Abstract
Campylobacter jejuni is a leading cause of human foodborne illness, and pet dogs can be a source of infection. Despite recognition of this zoonotic potential, tests for detection of C. jejuni are not performed routinely in dogs. We compared bacterial culture and molecular methods for the detection of C. jejuni directly from canine feces and from Bolton enrichment broth inoculated with feces in a multi-laboratory randomized-method test. Although culture offers the advantage of collection of isolates for further epidemiologic study, real-time PCR (rtPCR) to detect cpn60 from DNA isolated directly from canine feces was more sensitive than culture and more accurate than rtPCR using DNA isolated from enrichment cultures. The limit of detection for our cpn60 rtPCR was 320 cfu/g.
Keywords
Campylobacter is the leading cause of foodborne gastrointestinal disease in people worldwide. 44 Campylobacteriosis affects an estimated 1.3 million people in the United States each year, and the incidence of Campylobacter infections appears to be increasing based on the Centers for Disease Control and Prevention FoodNet data.40,41 People infected with Campylobacter develop diarrhea, cramping, and abdominal pain 2–5 d after exposure. 11 These signs usually last about one week. People typically become infected through the consumption of raw or undercooked poultry or beef, but outbreaks have also been associated with unpasteurized dairy products, contaminated water, and produce. 42 Until recently, most people would not consider dogs as a likely source of Campylobacter despite documented cases of transmission and evidence that genetically identical strains may be found in people and their pets.3,5,33 An outbreak of multidrug-resistant C. jejuni associated with puppies in pet stores, affecting a total of 143 people in 21 states, has increased awareness of the potential for transmission of Campylobacter from dogs to people.12,13,23,32
The 3 most common species of Campylobacter found in fecal samples from dogs are C. jejuni, C. upsaliensis, and C. helveticus.16,25,29,38 Multiple studies have demonstrated the variability of C. jejuni prevalence in dogs. One study found that among 251 dogs that visited dog parks in Ontario, Canada, 108 (43%) were shedding Campylobacter spp.; of these, 14 (13%) were shedding C. jejuni. 38 Another study in Saskatoon, Canada, found that 5 of 70 (7%) healthy dogs carried C. jejuni; 30 of 65 (46%) dogs with diarrhea were shedding the organism. 16 In a study from Colorado, 10 of 130 (0.8%) dogs were shedding C. jejuni. 25 In Texas, 10 of 185 (5.4%) dogs at animal shelters were positive for C. jejuni, 29 and isolates from 1 dog were resistant to ciprofloxacin, 27 which is commonly used to treat people with campylobacteriosis.
As with humans, young animals more commonly have clinical signs associated with Campylobacter infection; most adult dogs exposed to Campylobacter do not become clinically ill. 38 However, adult animals may become colonized and shed Campylobacter in their feces for several weeks following exposure. 37 Animals with campylobacteriosis have clinical signs similar to those in people, including fever, anorexia, vomiting, signs of abdominal pain, and diarrhea. 30 Also, like humans, dogs become infected by ingesting the organism in contaminated foods, including raw meat, which is frequently contaminated with C. jejuni.7,36,43 Dogs fed commercially prepared dry dog food are less likely to shed C. jejuni. 8 Despite the known risks of feeding raw-meat diets to dogs, up to 16.2% of pet dogs receive raw meat or bones as a treat or snack weekly. 26 Given the dietary risks and the popularity of dogs as pets (~57 million [45%] of U.S. households include dogs), 6 as well as the recent outbreak associated with dogs,12,23 tests for the rapid detection of C. jejuni from specimens from dogs are needed.
The American College of Veterinary Internal Medicine consensus statement on the detection of enteropathogens in dogs recommends culture or fecal real-time PCR (rtPCR)-based methods for the detection of Campylobacter. 30 Culture methods take up to 5–7 d for results, and C. jejuni growth may be obscured by normal microbiota.9,18 Culture offers the benefit of recovery of bacteria that can be tested for antimicrobial susceptibility; rtPCR is more sensitive. A variety of methods have been used to test populations of dogs for C. jejuni.2,4,10,37 With culture methods, fecal samples are plated on selective media containing antibacterial and antifungal agents that limit the growth of other organisms, which could obscure the observation of C. jejuni. 4 A combination of culture and PCR methods has been reported to improve Campylobacter detection.10,37 These combinations included the use of PCR for confirmation of cultured isolates, 10 as well as direct detection of Campylobacter spp. in DNA extracted from canine feces16,37 or enrichment broth. 37
C. jejuni can be difficult to culture and detect. Evaluations of proficiency testing have found that the organism is frequently not detected (or is most likely to be missed) in performance evaluation studies.20,24 A 1986 study from the Centers for Disease Control and Prevention summarizing 6 y of data found that 10% of medical laboratories were unable to isolate C. jejuni based on proficiency tests. 24 A 2009 study evaluated 9 y of qualitative proficiency test data from food microbiology laboratories and found an average false-negative rate of 13.6% for samples containing Campylobacter spp., nearly double the rate for other enteric pathogens. 20 These studies confirm that C. jejuni is a difficult organism to use in proficiency testing and may be challenging for laboratories to detect.
