Abstract
Background:
Hemophilia A is a X-linked recessive bleeding disorder characterized by qualitative and quantitative deficiency of factor VIII resulting from heterogeneous mutations in the factor VIII gene located in the Xq28 region. Intron 22 inversion (Inv22) mutation is one of the major causes of the protein alteration in factor VIII; its frequency is 40% to 50% in severe patients. Long polymerase chain reaction (PCR) and inverse PCR (I-PCR) have been used for the detection of Inv22 mutation.
Objective:
Development of new protocol for detection of Inv22 mutation.
Method:
We have designed a new method for the detection of Inv22 mutation in complementary DNA (cDNA) of patients. Real-time PCR targeting exons 21 to 22, 22 to 23, and 23 to 24 of factor VIII gene were used in cases with hemophilia A. Samples that were inversion positive by this new method were cross-checked by the conventional I-PCR method. We observed that region between exons 22 and 23 could not be amplified, while in negative cases and controls a 480 bp product is obtained.
Result:
The method was validated in 20 cases with severe hemophilia A by the new cDNA method, and 8 cases were inversion positive, whereas 12 were negative cases. The findings were confirmed by standard I-PCR method. Complete correlation was observed.
Conclusion:
Conventional long PCR and I-PCR methods are work intensive, prolonged, and sometimes difficult to be standardize. The cDNA method is short, involves 3 short-segment amplifications, and is easy to reproduce.
Introduction
Hemophilia A is an X-linked recessive bleeding disorder characterized by qualitative and quantitative deficiency of FVIII resulting from heterogeneous mutations in the FVIII gene located in the Xq28 region. 1 FVIII gene is 186 kb in length and has 26 exons. The organization of the gene is very complex with large introns and genes within introns, such as Int22h-1 and Int22h-2 in intron 22. 2,3 On the basis of FVIII clotting activity, hemophilia is characterized as (1) mild in which FVIII activity ranges between 5% and 35%, (2) moderate with FVIII activity of 1% to 5%, and (3) severe cases with 1% FVIII activity. Cases with hemophilia A experience frequent spontaneous bleeding episodes in joints or soft tissues. 4,5
The most frequent mutation in cases with hemophilia A (CWH) is the intron 22 inversion (Inv22) 6,7 that accounts for 40% to 50% of severe cases worldwide. 8 The Inv22 originates almost exclusively in male meiosis and is mediated by an event of intrachromosomal homologous recombination between a region of 9.5 kb within FVIII intron 22 (int22h-1) and 1 of 2 inversely oriented copies of this sequence (int22h-2, proximal or int22h-3, distal). 9 The Inv22 can be detected by Southern blot analysis, 10 inverse polymerase chain reaction (I-PCR), 6 or long-distance PCR (LD-PCR). 11 The LD-PCR and I-PCR are currently in use. The LD-PCR is difficult to standardize and inter- as well as intra-laboratory variations may occur. We have attempted to device a complementary DNA (cDNA)-based technique for direct detection of Inv22 mutations in CWH.
Materials and Methods
Patient Selection
A total of 20 cases with severe hemophilia A, with FVIII bioassay showing <1% activity were selected for test validation. The FVIII assay was done in all the patients, and written informed consent was obtained prior to the study. The study was approved by the institutional ethical committee. Blood samples were collected in Trizol reagent (Gibco BRL, Thermo Fisher, Germany) in a ratio of 1:6. One part of blood was mixed well with 6 parts of trizol until a homogenized mixture appeared. The mixture was stored in −80°C till further use.
RNA Extraction
Total RNA was extracted from the homogenized mixture of blood and trizol using chloroform and iso-amyl extraction protocol.
Complementary DNA Preparation
The cDNA was prepared from extracted RNA using Fermentas Revert Aid cDNA Extraction kit (Thermo Fisher, Germany). A total reaction mix of 20 μL was prepared using 1 μL of 450 ng of transfer RNA, 2 μL of oligo dT primers, 1 μL of ribonuclease inhibitor, and 1 μL reverse transcriptase enzyme. Volume was made up by DEPC-treated water. The PCR was initially run at 70°C for 5 minutes followed by 42°C for 60 minutes, 70°C for 10 minutes, and 4°C for 10 minutes. The cDNA was stored at −20°C till further use.
