Abstract
Background
Neuroblastoma (NBL) is the most common extracranial solid tumor in childhood, and patients with high-risk neuroblastoma had a relatively poor prognosis despite multimodal treatment. To improve immunotherapy efficacy in neuroblastoma, systematic profiling of the immune landscape in neuroblastoma is an urgent need.
Methods
RNA-seq and according clinical information of neuroblastoma were downloaded from the TARGET database and GEO database (GSE62564). With an immune-related-gene set obtained from the ImmPort database, Immune-related Prognostic Gene Pairs for Neuroblastoma (IPGPN) for overall survival (OS) were established with the TARGET-NBL cohort and then verified with the GEO-NBL cohort. Immune cell infiltration analysis was subsequently performed. The integrated model was established with IPGPN and clinicopathological parameters. Immune cell infiltration was analyzed with the XCELL algorithm. Functional enrichment analysis was performed with clusterProfiler package in R.
Results
Immune-related Prognostic Gene Pairs for Neuroblastoma was successfully established with seven immune-related gene pairs (IGPs) involving 13 unique genes in the training cohort. In the training cohort, IPGPN successfully stratified neuroblastoma patients into a high and low immune-risk groups with different OS (HR=3.92,
Conclusion
Herein, we presented an immune landscape with IPGPN for prognosis prediction in neuroblastoma, which complements the present understanding of the immune signature in neuroblastoma.
Keywords
Introduction
Neuroblastoma (NBL) is the most common extracranial solid tumor in childhood, originating from neurons of the sympathetic nervous system, 1 and accounting for 8–10% of pediatric cancers in the USA and Europe.2,3 According to age at diagnosis, the extent of disease, tumor subtype, and disease course can vary from spontaneous regression to implacable progression even with intensive multimodality therapy.1,4 With precise stratification generally accepted, neuroblastomas can be divided into three major subtypes, subtypes 1, 2A, and 2B, based on genetic features. 4 While subtype 1 neuroblastoma is associated with favorable features such as young age and lower tumor stage, subtypes 2A and 2B are associated with advanced tumor stage and worse prognosis, with subtype 2B being the most aggressive type. 5 However, despite the individualized therapeutic strategy brought by a well-defined subset, metastasis occurred in nearly half of the neuroblastoma patients. The five-year event-free survival (EFS) in high-risk neuroblastoma patients remains less than 50%. 6
Dysregulated tumor immune microenvironment is a critical hallmark of neuroblastoma. The tumor microenvironment is a rather complex composition of surrounding blood vessels, fibroblasts, immune cells, signal molecules, and extracellular matrix, 7 and interaction between tumor cells and these microenvironment components could significantly modulate the malignant phenotype of tumor cells, including proliferation, cancer cell stemness, and metastasis potential.8-10 In contrast, tumor cells could also shape the surrounding microenvironment to form a more suitable tumor niche through angiogenesis promotion. 11 Given immune signatures in neuroblastoma, numerous immune molecules, including NF-κB, 12 tumor necrosis factor (TNF), 13 and interleukins, 14 have been reported to be associated with tumorigenesis, apoptosis, and chemoresistance in neuroblastoma. Moreover, the mutual interaction between neuroblastoma cells and immune cells, such as tumor-associated macrophages (TAMs) and tumor-infiltrating lymphocytes (TILs), may further contribute to tumor proliferation, metastasis, and adaptive chemoresistance.15,16 It has come to the researchers that neuroblastoma could exhibit either immunogenic or non-immunogenic type, 17 but there is still a lack of ample evidence considering the immune landscape in neuroblastoma.
Our team has previously established a prognostic model in neuroblastoma with TARGET-NBL cohort and GEO-NBL cohort (GSE62564), 18 but the immune landscape of neuroblastoma remained unraveled. Herein, in the present study, we established and validated a prognostic immune-signature in neuroblastoma based on RNA sequencing data of totally 644 cases from these two NBL cohorts, which exhibited satisfactory stratification for both overall survival (OS) and event-free survival (EFS). Compared with the prognostic NBL signature reported previously, 18 though the same participant cohorts were used, our prognostic model was established with a different gene set and provided a more specific and comprehensive view focused on the NBL immune landscape, which is not elaborated in the previous publications.
Materials and Methods
RNA-Seq and Clinical Data Curation
The overall workflow was shown in Figure 1. Neuroblastoma RNA-seq data and corresponding clinical data were downloaded from the TARGET database (https://ocg.cancer.gov/programs/target) and GEO database (https://www.ncbi.nlm.nih.gov/geo, GSE62564).
19
The TARGET neuroblastoma cohort was set as a training cohort (n=152), while the GEO neuroblastoma cohort was selected as a validation cohort (n=492). No significant bias was found between the TARGET training cohort and the GEO validation cohort. The overall workflow of the study.