The U.S. Food and Drug Administration, Veterinary Laboratory Investigation and Response Network (FDA Vet-LIRN) supports the development and standardization of methods for the detection of pathogens from veterinary patient specimens.31,34,39 Our study, funded by Vet-LIRN, was designed to establish a rapid, robust, straightforward method for the identification of C. jejuni in canine fecal samples. Using the cpn60 gene identified in previous studies14,15,16,29 as the target for rtPCR, we compared culture and rtPCR methods with and without enrichment in a multi-laboratory randomized method test in 2015. Afterward, an inter-laboratory comparison exercise (
A randomized method test (
Materials and methods
Preparation of fecal samples inoculated with Campylobacter jejuni
Voluntarily voided, fresh canine fecal samples were collected and confirmed to be negative for Campylobacter species, including C. jejuni, by culture and PCR ( Fig. 1 ). Because feces consisted exclusively of voided specimens, our study was considered exempt from the Texas A&M University Institutional Animal Care and Use Committee guidelines. The feces were then mixed using a stomacher to create a uniform canine feces matrix. In a preliminary test, C. jejuni ATCC 33560T and wild-type isolates of C. jejuni from dogs were evaluated for their ability to survive in canine feces held at 4°C for 7 d. C. jejuni ATCC 33560T was viable for 9 d during this testing and was selected for further use. To inoculate feces, cultures of C. jejuni ATCC 33560T were grown in Bolton enrichment broth (Hardy Diagnostics), diluted in PBS (sourced from either Sigma-Aldrich or ThermoFisher), and used to inoculate fecal samples at an average of ~1,000, 100, 10, and 0 cfu/g of feces. To confirm the number of C. jejuni cfu/g of feces inoculated, a 100-µL aliquot of the Bolton broth culture was used to create 10-fold serial dilutions in Butterfield phosphate buffer (Weber Scientific) and plated in duplicate to either trypticase soy agar plates supplemented with 5% sheep blood (BAP; BD) or Campyfood agar plates (bioMérieux). Plates were incubated at 42 ± 2°C under microaerophilic conditions for 48 h, manually counted, and averaged to determine the count.
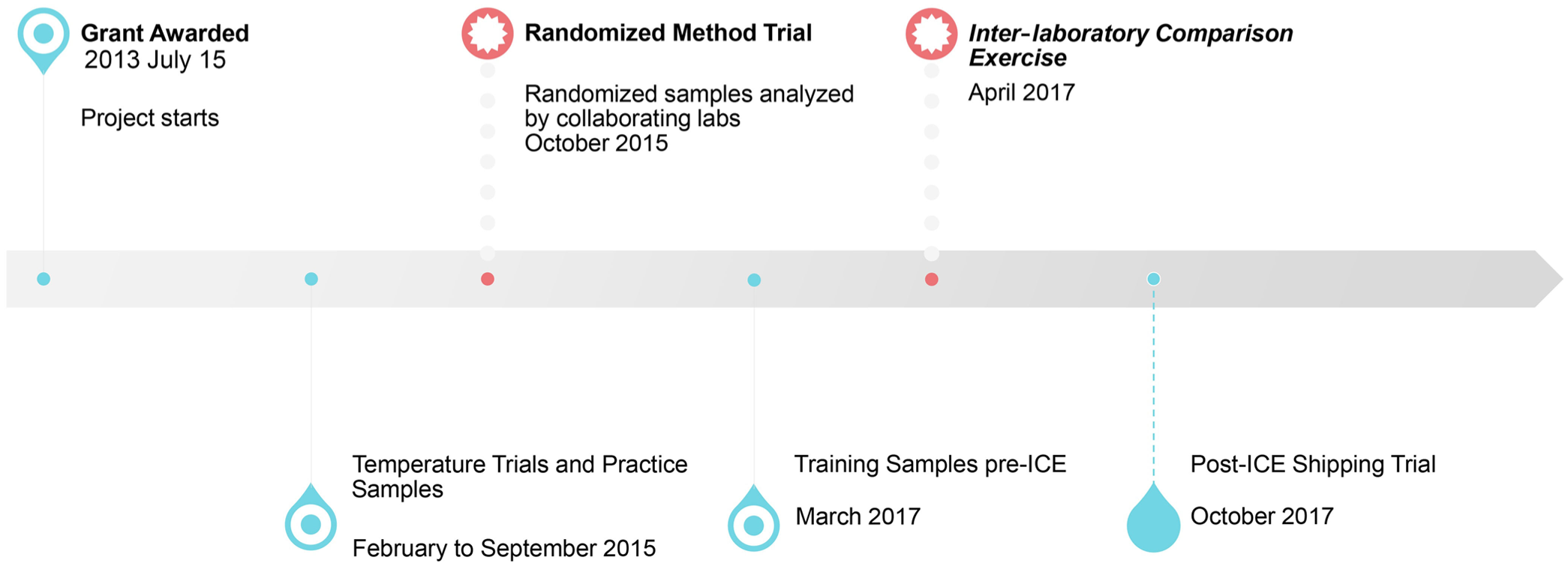

Four-year timeline for single-laboratory and multi-laboratory trials to evaluate the culture and molecular methods for the detection of Campylobacter jejuni in canine feces.