Primers
New primers have been designed using DNA Star and Oligo Software (Bosto, MA, USA). Blasting has been done by NCBI software (Nanos, Singapore). The primers were designed to include DNA fragments extending across exonic regions 21 to 22, 22 to 23, and 23 to 24, with the presumption that PCR across 22 to 23 will not yield DNA in Inv22 (Table 1).
Newly Designed Primers for the cDNA Amplification.
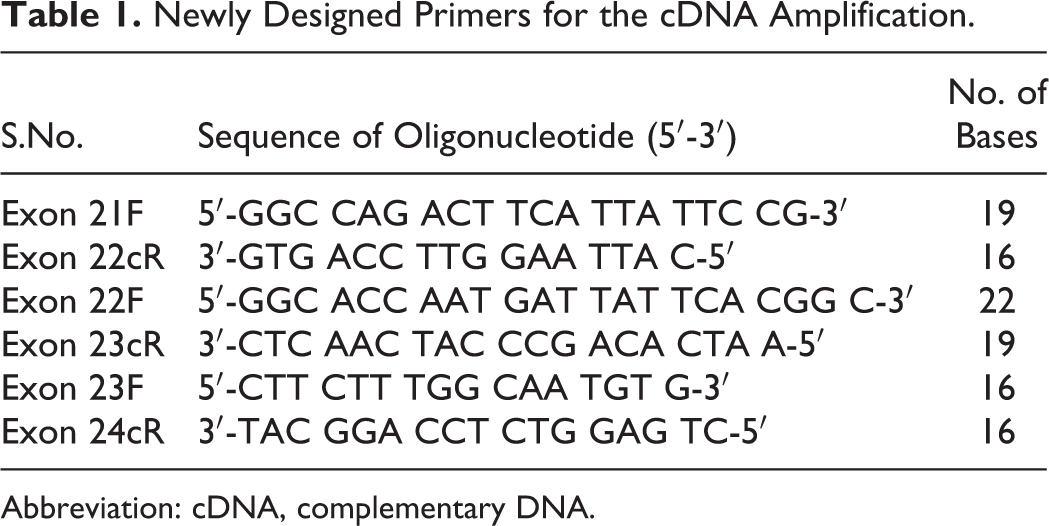

Abbreviation: cDNA, complementary DNA.
Real-Time PCR Amplification
Strata gene Mx3000P (Pro Gene, USA) was used for amplification. Total reaction volume was 20 μL, which comprised 1 μL cDNA and 10 μL of the SYBR GREEN (Takara, Germany) and 2 μL of forward and reverse primers. Final volume was maintained with Milli-Q water, and amplification plot was checked. Exons 21 to 22 gave a PCR product of 333 bp, PCR product of the exons 22 to 23 was 480 bp, and PCR product of the exons 23 to 24 is 393 bp. Good Ct values were also been obtained in the amplified samples, and no Ct values in PCR for exon 22 were obtained in samples positive for Inv22 (Figure 1A-C).