Identification of Immune-Related Gene Pairs (IGPs)
Identification of prognostic IGPs was performed as described previously. 20 A total of 1811 immune-related genes (IGs) were retrieved from ImmPort database, 21 all measured in the TARGET and GEO cohort. IGs with relatively high variation, as determined by median absolute deviation (MAD) more than .5, were selected. Each IGP was calculated by pairwise comparison of each sample’s gene expression level. Precisely, during a pairwise comparison, the output is 1 if the first gene expression level is higher than the later one, and 0 for other different orders. After removing IGPs with constant ordering, 512 IGPs remained and were pooled into further prognosis prediction to establish IPGPN.
Establishment and Validation of Immune-Related Prognostic Gene Pairs for Neuroblastoma (IPGPN)
The establishment of a prognostic signature based on IGPs was performed as described previously. 20 With the remaining IGPs, an immune signature IPGPN was established with the least absolute shrinkage and selection operator (LASSO) regression. To stratify patients into high/low-risk groups, the optimal cut-off value of the IPGPN was determined with time-dependent receiver operating characteristic (ROC) curve analysis, 22 at 5 years in the TARGET training cohort for overall survival. The prognostic prediction value for the overall survival of IPGPN was validated in the GEO cohort using the log-rank test. The sensitivity and specificity of IPGPN were also determined using ROC curve analysis.
Immune Cell Infiltration Analysis
To profile immune cell infiltration in different risk groups, Immunedeconv (2.0.3) package in R was used to characterize the immune-cells composition based on tumor gene expression profiles.
Functional Enrichment Analysis
To perform functional enrichment analysis, including KEGG pathway analysis and GSEA, differentially expressed genes (DEGs) were first confirmed with limma package in R and then analyzed with clusterProfiler package in R.
Statistical Analysis
Continuous variables were presented as mean ± SD, and categorized variables were presented as frequency (n) and proportion (%). The prognostic value for OS and EFS was evaluated by the multivariate Cox regression model in the survival analysis. Kaplan–Meier survival analysis was performed to compare different subgroups. Single, double, and triple asterisks indicate statistical significance, *P< .05, **P< .01, and ***P< .001, respectively.
Results
Identification and Definition of the IPGPN
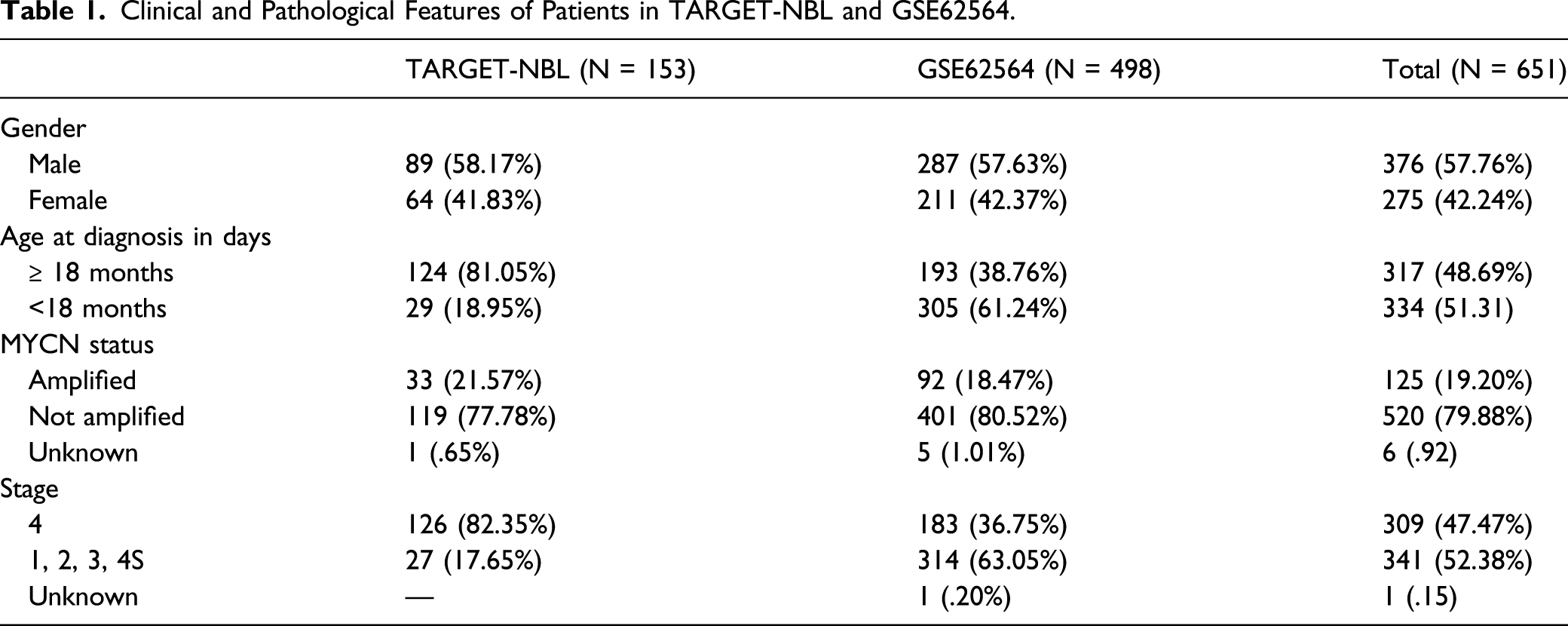
Clinical and Pathological Features of Patients in TARGET-NBL and GSE62564.