For the RMT conducted in 2015, fecal samples were divided into 2 ± 0.1-g aliquots in 50-mL conical tubes (VWR), packaged, and shipped on ice to the FDA Vet-LIRN Program Office (
Testing of shipping conditions
Before the 2015 RMT, the viability of C. jejuni ATCC 33560T during shipping was evaluated by shipping 12 inoculated canine fecal samples and 4 control fecal samples from Texas to the VPO, where they were held for 1 wk at 4°C and then shipped back to Texas for testing. Homogeneity and stability testing were performed before the ICEs. Homogeneity was tested 3 d after inoculation, and stability was tested at 2 and 9 d after inoculation. Before the shipping exercise, conditions inside the shipping containers were measured using different combinations of frozen and refrigerated ice packs (38-oz re-usable ice pack, Cryopack; 12-oz U-tek gel packs, Sonoco, Thermosafe) and frozen pellets (U-tek singles, Sonoco, Thermosafe), including combinations of 2 frozen ice packs with 4 refrigerated ice packs, 4 frozen with 2 refrigerated ice packs, and 4 frozen with 2 refrigerated ice packs with ~20 frozen pellets.
Microbiologic procedure for Campylobacter jejuni detection
The culture protocol with modifications can be found on protocols.io—open access repository platform DOI: dx.doi.org/10.17504/protocols.io.kxygx91wwg8j/v1. 28 For the 2015 RMT, samples were shipped to laboratories where 8 mL of 1× PBS was added to each 2-g sample to create a fecal slurry within 48 h of receipt of the samples. Using a sterile polyester swab, the fecal slurry was inoculated onto agar plates consisting of Campy-Cefex agar (BD) and modified charcoal cefoperazone deoxycholate agar (HiMedia Laboratories), supplemented with 8 mg/L cefoperazone, 4 mg/L amphotericin B, and 10 mg/L teicoplanin (mCCDA-CAT). Positive (C. jejuni ATCC 33291) and negative (E. coli ATCC 29522) controls were inoculated onto media for comparison. Plates were incubated at 42 ± 2°C for 48 h in a gas-tight canister or box under microaerophilic conditions (Mitsubishi Gas Chemical America). In addition to the direct culture of the fecal slurry described above, a 1-mL aliquot of the fecal slurry was added to 9 mL of Bolton broth (Hardy Diagnostics) and incubated at 42 ± 2°C for 48 h under microaerophilic conditions (enrichment culture). Following incubation, a swab was used to inoculate the enriched Bolton broth culture onto mCCDA-CAT and Campy-Cefex plates, which were incubated at 42 ± 2°C for 48 h under microaerophilic conditions, followed by examination for suspect colonies. Up to 4 suspect colonies from each sample were isolated on BAP, incubated again at 42 ± 2°C for 48 h under microaerophilic conditions, and tested for catalase and oxidase production. Isolates that were both catalase- and oxidase-positive were presumptively considered to be C. jejuni.
Molecular procedure for Campylobacter jejuni detection
For molecular testing, DNA was extracted from a 1-mL aliquot of the fecal slurry (PowerFecal DNA isolation kit; MO BIO Laboratories [Qiagen]) for direct detection of C. jejuni from the feces (direct PCR). For detection from enrichment (enrichment PCR), DNA was also isolated from a 1-mL aliquot of the Bolton enrichment broth (PowerFecal DNA isolation kit). rtPCR was performed (Applied Biosystems 7500 Fast real-time PCR system; ThermoFisher). All primers and probes were obtained from Sigma-Genosys. The rtPCR procedure was described previously. 29 Briefly, 2 μL of each DNA sample was combined with 6.75 μL of nuclease-free water (ThermoFisher), 12.5 μL of PCR master mix (TaqMan Fast universal PCR master mix; ThermoFisher), 1.25 μL (10 μM working stock; 0.5 μM final concentration in total reaction volume) of forward primer cpn60F (5′-TTAATGACGCGGTAAAAGTAACTATGG-3′), 1.25 μL (10 μM working stock; 0.5 μM final concentration) of cpn60 reverse primer (5′-TGCTTGGAGCACCAAAGCT-3′), and 1.25 μL (10 μM working stock; 0.5 μM final concentration) of the cpn60 probe (5′-[6FAM]CCAAGAGGACGCAATGT[BHQ1]-3′), for a reaction volume of 25 μL per sample. A positive control of 2 μL of C. jejuni ATCC 33560T DNA, a negative control of DNA from E. coli ATCC 25922, and a no-template negative control of 2 μL of nuclease-free water (ThermoFisher) were used. All reaction mixtures underwent quantification according to the following amplification cycle: 1 cycle at 95°C for 20 s, followed by 40 cycles of 1 s at 95°C and 20 s at 60°C. Based on preliminary experiments (data not shown), all samples yielding Ct values of 14–35 were considered positive; Ct values >35 but <38 were considered suspect positive, and Ct values ≥38 or samples in which no signal was detected were considered negative. 29
Management of multi-laboratory testing
Before the performance of all testing exercises (RMT, ICE, and shipping), practice samples were shipped to the participating laboratories to allow them to become familiar with the method. Following the testing of the practice samples, a second conference call was held to discuss the method and to address questions that arose during the practice testing. After this conference call, samples for the multi-laboratory randomized method test (
Following the M-RMT, in April 2017, the Vet-LIRN laboratories were offered the opportunity to test the method along with their own laboratory-developed methods in the Moffett ICE. Twelve samples were sent to 57 analysts in 28 laboratories. Afterward, a shipping exercise was conducted by Moffett in 2017 Oct with 12 analysts at 5 laboratories with an established record for successful detection of C. jejuni. Shipping conditions were tested by increasing the sample volume from 2 g to 8 g, using shipping containers with additional insulation, providing more detailed instructions with an emphasis on adequate vortexing during DNA extraction, and using temperature data loggers to actively record temperature throughout transport.
