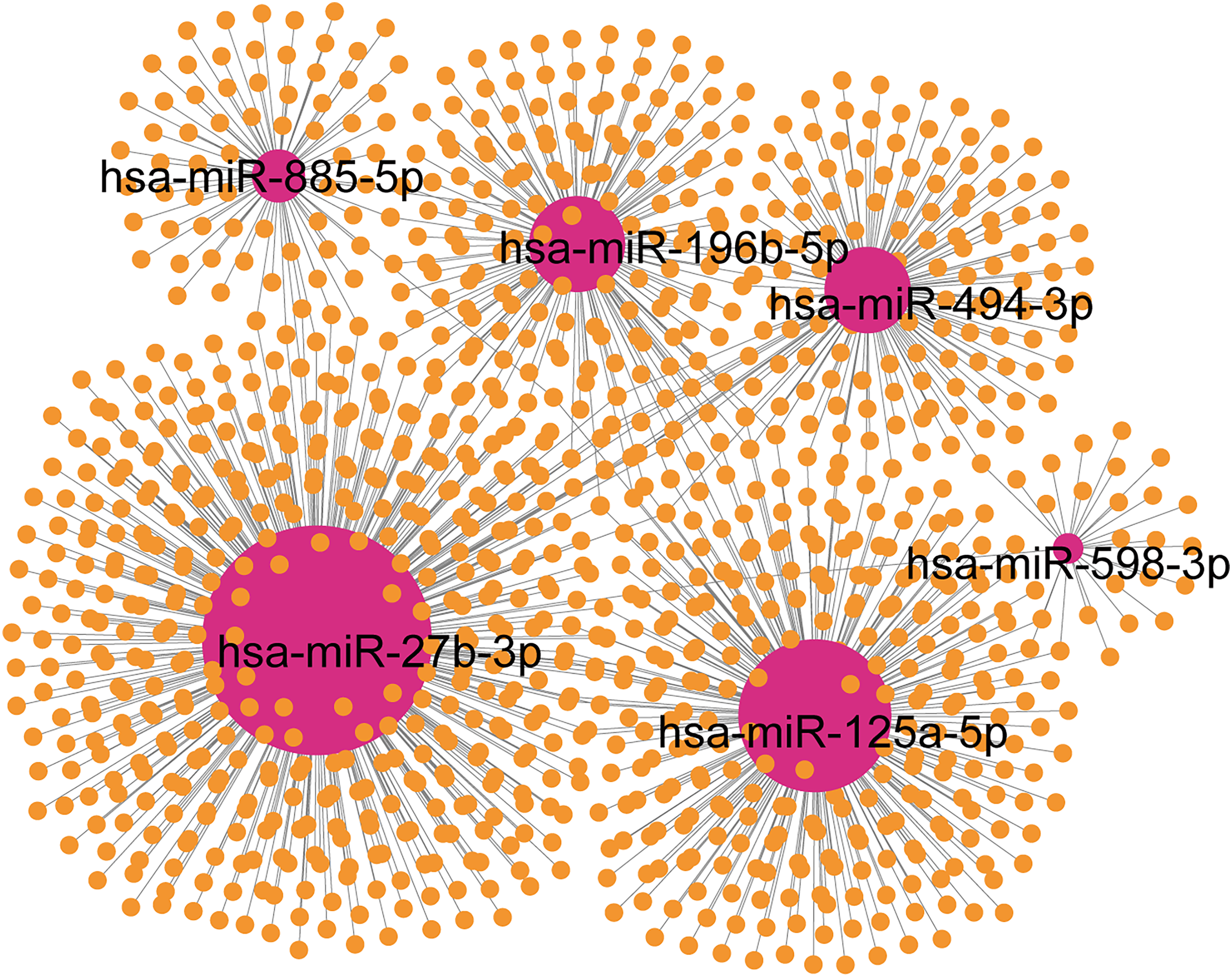
The miRNA-Gene Network of the Causal miRNAs Associated with Sepsis.

Significantly Enriched KEGG Pathways for the Targets of Causal miRNAs.

Significantly Enriched Biological Processes for the Targets of Causal miRNAs.
Validation of Circulating miRNAs in an Independent Cohort
Next, we used the cohort from Nikpay et al to validate potential miRNAs. We collected genome-wide miRNA eQTL data for all six candidate causal miRNAs. A P-value threshold of 1 × 10−6 was set to select relevant IVs. The same quality control procedures applied in the original study were also performed here. After data preprocessing, all six miRNAs were validated in this new study, without observing significant horizontal pleiotropy (Figure 6, Table S3). IVW analysis showed that hsa-miR-27b-3p exhibited a detrimental effect on sepsis, with an OR of 1.22 (95% CI: 1.061-1.403), consistent with observations from the discovery dataset. Other miRNAs did not show significant associations with sepsis risk in this cohort.