Details of 7 Gene Pairs.

IPGPN Exhibited Satisfactory Performance on OS and EFS Prediction
For OS prediction, the optimal cut-off of the IPGPN was set as −2.084, determined by time-dependent ROC curve analysis (Figure 2). Of note, IPGPN successfully stratifies neuroblastoma patients in the TARGET training cohort into low and high immune risk groups with diametrically different OS in the training set (Figure 3A), in which high IPGPN score predicted an unfavorable prognosis (HR=3.92, 95%CI=2.35-6.53; Establishment of IPGPN based on Immune-related Prognostic Gene Pairs. (A-B) The risk score considering IPGPN of each NBL patient in the training and validation cohort, and the cut-off value between high- and low-risk groups. (C-D) Survival status of each NBL patient in the training and validation cohort. (E-F) Heatmap of the seven immune-related gene pairs in the training and validation cohort. Abbreviations: IPGPN, Immune-related Prognostic Gene Pairs for Neuroblastoma; NBL, neuroblastoma. IPGPN is prognostic for OS in NBL patients. (A) OS analysis of NBL patients in the low- and high-risk group in the training cohort. (B) ROC curve demonstrating the prognostic value of IPGPN in predicting OS in the training cohort. (C) OS analysis of NBL patients in the low- and high-risk group in the validation cohort. (D) ROC curve demonstrating the prognostic value of IPGPN in predicting OS in the validation cohort. Abbreviations: IPGPN, Immune-related Prognostic Gene Pairs for Neuroblastoma; NBL, neuroblastoma; ROC; receiver operating characteristic.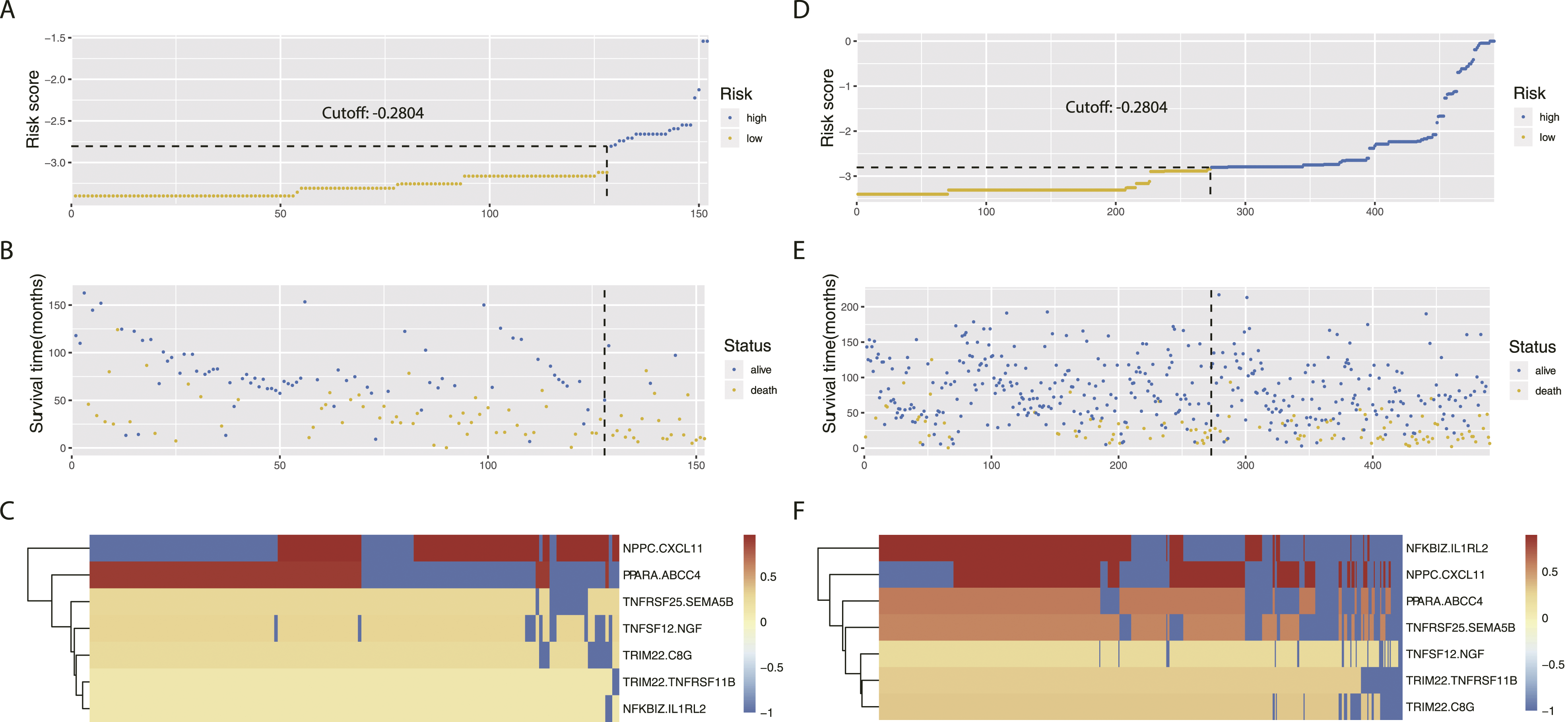

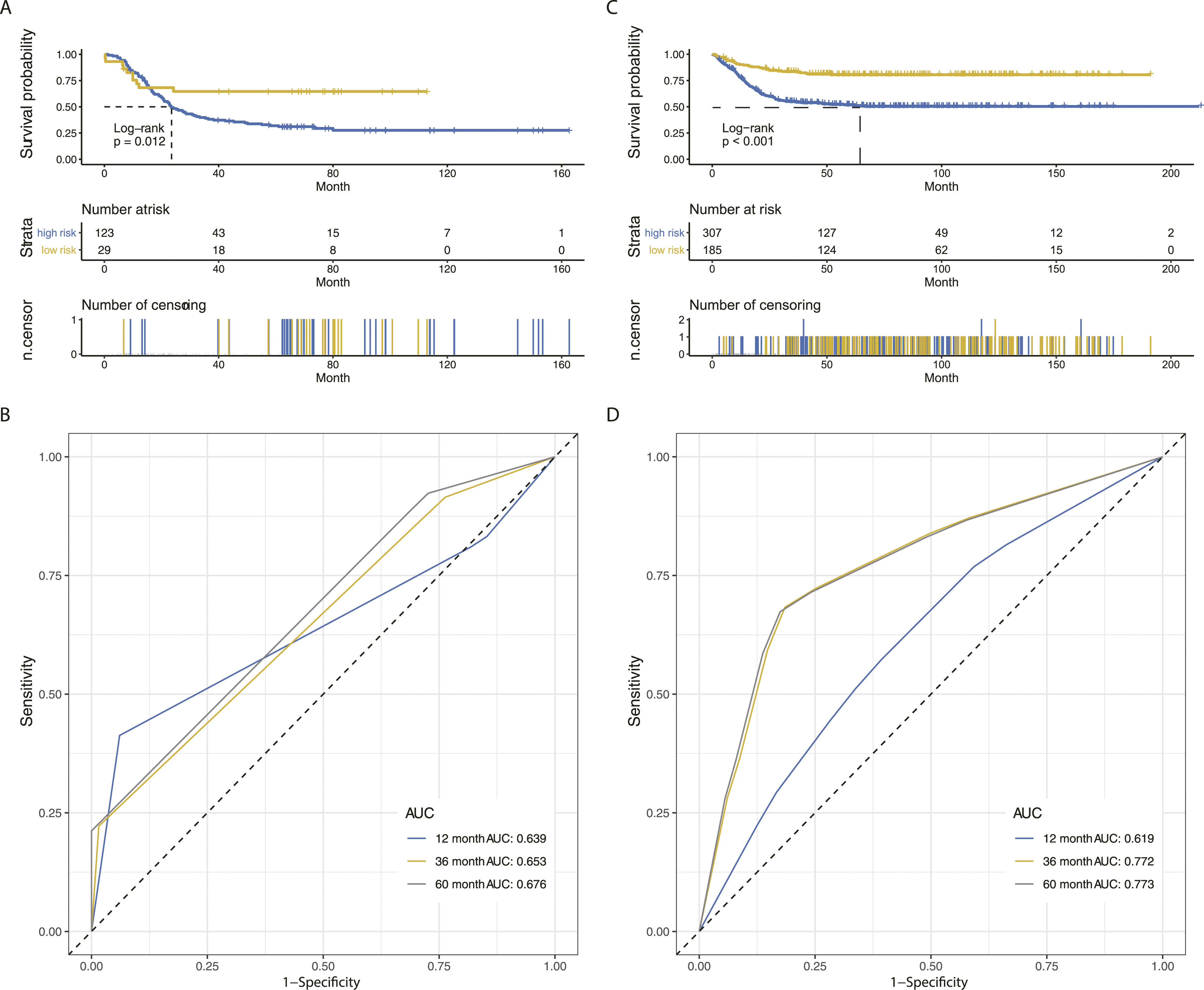