Inclusivity and exclusivity testing
To confirm the specificity of both the cpn60 primers and probes, inclusivity testing was performed using genomic DNA from C. jejuni ATCC 33291 and C. jejuni ATCC 33560T, as well as 20 clinical isolates that were confirmed as C. jejuni. We used matrix-assisted laser-desorption/ionization–time-of-flight mass spectrometry (MALDI-TOF MS; Bruker) and/or sequencing of a portion of the 16S ribosomal DNA subunit using the primers 515F (5′-GTGCCAGCAGCCGCGGTAA-3′) and 13R (5′-AGGCCCGGGAACGTATTCAC-3′). 35 Exclusivity testing was performed using genomic DNA from C. coli ATCC 33559T and 1 clinical isolate of C. coli; a clinical isolate of C. fetus, C. helveticus ATCC 51210, and C. upsaliensis ATCC 43954T; and 3 clinical isolates of C. upsaliensis. Additionally, the following strains were used for testing exclusivity: Actinobacillus equuli ATCC 19392T, Bacteroides fragilis ATCC 25285T, Bacteroides thetaiotamicron ATCC 29741, Bordetella bronchiseptica ATCC 19395T, Enterococcus faecalis ATCC 29212, Escherichia coli ATCC 25922, E. coli ATCC 35218, E. coli ATCC 51446, E. coli O157:H7 ATCC 43895, Klebsiella pneumoniae ATCC 13883T, a clinical isolate of Listeria ivanovii, a clinical isolate of L. monocytogenes, Mannheimia haemolytica ATCC 33396T, Pasteurella multocida ATCC 43137T, Proteus mirabilis ATCC 12453, Pseudomonas aeruginosa ATCC 27853, Rhodococcus equi ATCC 6939T, Salmonella enterica subsp. enterica serovar Braenderup ATCC BAA-664, Salmonella enterica subsp. enterica serovar Choleraesuis ATCC 13312, Salmonella enterica subsp. enterica serovar Dublin ATCC 15480, Salmonella enterica subsp. enterica serovar Typhimurium ATCC 14028, Staphylococcus aureus ATCC 25923, S. aureus ATCC 29213, S. aureus ATCC 43300, and Vibrio campbellii ATCC BAA-1117. It was not possible to follow the FDA guidelines 21 to include 50 isolates of the target organism for inclusivity (22 were obtained) and 30 strains of non-target organisms for exclusivity testing (33 isolates representing 26 organisms were used) because of the limited number of Campylobacter strains available; however, efforts were made to collect a variety of strains for testing.
Statistical analysis
Kappa statistics (κ) were calculated to assess agreement between testing methods, using a commercial statistical software program (SAS v.9.4; SAS Institute). Standard interpretive criteria were utilized. 19 Sensitivity (Se) was calculated by dividing the number of true-positives by the total number of positive samples multiplied by 100%. The specificity (Sp) was calculated by dividing the number of true-negatives by the total number of negative samples multiplied by 100%. The accuracy was calculated by adding the true-negative and true-positive samples, dividing the sum by the total number of tests, and multiplying the result by 100%.
Results
Using the cpn60 rtPCR, products were amplified from gDNA isolated from 22 strains of C. jejuni, including 2 commercial C. jejuni strains and 20 clinical isolates confirmed as C. jejuni using MALDI-TOF MS and sequencing of the 16S rDNA. No products were amplified from gDNA of any other bacteria tested, including C. coli, C. upsaliensis, C. fetus, and C. helveticus.