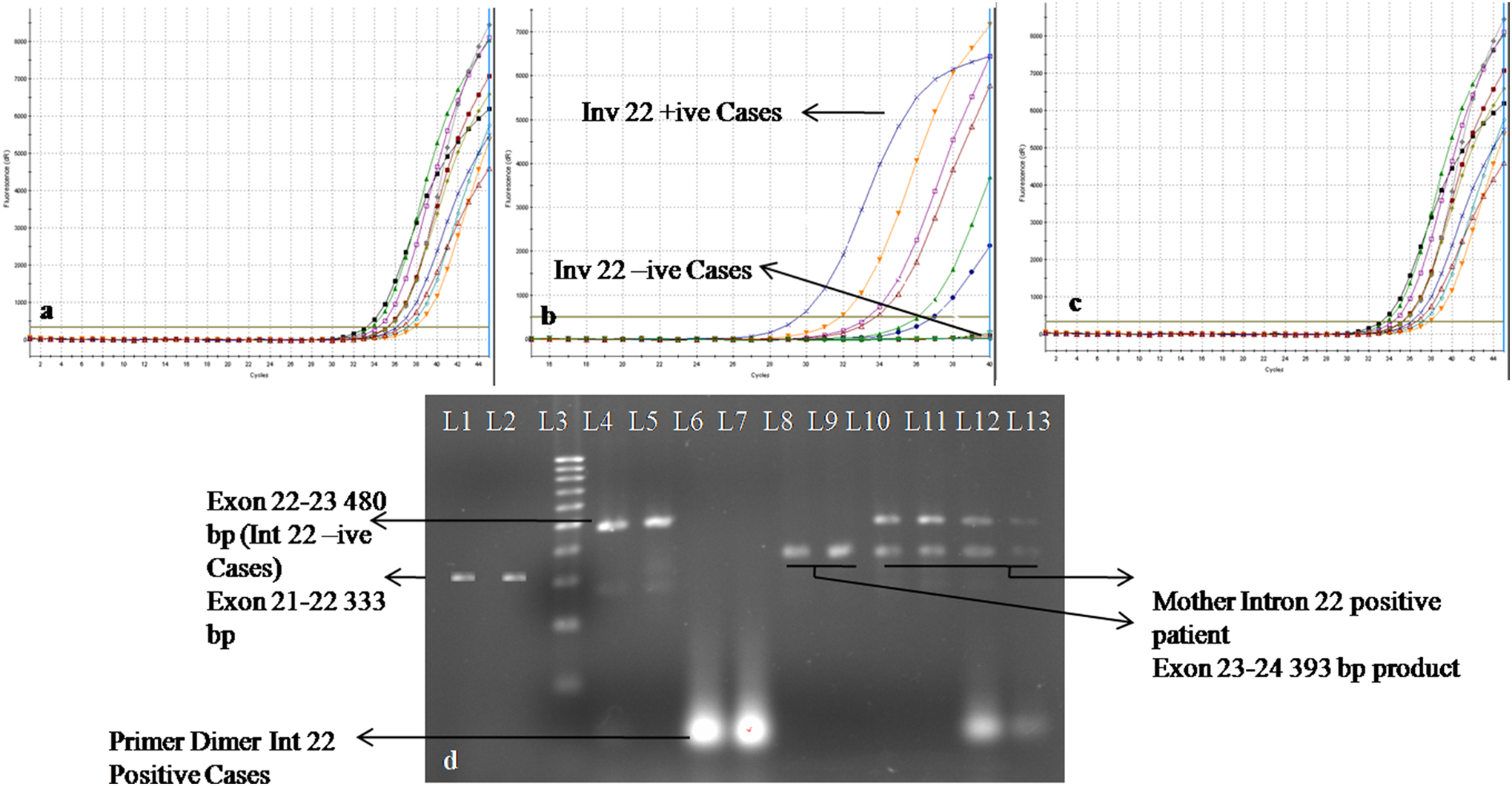
A, Real-time PCR amplification of exon 21 with positive amplification in 10 CWH. B, Real-time PCR amplification of exon 22 with 6 Inv22 negative and 4 Inv22 positive CWH C, Real-time PCR amplification of exon 23 with positive amplification in 10 CWH. D, L1 and L2 shows the gel picture of exons 21 to 22 with a PCR product of 333, L3 = 100 bp ladder (Fermentas), L4 to L5 show the amplification product of Inv22 negative case with a product size of 480 bp band for exons 22 to 23 obtained in case and normal controls, L6 to L7 show the dimer formation of Inv22 positive case, L8 to L9 show the amplification of exons 23 to 24 with the amplification product of 393 bp, and L10 to L13 = mother of CWH showing heterozygous 480 bp and 393 bp bands. CWH indicates cases with hemophilia A; Inv22, intron 22 inversion; PCR, polymerase chain reaction.
Sequencing
The PCR products of exon 21, 22, and 23 were sequenced in 2 controls, 2 Inv22 positive (excl. 22-23 product), and 2 Inv22 negative cases. The sequencing results matched the wild sequence of the FVIII gene using Gene Runner software (Agar Soft Technology, Japan).