We also interrogate the predictive value of IPGPN on EFS. Intriguingly, the TARGET-NBL cohort was similarly divided into two distinct groups in terms of EFS, and a low IPGPN score is correlated with a better EFS (Figure 4A; HR=3.66, 95%CI 2.25-5.95; IPGPN is prognostic for EFS in NBL patients. (A) EFS analysis of NBL patients in the low- and high-risk group in the training cohort. (B) ROC curve demonstrating the prognostic value of IPGPN in predicting EFS in the training cohort. (C) EFS analysis of NBL patients in the low- and high-risk group in the validation cohort. (D) ROC curve demonstrating the prognostic value of IPGPN in predicting EFS in the validation cohort. Abbreviations: EFS, event-free survival; IPGPN, Immune-related Prognostic Gene Pairs for Neuroblastoma; NBL, neuroblastoma; ROC; receiver operating characteristic.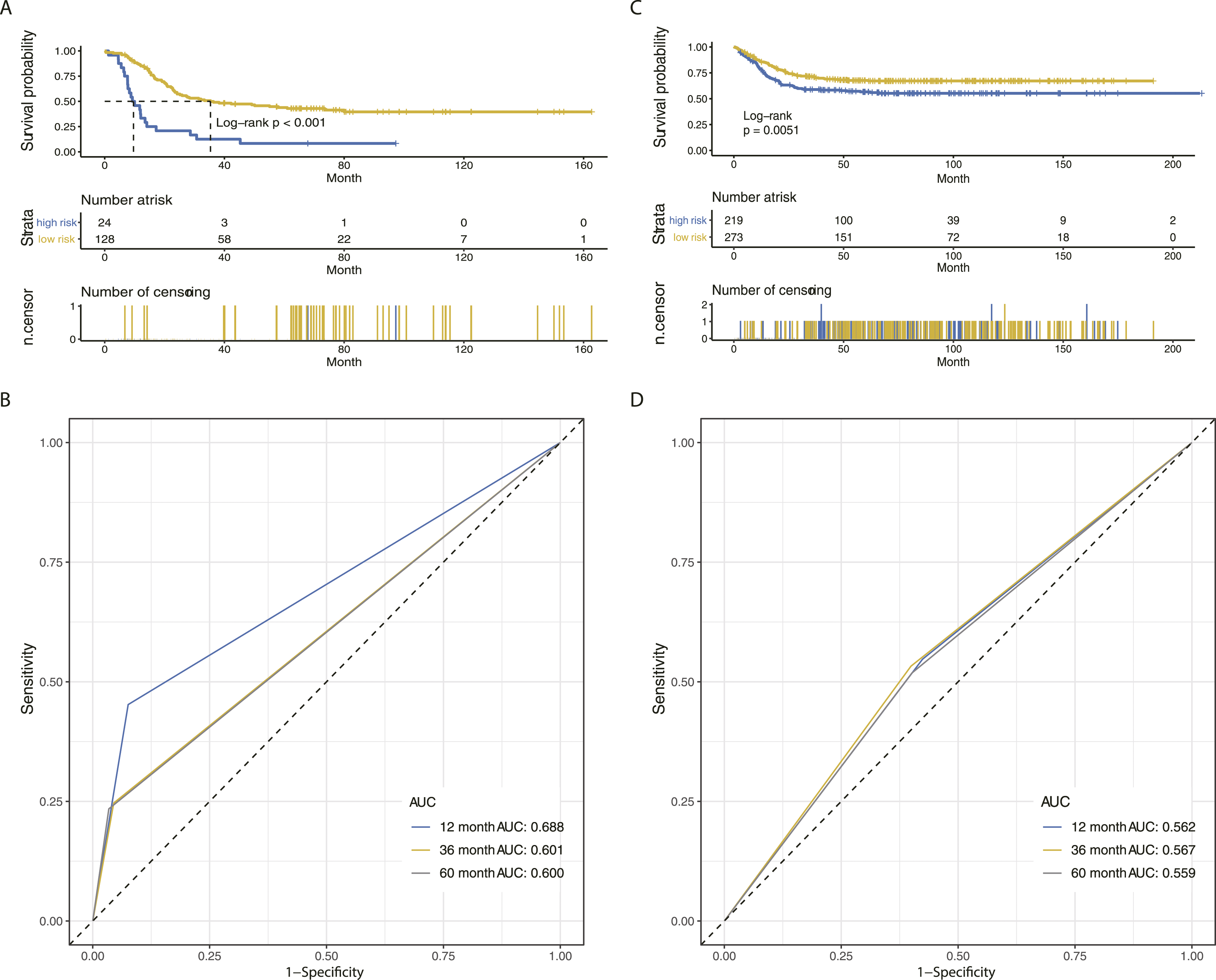
Validation of IPGPN
To further interrogate the predictive value of IPGPN in other populations, we then applied the same formula to the GEO-NBL cohort (GSE62564). Consistently, for OS, IPGPN significantly divided GEO-NBL patients into a low and high-risk group, in which a high IPGPN score predicted an extended OS (Figure 3C; HR=1.84, 95%CI=1.24-2.71;
Immune Cell Infiltration Analysis Based on IPGPN
Emerging evidence links increased immune cell infiltration in the tumor microenvironment with tumorigenesis and metastasis. To further profile the infiltration of different immune cells, including macrophages, B cells, and T cells, the XCELL algorithm was applied in both training and validation sets. As shown in Figure 5, activated dendritic cells (P=7.27 × 10−4 and 