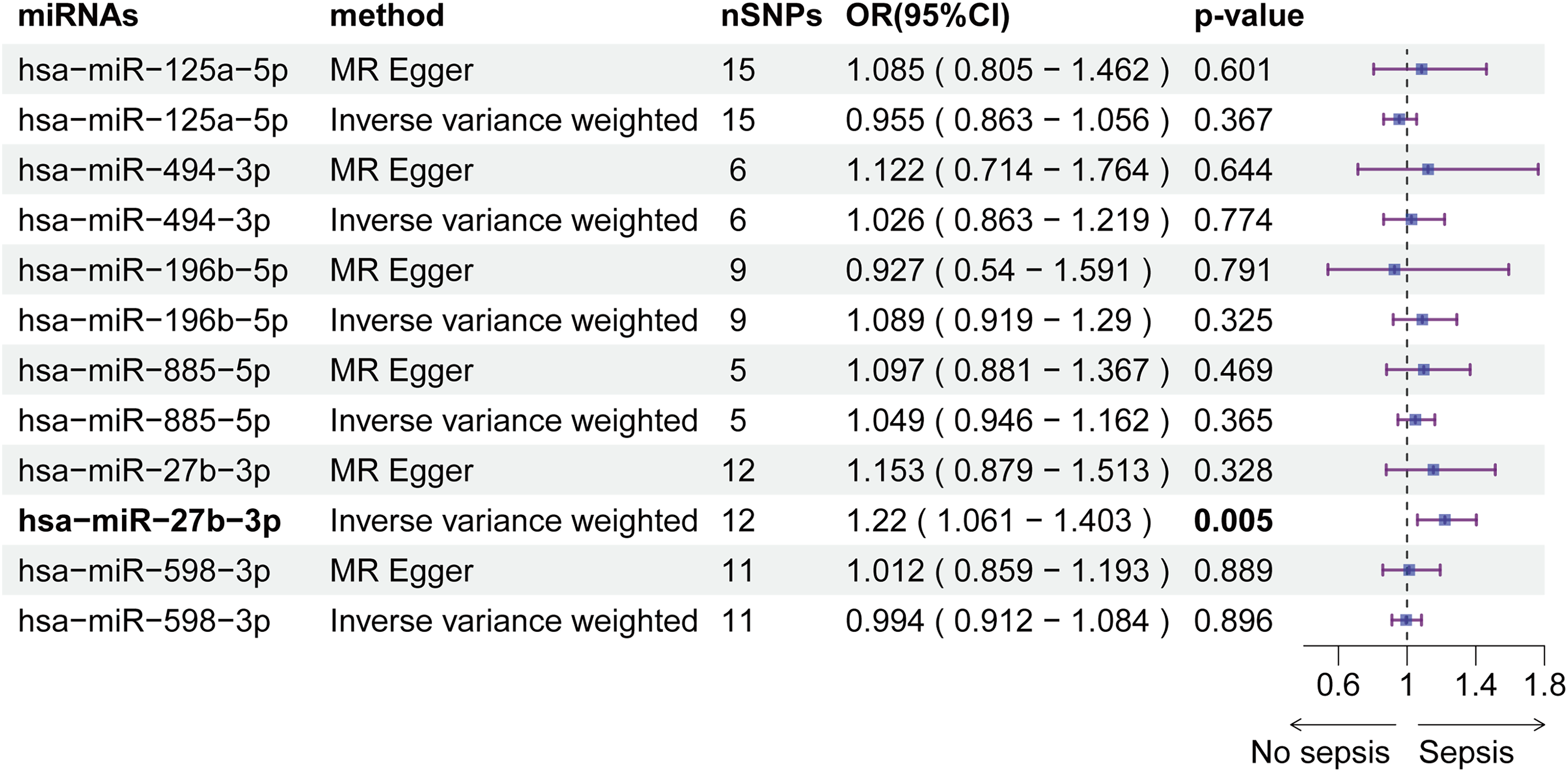
Associations of serum-Circulating miRNAs with Protective and Harmful Effects on Sepsis in the Validation Dataset. nSNP: Number of Single Nucleotide Polymorphism; CI: Confidence Interval; MR: Mendelian Randomization.
Discussion
This study employed MR to systematically investigate, for the first time, the causal relationship between circulating miRNAs and sepsis, providing new insights for early diagnosis and therapeutic strategies. Sepsis is a severe systemic inflammatory response syndrome with a complex pathogenesis involving alterations in various biomarkers. We identified a panel of circulating miRNAs that may exert causal effects on sepsis risk. Although their precise mechanisms require further experimental validation, these findings offer potential directions for improving sepsis management.