Before the randomized method test in 2015, the viability of the C. jejuni strain used for spiking samples during shipping was tested by shipping 12 spiked canine fecal samples and 4 control fecal samples from Texas to the VPO, where they were held for 1 wk at 4°C and then shipped back to Texas for testing. Four samples each were spiked with 2.9 × 104 cfu/g, 2.9 × 103 cfu/g, or 2.9 × 102 cfu/g C. jejuni. Upon their return to Texas, testing found that only 1 sample, spiked at 2.9 × 104 cfu/g, was culture positive on direct plating, but 11 of 12 spiked samples were positive on enrichment culture. The same 11 of 12 samples were positive by PCR of direct samples (Ct < 35). rtPCR testing using DNA from enrichment broths was positive for the same 11 of 12 samples, with Ct < 20.
In 2015 Oct, 7 FDA Vet-LIRN laboratories participated in a M-RMT to compare the sensitivity and specificity of culture-based and molecular detection of C. jejuni in canine feces. A total of 32 samples consisting of 8 samples at each of 4 bacterial concentrations (~3,200, 320, 32, and 0 cfu/g) were shipped to each laboratory and processed as described above. Culture of canine feces with and without enrichment in Bolton broth under microaerophilic conditions for 48 h (enrichment culture and direct culture, respectively) was tested and compared with rtPCR detection of cpn60 using DNA isolated directly from feces (direct PCR) or DNA isolated from the feces-inoculated Bolton enrichment broth following incubation under microaerophilic conditions for 48 h (enrichment PCR).
Data from 2 of the laboratories were excluded because of inconsistencies in the results of the positive and negative controls, as well as the samples. For the remaining 5 laboratories, the limit of detection (LOD) using PCR to detect C. jejuni from DNA isolated directly from feces without enrichment with Bolton broth was 320 cfu/g of feces, although some laboratories were able to detect C. jejuni at 32 cfu/g of feces (
Positive test results of culture and cpn60 real-time PCR for detection of Campylobacter jejuni from the October 2015 randomized method test.
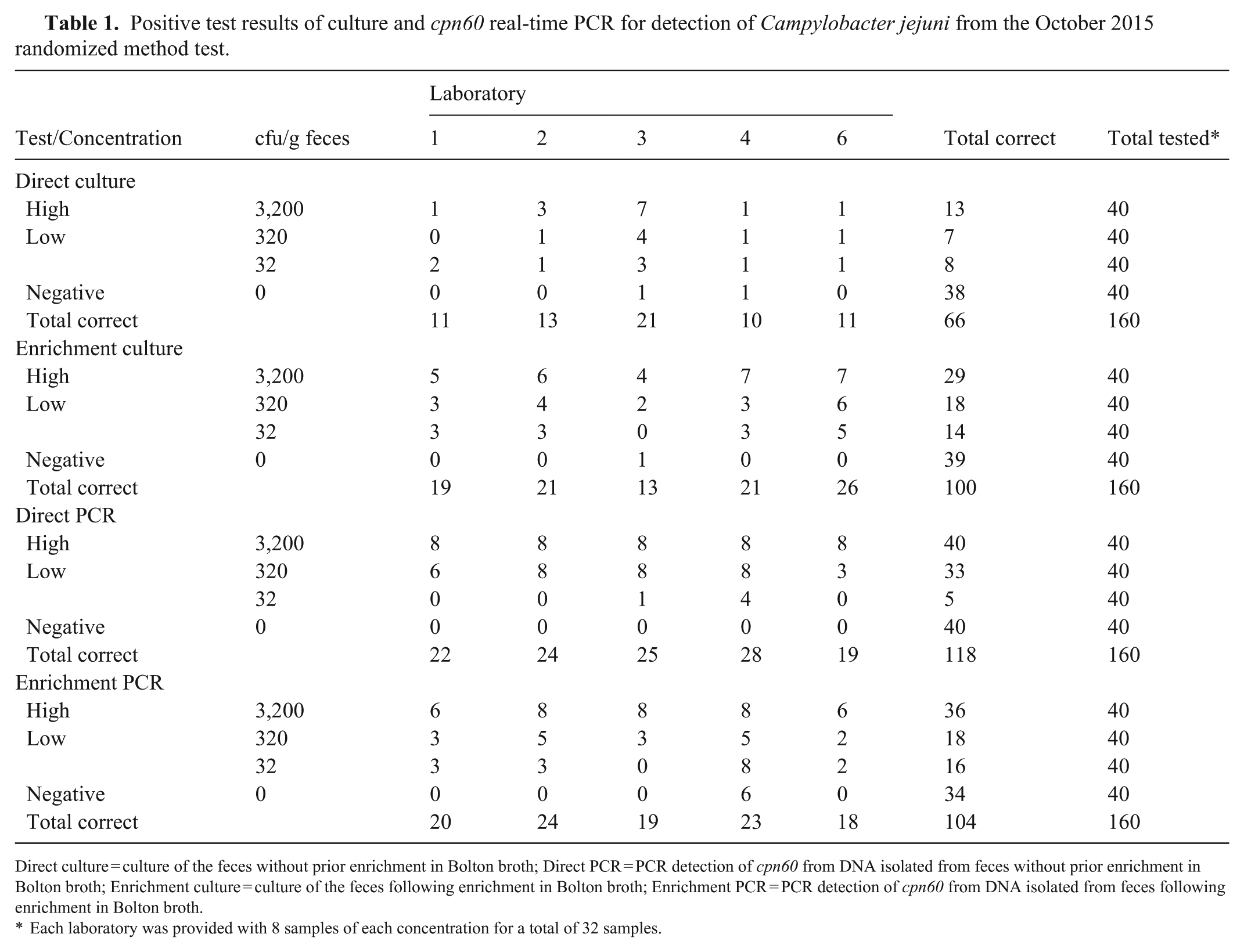
Direct culture = culture of the feces without prior enrichment in Bolton broth; Direct PCR = PCR detection of cpn60 from DNA isolated from feces without prior enrichment in Bolton broth; Enrichment culture = culture of the feces following enrichment in Bolton broth; Enrichment PCR = PCR detection of cpn60 from DNA isolated from feces following enrichment in Bolton broth.