3.51 × 10−7 in the training and validation cohort, respectively) and M1 macrophage (P=6.83 × 10−6 and 1.61 × 10−9 in the training and validation cohort, respectively) were enriched in the low immune risk group, while Th1 CD4+ (no significant difference was observed in the training cohort while P=4.51 × 10−15 in the validation cohort) and Th2 CD4+ T cells (P=.01 and 5.85 × 10−12 in the training and validation cohort, respectively) were enriched in the high immune risk group, indicating a more active immune response predicted a favorable prognosis. Immune cell infiltration analysis based on Immune-related Prognostic Gene Pairs for Neuroblastoma. (A-D) In the training cohort, enrichment of myeloid dendritic cells and M1 macrophages was observed in the low immune risk group, while enrichment of Th1 and Th2 subsets of CD4+ T cells was observed in the high immune risk group. (E-H) In the validation cohort, enrichment of myeloid dendritic cells and M1 macrophages was observed in the low immune risk group, while enrichment of Th1 and Th2 subsets of CD4+ T cells was observed in the high immune risk group.
Integrated Prognostic Signature Combing IPGPN and Clinicopathological Characteristics
Univariate and Multivariate Cox Regression Analyses of Clinicopathological Factors in OS.

Abbreviations: IPGPN, Immune-related Prognostic Gene Pairs for Neuroblastoma.

Integrated prognostic signature combing IPGPN and clinicopathological characteristics is prognostic for OS in NBL patients. (A-B) OS analysis of NBL patients in the high- and low-risk group considering integrated prognostic signature in the training and validation cohort. (C-D) ROC