The most immediate application lies in the development of biomarkers. A multi-miRNA signature, constructed based on the protective miRNAs (eg, hsa-miR-125a-5p, hsa-miR-494-3p) and risk miRNAs (eg, hsa-miR-27b-3p) identified here, could be utilized for early diagnosis and risk stratification in patients with suspected infections. 15 This would assist clinicians in identifying high-risk individuals and initiating timely, targeted interventions. Furthermore, protective miRNAs represent promising candidates for novel therapeutic strategies, such as miRNA mimics. 16 Conversely, risk miRNAs may indicate the need for closer monitoring or interventions targeting their downstream pathways (eg, apoptosis regulation).
We found that hsa-miR-125a-5p, hsa-miR-494-3p, and hsa-miR-885-5p significantly reduce the risk of sepsis. Studies have shown that in sepsis patients, serum extracellular vesicles contain elevated levels of hsa-miR-125a-5p, which are transported to lung macrophages to inhibit Tnfaip3, thereby promoting reactive oxygen species (ROS) production and pro-inflammatory cytokine release, leading to acute lung injury. 17 Additionally, sepsis is a major cause of acute kidney injury, and hsa-miR-125a-5p may mitigate renal inflammation through the TRAF6/NF-κB axis. 18 Compared to healthy controls, serum levels of hsa-miR-494-3p are significantly lower in sepsis patients; this miRNA can regulate inflammation by targeting PTEN mRNA's 3′-untranslated region 19 and modulate TLR6 expression to alleviate sepsis-related inflammation. 20 Moreover, miR-494-3p targets Lp-PLA2 and inhibits its secretion, thereby reducing sepsis-associated inflammation. 21 MiR-885-5p has been shown to regulate cell growth, senescence, and/or apoptosis. Research indicates that exosomes from sepsis patients promote cardiomyocyte apoptosis via HMBOX1-mediated miR-885-5p, suggesting new therapeutic directions for sepsis-induced cardiac suppression. 22
Furthermore, we found that hsa-miR-196b-5p, hsa-miR-27b-3p, and hsa-miR-598-3p are associated with increased sepsis risk. Chrysophanol has been shown to inhibit inflammation and apoptosis through the miR-27b-3p-PPARG axis, offering protective effects against sepsis-induced acute myocardial injury. 23 In sepsis patients, miR-27b-3p is upregulated in both cellular and extracellular vesicle compartments and exhibits different expression patterns at various disease stages, suggesting its potential role in diagnosing and assessing sepsis severity. 24 However, there is no direct experimental evidence linking hsa-miR-196b-5p and hsa-miR-598-3p to sepsis, although they are widely implicated in regulating inflammation and apoptosis.25,26 Future research should further explore the roles and mechanisms of these miRNAs in sepsis.
Our MR analysis identifies a causal link, but the underlying mechanisms are derived from computational predictions and thus are speculative. For instance, while bacterial infection is a primary cause of sepsis and the targets of causal miRNAs were significantly enriched in related pathways, this association is suggestive rather than proven, and experimental confirmation is necessary. 27 Similarly, the proposed associations with myeloid cell differentiation, apoptosis, and the FoxO signaling pathway represent interesting hypotheses generated from our analysis that merit further investigation in mechanistic studies. 28 Multiple apoptosis regulatory mechanisms are affected in sepsis patients. For instance, the expression of pro-apoptotic proteins like Bax increases, while anti-apoptotic proteins such as Bcl-2 decrease, promoting apoptosis. 29 Targeting apoptosis regulation pathways could be a potential therapeutic strategy. FoxO transcription factors can regulate inflammation, 30 apoptosis and survival, 31 and immune cell function, including T lymphocytes and macrophages. 32 Furthermore, sepsis often involves severe metabolic disturbances, and FoxO transcription factors also participate in cellular metabolism regulation, particularly adjusting energy metabolism under nutrient deficiency or stress conditions to maintain cell survival. 33
Moreover, the independent validation cohort results showed that hsa-miR-27b-3p exhibits detrimental effects on sepsis, consistent with findings from the discovery dataset, enhancing the reliability of our conclusions. Notably, however, other candidate miRNAs failed to exhibit statistically significant associations. This discrepancy may stem from multiple contributing factors: First, the validation cohort's relatively modest sample size (n = 710) compared to the discovery set (n = 5329) may have compromised the statistical power to detect miRNA associations with moderate effect sizes. 34 Second, inherent biological variability in miRNA expression profiles across diverse populations, compounded by technical discrepancies in eQTL profiling platforms, could undermine reproducibility. 35 Third, certain miRNA effects might be context-dependent, influenced by specific genetic architectures or environmental exposures that differ between cohorts. 36
Several limitations of this study should be acknowledged. First, the GWAS and eQTL data utilized were primarily derived from European populations, which may limit the generalizability of our findings to other ethnic groups and necessitate further validation in diverse cohorts. Second, although the MR analysis suggests causal relationships and pathway analyses reveal potential biological implications, these findings remain hypothesis-generating and are not yet directly translatable to clinical practice. Future work should include functional validation in experimental models of sepsis, as well as well-designed prospective cohort studies to evaluate the diagnostic and prognostic utility of these miRNAs before clinical application can be considered.