Inverse PCR
The Inv22 positive cases were cross-checked by I–PCR that involves the following steps: (1) DNA extraction: genomic DNA was isolated by the Invitrogen Column-based DNA extraction Kit (Invitrogen, Germany). Protocol described by Rosette et al was used with some modifications, (2) BCl I restriction digestion was done overnight. DNA was extracted in phenol chloroform and chloroform iso-amyl alcohol and precipitated in 0.3 mol/L KCl instead of NaCl as used in Rosette et al’s protocol. (3) DNA ligation was performed overnight with T4 DNA Ligase (Fermentas, Germany) followed by PCR amplification of the ligated DNA (Figure 2A and B).
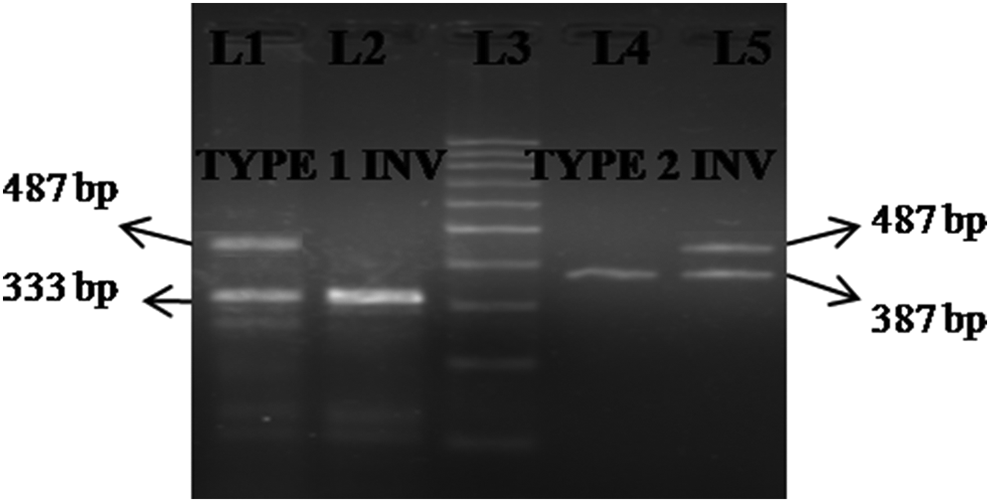
L1 = heterozygous mother of CWH showing 487 and 333 bp bands, L2 = Inv22 positive case type 1 showing 333 bp band, L3 =100 bp ladder (New England Biolabs), L4 = mother of CWH showing 487 and 387 bp bands, and L5 = Inv22 positive case type 2 with 387 bp band. CWH indicates cases with hemophilia A; Inv22, intron 22 inversion.
Interpretation of Results of the New Protocol
For diagnosis of Inv22 3 independent real-time PCR reaction were performed on cDNA using newly designed primers (Table 1). In the first reaction, primers specific for exon 21F-22cR yielded a 333-bp product on gel electrophoresis. In the second reaction using primers specific for Exon 22F-23cR, a 480-bp product is obtained, and in the third reaction using primers specific for 23F-24cR, gel electrophoresis yields a 393-bp product in all the cases. Cases positive for Inv22 failed to show PCR product in second reaction viz exon 22F-23cR. In control samples, amplification in all 3 reactions was evident (Figure 3).