Each laboratory was provided with 8 samples of each concentration for a total of 32 samples.
Summary of sensitivity, specificity, and accuracy rates of the direct PCR for detection of Campylobacter jejuni cpn60 in 5 independent multi-laboratory testing exercises.
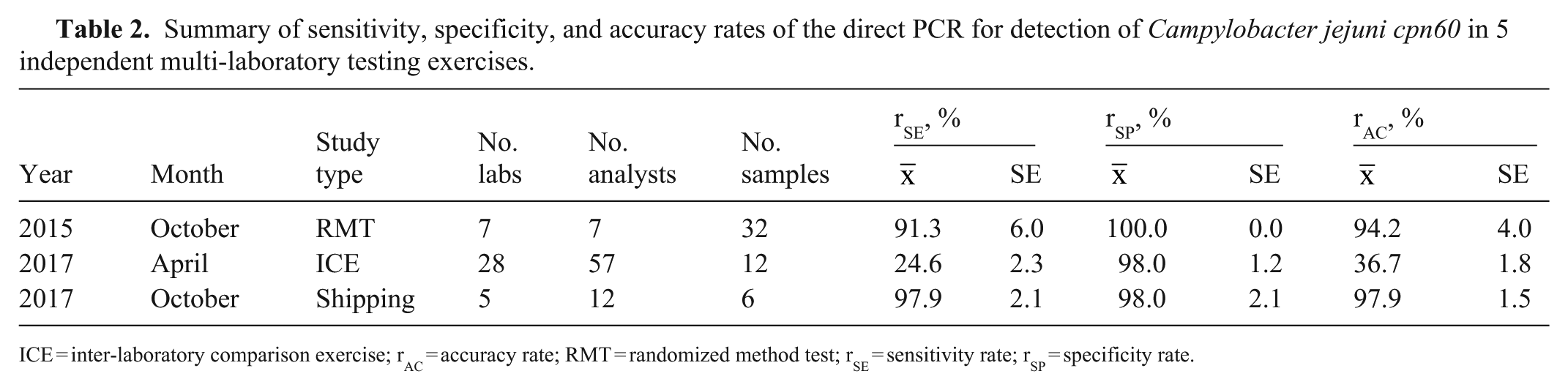
ICE = inter-laboratory comparison exercise; rAC = accuracy rate; RMT = randomized method test; rSE = sensitivity rate; rSP = specificity rate.
In 2017 Apr, Moffett offered the Vet-LIRN laboratories the opportunity to test the method along with their own laboratory-developed methods in an ICE. Twelve samples were sent to 57 analysts. Of the 32 analysts who tested the cpn60 direct PCR method, 14 did not detect C. jejuni in any sample, and 11 of the 18 analysts who detected C. jejuni in at least 1 sample failed to detect the organism in at least 7 of the 10 positive samples. Analysis of the temperature indicator data indicated that 79% of the packages reached a temperature of 5°C or greater during transport, possibly causing sample instability. Many false-negative results were detected, and the sensitivity of the test was reduced to 24.6% (Table 2). As a result, shipping methods were re-evaluated.