curve demonstrating the prognostic value of integrated prognostic signature in predicting OS in the training and validation cohort. Abbreviations: IPGPN, Immune-related Prognostic Gene Pairs for Neuroblastoma; NBL, neuroblastoma; ROC; receiver operating characteristic.
Integrated prognostic signature combing IPGPN and clinicopathological characteristics is prognostic for EFS in NBL patients. (A-B) EFS analysis of NBL patients in the high- and low-risk group considering integrated prognostic signature in the training and validation cohort. (C-D) ROC curve demonstrating the prognostic value of integrated prognostic signature in predicting EFS in the training and validation cohort. Abbreviations: EFS, event-free survival; IPGPN, Immune-related Prognostic Gene Pairs for Neuroblastoma; NBL, neuroblastoma; ROC; receiver operating characteristic.

Nomogram for both OS and EFS prediction. (A) Nomogram combing IPGPN and clinicopathological characteristics for 1-year, 3-year, and 5-year OS prediction in the training cohort. (B) Calibration plot showing the ideal prediction (black dotted line) and the nomogram-prediction (blue line with the bars representing the 95% CI) in the training cohort. (C) Nomogram combing IPGPN and clinicopathological characteristics for 1-year, 3-year, and 5-year OS prediction in the validation cohort. (D) Calibration plot showing the ideal prediction (black dotted line) and the nomogram-prediction (blue line with the bars representing the 95% CI) in the validation cohort. Abbreviations: EFS, event-free survival; IPGPN, Immune-related Prognostic Gene Pairs for Neuroblastoma; OS, overall survival.