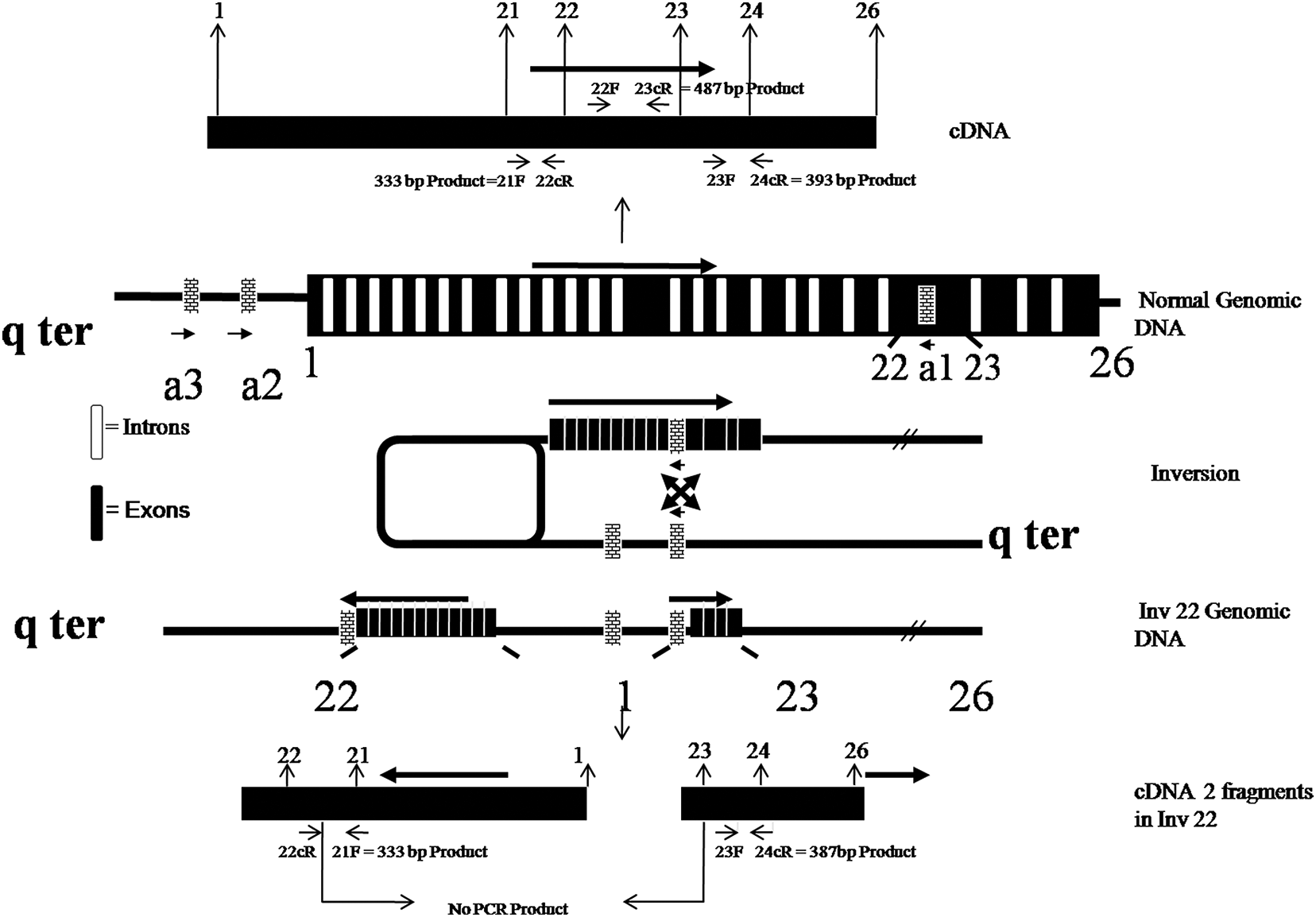
Illustrates the effect of intron 22 inversion (Inv22) on DNA and complementary DNA (cDNA) and shows primers sites of the new protocol.
Results and Discussion
A total of 20 cases with severe hemophilia A and 10 controls have been taken for the study. Using the standard I-PCR method, 8 cases were Inv22 positive, and 12 cases were Inv22 negative. In the new real-time PCR of the cDNA extracted from these cases, 8 CWH with Inv22 showed no amplification of exon 22 and produced no Ct values, while 12 cDNA from CWH with no Inv22 showed positive amplification. Exons 22 and 23 were amplified in all 20 cases with Ct values ranging from 21.06 to 28.8. Controls also showed amplification of all 3 pairs of exons with Ct values ranging from 21.06 to 28.8. The newly designed cDNA protocol matched completely with the results of I-PCR. No abnormal results were obtained in controls. The new method appears to be valid for the detection of Inv22 mutation in CWH (Figure 1A-D).