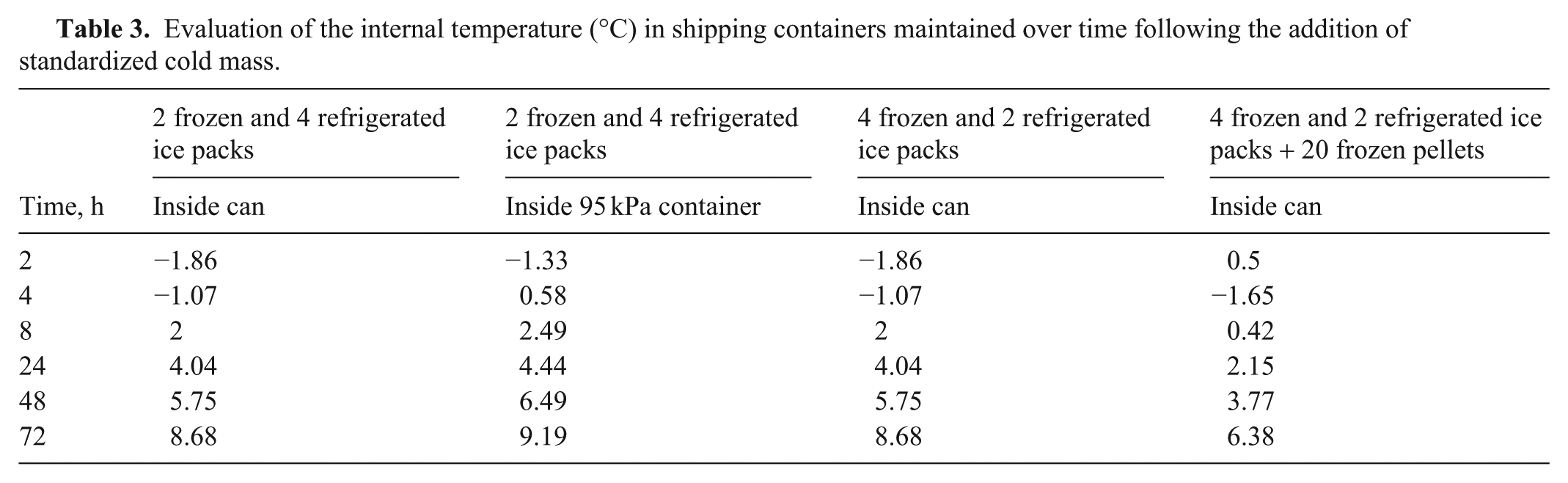
A shipping exercise was conducted by Moffett in October 2017 with 12 analysts at 5 laboratories with an established record for successful detection of C. jejuni. During the shipping exercise, improvements to the study included increasing sample volume from 2 g to 8 g, using shipping containers with additional insulation, providing more detailed instructions with an emphasis on adequate vortexing during DNA extraction, and using temperature data loggers to actively record temperature throughout transport. Before shipping samples to laboratories, the internal temperature of the shipping container was measured over 3 d (
Evaluation of the internal temperature (°C) in shipping containers maintained over time following the addition of standardized cold mass.
Discussion
In our M-RMT, we experienced the usual challenges in the culture and detection of C. jejuni that other laboratories have experienced.20,24 We found that false-negatives were common in culture-based methods, even with enrichment culture. One laboratory had a high rate of false-positives with the enrichment culture. Additionally, 2 laboratories were unable to provide reliable results, and their data were excluded from analysis. Ultimately, molecular detection of DNA isolated directly from feces provided greater sensitivity than culture or molecular detection from enrichment samples, without compromising specificity.
Shipping conditions were a factor in the 2017 Apr ICE. Temperatures in 79% of the shipped containers reached >5°C. This affected the ability of the analysts to detect C. jejuni in samples. Sensitivity of that trial assay was 24.6%, compared with 97.9% in the subsequent shipping exercise (conducted in 2017 Oct, after successfully establishing a shipping method to consistently maintain samples at <4°C for 48 h).
It was not surprising that detection of C. jejuni was lower in samples inoculated with <100 cfu/g of feces. This low inoculation level was used to gather information on the fractional recovery level of the final samples. The ideal inoculation level for fractional recovery is the level at which half of the inoculated samples test positive and the other half test negative, and fractional recovery should be ~50 ± 25% of the total number of replicates in the sample set. 21 In the 2015 M-RMT, 8 samples at the 32 cfu/g concentration were tested by 5 laboratories; a total of 40 samples were tested. This level was determined to be below the level of fractional recovery, as none of the methods used detected half of the samples at this inoculation level. The growth and recovery of C. jejuni by participating laboratories is part of the challenge in using this organism in performance evaluation studies.20,24
An additional potential limitation in molecular detection of pathogens in patient specimens is the possible presence of PCR inhibitors in feces and body fluids. To control for this issue, laboratories may use either a universal inhibition-monitoring strategy or include internal controls specific to each pathogen-detection assay. This process has been reviewed in detail. 45 We did not use an endogenous internal PCR control in our study; however, the status of all samples was known in the M-RMT, ICE, and shipping exercise. Additionally, in the M-RMT, 3 of the 5 laboratories detected C. jejuni in 8 of 8 samples in the high- and low-concentration samples, suggesting that PCR inhibition was not a significant issue. Subsequent testing has confirmed that an internal endogenous DNA extraction and PCR control can be multiplexed with rtPCR for C. jejuni testing and is part of a subsequent study (data not shown). PCR detection of C. jejuni should be followed up with culture, susceptibility testing, and whole-genome sequencing of recovered isolates for outbreak monitoring.