Functional Enrichment Analysis
A total of 10 009 DEGs were confirmed between the low and high immune-risk groups in the TARGET-training cohort. Functional enrichment analysis was then performed between low and high immune risk groups in the TARGET-training cohort. GSEA demonstrated that gene sets of NK cells mediated cytotoxicity and Th1/Th2 CD4+ cells were significantly activated in the low immune risk group, indicating the critical roles of NK cells and CD4+ cells in neuroblastoma (Figure 8A and 8B). Enrichment of other cancer-immunity-related gene sets, such as antigen processing and lysosome, was also observed (Supplementary Table 1).
Discussions
There is emerging attention on tumor-associated immune cells and molecular events governing immune responses in NBL. Frequent spontaneous regression mediated by cellular immunity was observed in children less than 18 months.
23
Notably, neuroblastoma is the first pediatric tumor for which immunotherapy (dinutuximab, a chimeric anti-GD2 monoclonal antibody) was approved. Still, the cure rates remain less than 50%, and many patients failed to achieve a minimal residual disease state.
24
Therefore, systematic profiling of NBL immune signature is warranted to display the immune landscape in NBL and guide the following immunotherapy. This study established IPGPN, a prognostic immune signature for neuroblastoma, through the specific pairwise correlation between different immune-related genes' expression. With the TARGET-NBL cohort, we finally selected 7 IGPs with 13 IGs to establish IPGPN, which successfully stratified NBL patients into high and low immune risk groups with distinctively different survival outcomes. Subsequently, the predictive value of IPGPN for both OS and EFS in neuroblastoma was verified in the GEO-NBL cohort. Moreover, immune cell infiltration analysis demonstrated that the low immune risk group might possess a more active immune response. Furthermore, an integrated model was established by including age, stage, and MYCN to IPGPN, which exhibited better predictive power than IPGPN alone (Figure 9). Functional enrichment analysis of DEGs between low- and high-risk groups. (A) Enrichment plots for the nature killer cell mediated cytotoxicity signature. (B) Enrichment plots for the Th1 and Th2 cell differentiation signature.
Expression of immune-related genes could be a reliable predictor in neuroblastoma. It was found that 14% of neuroblastoma expressed programmed death-ligand 1 (PD-L1), and these patients were more prone to unfavorable prognosis than those PD-L1 negative ones. 25 Complementary to the current understanding of tumor immune in neuroblastoma, IPGPN was established with seven gene pairs involving 13 immune-related genes. Compared to a single-gene signature, a gene signature built on the pairwise correlation of gene expression is more robust and accountable across different datasets. 26 A handful of these genes are involved in the immune response under specific pathological conditions. For example, mutation of NFKBIZ was found to be associated with downregulation of pro-inflammatory factor and against colorectal carcinogenesis. 27 As a tripartite motif-containing (TRIM) superfamily member, TRIM22 plays a vital role in immune response facing viral infection by modifying monocyte functions.28,29 Fewer of them have been discussed in neuroblastoma. CXCL11, a potent CXCR3 ligand released by activated NK cells, has already been reported to mediate the cytotoxicity of anti-GD2 antibody in neuroblastoma. 30 Other genes, including ABCC4, 31 TNFRSF25, 32 and NGF 33 are found to be dysregulated in neuroblastoma, yet future study into their immune response role is needed.
Interaction between immune-related genes and immune cells also attracted attention from researchers. On the one hand, several immune cells were found to be oncogenic in neuroblastoma. A subset of neuroblastoma cell lines was found to express membrane-bound TNF-α, which further induced TAMs to release CCL20. 34 Macrophages were prone to be identified in PD-L1 positive neuroblastoma, associated with poor prognosis. 25 In particular, M2 macrophage infiltration was correlated with both the extent of disease and event-free survival in patients with MYCN-non-amplified neuroblastoma. 35 Depletion of TAMs by blockade of colony-stimulating factor 1 receptor (CSF-1R) could sensitize neuroblastoma cells to cyclophosphamide and topotecan regimen. 36 Likewise, the lack of regulatory T cells, which are immune suppressive, improved the efficacy of anti-GD2 antibody treatment. 37
On the other hand, activation of a specific type of immune cell could synergize immunotherapy and was associated with a favorable outcome. Increased signatures of activated NK cells and CD8+ T cells were associated with a favorable prognosis in high-risk neuroblastoma. 38 Therefore, a reversal of the immune-suppressive environment through NK cells' modulation is promising. NKG2D.ζ–NK cell, a gene-modified type of NK cell that targeted myeloid-derived suppressor cells, enhances CAR-T cell activity in neuroblastoma. 39 Likewise, enhanced uptake of cancer-derived neoantigens by dendritic cells could stimulate CD8+ T cells' antitumor effect, 40 indicating the vital role of antigen processing in cancer immunity. Following this, we found prominent enrichment of NK cells, Th1/Th2 CD4+ cells, and antigen processing gene sets in the immune low-risk group, indicating an active immune response positively correlated with a favorable prognosis. Therefore, immune modulation focused on NK cells and CD4+ cells may synergize immunotherapy in the high immune risk group.