The size and genomic complexity of the factor VIII gene along with the demonstrated mutational heterogeneity of hemophilia A has been a major limitation for genetic studies of this disorder. 12 The Inv22 is the most common mutation found in cases with severe hemophilia A, occurring in approximately 40% to 50% of cases. 13 This mutation occurs when homologous recombination occurs between the Int22h-1 in intron 22 of FVIII and 1 of 2 homologous regions (Int22h-2 or Int22h-3) telomeric to the FVIII gene. The mutation frequently occurs in the meiotic cell division during the crossover. Somatic mosaicism of the Inv22 mutation caused by a postzygotic de novo mutation has been reported by Lakich et al. 11 The Inv22 mutation was first described in 1993 and was initially detected by Southern blot analysis. This procedure gradually phased out, because it is more time consuming with use of hazardous radio chemicals, and it has become a technique of the past. Further, it is less sensitive and may not detect mosaic mutation. 2 In 1996, Liu et at developed a long PCR for the detection of Inv22. The original protocol used Expand long template PCR kit (Roche, Germany) with 4 primers P, Q, A, and B in a single PCR reaction utilizing a 3-temperature PCR protocol. 14 Reliable consistent results were difficult to obtain with the original protocol and a high background smear, or no product was obtained in many reactions. 14 The long PCR amplification was later modified in 2003 by Poláková et al using a combination of 3 primers in 2 sets of reactions instead of 4 primers in 1 reaction. 15 This decreased the failure rate of the PCR. Although the protocol is now frequently used as the gold standard for diagnosis of Inv22, it has several drawbacks. The fragments to be amplified are very long, and the region is GC rich. We have also faced difficulty in standardization of long PCR, because the amount and quality of DNA required are high, and fragmentation of DNA may occur in extraction with phenol–chloroform method. Therefore, we switched to column-based DNA extraction. 7-Deaza-dGTP is also used in the reaction mixture, as the region is GC rich with 27.5 mmol/L MgCl2 buffer. Reaction failure and smear formation were also observed frequently in our experience. The protocol is time consuming, and the long PCR amplification time extends up to 8 to 9 hours. DNA polymerization occurs, and to resolve signals of 10-, 11-, and 12-kb fragments obtained, a slow electrophoresis run is necessary for 8 to 12 hours. In 2003, Rossette et al developed I-PCR for Inv22 genotyping. This requires an overnight restriction digestion of genomic DNA with Bcl I followed by extraction of fragmented DNA and overnight incubation to achieve Bcl I fragment ligation and subsequent PCR amplification and electrophoresis. 6 A critical step in the I-PCR is extraction of digested DNA. We have used KCl for fragment DNA precipitation instead of NaCl as the precipitation capacity of KCl is more than the NaCl. The I-PCR approach is rapid, robust, and recommended as a diagnostic tool for detection of Inv22 mutation. 15 We have used I-PCR in our study as a standard for comparing the new test. Major drawback in the I-PCR is the amount of time required for obtaining the results. Our protocol requires 2 hours for RNA extraction followed by cDNA synthesis and real-time PCR for exonic regions 21F-22cR, 22F-23cR, and 23F-24cR for diagnosis of this mutation. Although in real-time PCR, the amplification plot demonstrates positive amplification, we have also run gel electrophoresis on 2% agarose gel to check the expected product size in all exonic regions. Band sizes of 333 bp product in exons 21 to 22, 480 bp in exons 22 to 23, and 393 bp in exons 23 to 24 were observed. Further sequencing has been done in amplified DNA from controls as well as in cases to check that the sequence of the amplified product matched with the wild-type sequence of the exonic regions. Conventional PCR with gel electrophoresis may also be used to demonstrate the bands and to reduce the cost. Use of real time also eliminates the gel electrophoresis step. The limitation of this method, however, lies in its inability to detect carrier status, since cDNA would be present also in the carrier. Quantity of cDNA may vary, and Ct values in real-time amplification may play some role in suggesting a carrier status. Further, effect on the test in cases with other large mutations also needs to be checked.
Conclusion
The new method described is easy to standardize and yields quick results. In our hands, the results are unambiguous and provide a definitive diagnosis in CWH with no failure rate. External studies for its application in direct diagnosis of Inv22 mutation in prenatal diagnosis of CWH are required before its clinical implementation.
Footnotes
Authors’ Note
Ethical Approval has been taken by the institutional ethical committee.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was financially supported by CST Lucknow File no: CST/SEPRD/D-3824, and author Praveen Kumar acknowledges the financial support of University Grant Commission (RGNF), New Delhi, India, File no: F.16-786 (RGNF)2007.