Overall, the detection of C. jejuni DNA directly from canine feces was successful at a LOD of 320 cfu/g with the cpn60 rtPCR assay. Isolation of DNA directly from feces rather than from enrichment broth offered the advantage of fewer opportunities for cross-contamination during testing and decreased time between specimen collection and test completion. The tested enrichment protocol required an additional 48 h. During an outbreak situation, this would substantially increase the time needed to identify the etiologic agent. Based on the results of our study, shipping conditions should be considered when preparing ICE and routine specimen collection and submission for the detection of C. jejuni. C. jejuni inoculated at 320 cfu/g remains stable and detectable by rtPCR for 48 h, but detection may be hampered if testing is delayed. Detection of C. jejuni directly from canine feces using molecular tests is more sensitive than culture and offers a means of rapid identification of positive fecal samples.
Supplemental Material
sj-xlsx-1-vdi-10.1177_10406387261434179 – Supplemental material for Comparison of PCR and culture for detection of Campylobacter jejuni in canine feces
Supplemental material, sj-xlsx-1-vdi-10.1177_10406387261434179 for Comparison of PCR and culture for detection of Campylobacter jejuni in canine feces by Sara D. Lawhon, Kevin J. Cummings, Jing Wu, Ching Yuan Yang, Nicholas Whipple, Melanie Prarat Koscielny, Samantha Lindemann, Shannon Kiener, Matthew Kmet, Ravinder Reddy, Laura B. Goodman, Rebecca J. Franklin-Guild, Kelly Sams, Brittany Cronk, Jing Cui, Kerri Lawrence, Qirui Zhang, Yan Zhang, Steven Ensley, Karen Olsen, John Tomaszewski, Zhenyu Shen, Shuping Zhang, Sarmila Dasgupta, Shannon Mann, Amar Patil, Siddhartha Thakur, Shelley C. Rankin, Laura Ruesch, Nuria Garcia-Fernandez, Jake Guag, Sarah M. Nemser and Renate Reimschuessel in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank the laboratories and individual analysts who performed the work and patiently provided insights that improved the assays throughout the course of this effort. Of particular note, we thank the members of the CVM Vet-LIRN and Moffett PT Laboratory IIT/IFSH for organizing and preparing test samples for testing performed in 2017–2019. For all of their work participating in the ICE, we thank Zeinab Helal and Risatti Guillermo of the Connecticut Veterinary Medical Diagnostic Laboratory; Christina Weller and Kristy L. Pabilonia of the Colorado State University Veterinary Diagnostic Laboratory; Wanda Tirado, Alex Nemethy, and David Simon of the Bronson Animal Disease Diagnostic Laboratory; Ingrid Fernandez and Paula Bartlett of the University of Georgia, Athens Veterinary Diagnostic Laboratory; Dr. Lester Perez, Evette Vlach, and Vanessa DeShambo of the University of Illinois, College of Veterinary Medicine, Veterinary Diagnostic Laboratory; Tracie Jenkins and Dr. Shri Singh of the Murray State University Breathitt Veterinary Center; Tarra Harden and Keith Strother of the Louisiana Animal Disease Diagnostic Laboratory; Rinosh Mani and Niesa Kettler of the Michigan State University Veterinary Diagnostic Laboratory; members of the Veterinary Diagnostic Laboratory at the University of Minnesota; Rebecca Mackey, Lifang Yan and Candy Zhang of the Mississippi Veterinary Research and Diagnostic Laboratory; Solomon Odemuiwa of the University of Missouri Veterinary Medical Diagnostic Laboratory; members of the New Jersey Department of Agriculture, Animal Health Diagnostic Laboratory; Weihua Wang and Renee Anderson of the Animal Health Diagnostic Center, New York State Veterinary Diagnostic Laboratory, Cornell University; Aaron Briley, Melissa W. Byrd, and Dr. Megan Jacob of the North Carolina State College of Veterinary Medicine, Microbiology and Molecular Diagnostics Laboratory; Dominika Jurkovic of the Ohio Animal Disease Diagnostic Lab; Robin Madden and Dr. Akhilesh Ramachandran of the Oklahoma Animal Disease Diagnostic Laboratory; Li Fang, Donna Krouse, and Nagaraja Thirumalapura of the Pennsylvania Veterinary Laboratory; Laura Chen and Sylvia Weber of the Avian Health & Food Safety Laboratory of the Washington Animal Disease Diagnostic Laboratory; and Molly MacNab, Karen Wallace, and Jennifer Cooper of the Wisconsin Veterinary Diagnostic Laboratory.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our work was supported by U.S. FDA grants U18FD005013, U18FD006446, and U18FD006664. N. Whipple was supported by an NIH grant to A. Kier (5T35OD010991). Diagnostic materials (canine feces) were provided by Dwayne Schrunk and Dr. Steve Ensley at the Veterinary Diagnostic Laboratory, Iowa State University (Ames, IA, USA) under contract HHSF223201410256A for diagnostic sample collection and subject matter expertise for Vet-LIRN Proficiency Exercises.
ORCID iDs
Supplemental Material
Supplemental material for this article is available online.