It is also noteworthy that the effect sizes (odds ratios) of the identified miRNAs, while statistically significant, were modest—a common feature in MR studies of complex traits. This indicates that individual miRNAs likely contribute only limited effects to sepsis risk and are unlikely to serve as stand-alone diagnostic biomarkers. The principal value of these findings lies in highlighting potential causal players in sepsis pathophysiology, supported by the significant enrichment of their target genes in key biological pathways. Future research could explore the construction of a multi-miRNA signature or a polygenic risk score integrating these miRNAs. Combining such a profile with established clinical scores (eg, SOFA) and biomarkers (eg, PCT) may enhance the accuracy of risk stratification, early diagnosis, and prognosis assessment in sepsis.
Conclusion
In summary, this MR study provides genetic evidence supporting a causal role for specific circulating miRNAs in sepsis pathogenesis. Our findings highlight several promising candidates for therapeutic development. Specifically, miRNA mimics for the protective miRNAs (eg, hsa-miR-125a-5p, hsa-miR-494-3p) or antagomirs for the risk miRNAs (eg, hsa-miR-27b-3p) represent novel biological strategies worthy of investigation. Furthermore, the enriched pathways (eg, apoptosis, FoxO signaling) unveil potential downstream druggable targets. While these possibilities are compelling, they remain hypothetical and require rigorous functional validation in experimental models of sepsis to confirm the mechanisms and assess the efficacy and safety of such interventions before any clinical application can be considered.
Supplemental Material
sj-xlsx-1-cat-10.1177_10760296251386028 - Supplemental material for Exploring the Causal Relationship Between Circulating miRNAs and Sepsis Through Mendelian Randomization Analysis
Supplemental material, sj-xlsx-1-cat-10.1177_10760296251386028 for Exploring the Causal Relationship Between Circulating miRNAs and Sepsis Through Mendelian Randomization Analysis by Yan Jin and Yuan Zhang in Clinical and Applied Thrombosis/Hemostasis
Supplemental Material
sj-xlsx-2-cat-10.1177_10760296251386028 - Supplemental material for Exploring the Causal Relationship Between Circulating miRNAs and Sepsis Through Mendelian Randomization Analysis
Supplemental material, sj-xlsx-2-cat-10.1177_10760296251386028 for Exploring the Causal Relationship Between Circulating miRNAs and Sepsis Through Mendelian Randomization Analysis by Yan Jin and Yuan Zhang in Clinical and Applied Thrombosis/Hemostasis
Supplemental Material
sj-xlsx-3-cat-10.1177_10760296251386028 - Supplemental material for Exploring the Causal Relationship Between Circulating miRNAs and Sepsis Through Mendelian Randomization Analysis
Supplemental material, sj-xlsx-3-cat-10.1177_10760296251386028 for Exploring the Causal Relationship Between Circulating miRNAs and Sepsis Through Mendelian Randomization Analysis by Yan Jin and Yuan Zhang in Clinical and Applied Thrombosis/Hemostasis
Supplemental Material
sj-doc-4-cat-10.1177_10760296251386028 - Supplemental material for Exploring the Causal Relationship Between Circulating miRNAs and Sepsis Through Mendelian Randomization Analysis
Supplemental material, sj-doc-4-cat-10.1177_10760296251386028 for Exploring the Causal Relationship Between Circulating miRNAs and Sepsis Through Mendelian Randomization Analysis by Yan Jin and Yuan Zhang in Clinical and Applied Thrombosis/Hemostasis
Footnotes
Abbreviation
Ethics Approval
There is no need for informed consent in our study since the unidentified data were free from medical ethics review.
Consent for Publication
Institutional review board approval and informed consent were not required in the current study because research data are publicly available and all patient data are de-identified.
Authors’ Contributions
Yan Jin and Yuan Zhang designed, extracted, analyzed, and interpreted the data from databases. Yan Jin and Yuan Zhang wrote the manuscript and substantively revised it. All authors have read and approved the final manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of Data and Materials
The original contributions presented in this study are included in the article/Supplemental Material. Further inquiries can be directed to the corresponding author(s).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