Though immune checkpoint inhibitors have achieved satisfying efficacy in several adult cancers, most of them failed in neuroblastoma due to their cold immunogenic nature. 41 Several approaches have been suggested to improve the immune response of cold neuroblastoma by reprogramming innate immune systems. For those neuroblastomas with inadequate immune response, activation of the simulator of interferon genes (STING) pathway could convert them into a more T-cell enriched microenvironment and enhance their response to immune checkpoint blockade. 42 While the addition of antibody against cytotoxic T-lymphocyte-associated protein 4 (CTLA-4), an immune checkpoint blockade depleting T regulatory cells (Tregs), to combined therapy of radiation and intratumor injection of immunocytokine could not lead to complete regression of N-MYC driven cold neuroblastoma, further addition of CpG-oligodeoxynucleotides and anti-CD40 mAb to this combined regimen could result in complete response through activating innate immune system. 43 Furthermore, though inhibition of programmed cell death protein 1 (PD-1) alone could not lead to neuroblastoma regression, concurrent inhibition of CSF-1R could significantly promote T-cell infiltration through upregulating the expression of recruiting chemokines and downregulating suppressive myeloid cells. 44 Based on the comprehensive immune profiling of neuroblastoma in our research, neuroblastoma with low immune response and specific modification of innate immune to enhance their response to immune checkpoint blockade treatment could be further identified.
Besides of gene expression profiles, several clinicopathological characteristics could be predictive for prognosis and clinical efficacy in neuroblastoma. For example, patients with older age at diagnosis (>18 months), independent from the stage, usually had a worse prognosis.45-47 Moreover, MYCN status is associated with therapeutic response to immunotherapy. MYC and MYCN were found to regulate the expression of PD-L1 in neuroblastoma, 48 and MYCN could further inhibit NK cell activation in high-risk neuroblastoma. Consistently, we confirmed that age, stage, and MYCN are independent prognosis predictors. These three parameters could further complement IPGPN with better predictive power than IPGPN alone or clinicopathological factor alone.
Our study has several advantages compared with other existing models. First, compared with the previous NBL signature, 18 IPGPN focused on immune-related genes and delineated the immune cell composition in NBL. Second, compared with the previous models established with immune-related genes in NBL,49,50 IPGPN was established with pairwise correlation, reflecting the landscape of immune-related tumor microenvironment more accurately across different populations and reducing specific cross-study batch effects. Third, while IPGPN was established with the TARGET neuroblastoma cohort in terms of OS, it presents with a satisfactory performance for both OS and EFS prediction in the GEO neuroblastoma cohort, indicating the robustness of this model among different neuroblastoma cohorts.
Limitations of our study should also be noted. Though gene-pair calculation reduced the batch effects to some extent, it led to the low AUC values around 0.6. Moreover, further studies to validate this model both
In conclusion, our study provided a comprehensive understanding of immune signatures in neuroblastoma based on the pairwise correlation of immune-related genes and cellular compartment within different immune risk groups. IPGPN, a prognostic immune signature proposed, is promising for prognosis prediction and distinguishing neuroblastoma with different immune responsiveness, which guides immunotherapy to overcome immune therapy resistance.
Supplemental Material
sj-pdf-1-ccx-10.1177_10732748211033751 – Supplemental Material for Establishment and Validation of a Prognostic Immune Signature in Neuroblastoma
Supplemental Material, sj-pdf-1-ccx-10.1177_10732748211033751 for Establishment and Validation of a Prognostic Immune Signature in Neuroblastoma by Yunhu Yu, Yu Zeng, Xiangping Xia, Jian-Guo Zhou and Fang Cao in Cancer Control
Footnotes
Acknowledgments
The study funders had no role in the study’s design; the collection, analysis, or interpretation of the data; the writing of the manuscript; or the decision to submit the manuscript for publication.
Authors Contributions
YY and FC conceived, designed, or planned the study. YZ, YY, and JGZ analyzed the data. FC acquired data. JGZ and YZ helped interpret the results. JGZ and FC provided study materials or patients. YZ,YY, JGZ, XPX, and FC drafted the manuscript. All authors revised and reviewed this work, and all authors gave their final approval of the submitted manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the Natural Science Foundation of China (NSFC: 81660421, 81 860 450). Outstanding Youth Foundation of Zunyi medical university (2018-05).
Supplementary Material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
