Abstract
We developed a real-time (rt)PCR assay based on TaqMan probe technology for the specific detection of canine adenovirus 1 (CAdV-1). The assay is able to detect three 50% tissue culture infectious dose/mL in CAdV-1–containing cell culture supernatant. Viral genomes were not amplified of canine adenovirus 2 or of several bovine, porcine, and avian adenoviruses. In silico analysis provided no indication of amplification of other heterologous genomes. The sensitivity of the real-time assay exceeded that of a conventional gel-based CAdV-1 PCR by a factor of 100. Following the integration of the novel PCR into the Hessian wildlife-monitoring program, CAdV-1 DNA was detected in none of the tested raccoons (n = 48) but in 11 of 97 foxes.
Species Canine adenovirus 1 (CAdV-1; family Adenoviridae, genus Mastadenovirus) is the causative agent of infectious canine hepatitis. In contrast to canine adenovirus 2, which is mainly associated with infectious tracheobronchitis in dogs, 5 CAdV-1 is characterized by much higher pathogenicity because of differing tissue tropism. 5 Infection with CAdV-1 and -2 has been documented in dogs,3,7 foxes,2,6,12,17 and wolves, 16 among other species. In raccoons, 14 exposure to canine adenoviruses has been proven based on serologic studies, which do not allow for differentiation of antibodies against CadV-1 and -2.
Infection with CAdV-1 causes febrile disease accompanied by necrosis in various internal organs, petechial or ecchymotic hemorrhage, and neurologic disorders. In young, unvaccinated animals, infection may be peracute with sudden death and few clinical signs. In adult canids possessing CAdV-specific antibodies at the time of infection, it may also take a subclinical course, potentially leading to chronic viral shedding. 9 In most parts of Europe, the disease is now rare in dogs thanks to extensive vaccination of owned dogs against CAdV using CAdV-2 live vaccines that confer protection against both CAdV-1 and -2. However, in settings where vaccination of puppies is neglected, infections still occur. 8
For detection of CAdV-1 infection, different protocols have been developed for PCR.4,7,15,15 Most of those protocols are based on analysis of the amplification reaction by agarose gel electrophoresis. In order to simplify investigations dealing with the monitoring of CAdV-1 in wild animal species, we developed a real-time (rt)PCR, the validation of which we report herein. We present data from Hessian wildlife monitoring during which foxes (Vulpes vulpes) and raccoons (Procyon lotor) hunted in Hesse during autumn 2015 and winter 2015–2016 were examined postmortem and tested for CAdV-1.
Animals were collected for the German national rabies monitoring program in Hesse. They were killed by hunters after showing clinical signs of illness or were found dead. Organ samples from 97 foxes and 48 raccoons of various ages were screened with the newly developed CadV-1 rtPCR method between August 2015 and January 2016. The age of the animals was estimated at the time of autopsy based on tooth status and outer appearance. DNA was extracted from a pool of tissue containing liver, lung, and brain of each animal; 250 mg of this organ pool were homogenized in 1 mL of phosphate-buffered saline (PBS; FastPrep-24, MP Biomedicals, Santa Ana, CA) and centrifuged for 1 min at 20,000 × g; 200 µL of the resulting supernatant were transferred to a 2-mL V-bottom screw-cap vessel and DNA was extracted automatically using silica membrane–containing spin columns (QIAcube in combination with QIAamp cador pathogen mini kit, Qiagen, Hilden, Germany), according to the manufacturer’s instructions. Lysis was done using 20 µL of proteinase K per sample, and incubation was performed at room temperature for 15 min. DNA was eluted in a volume of 100 µL per sample. Sample material and DNA were stored at −20°C until use.
New primers and a hydrolysis probe were designed to specifically amplify a 170-bp, CAdV-1–specific fragment within the E3 region of the adenovirus genome (Table 1). To identify the appropriate location for CAdV-1–specific primers and probe, published sequences for CAdV-1-E3 and CAdV-2-E3 were retrieved from GenBank and their consensus sequences were compared. The CAdV-1-E3–specific part of the sequence was analyzed, and candidate primer and probe sequences were chosen with the help of OligoCalc (https://goo.gl/Q0gAMZ). Specificity was verified in silico by BLAST analysis of the oligonucleotide sequences. 1 Information on primers and probes of the beta-actin internal control system and on primers for a PCR capable of simultaneously detecting CAdV-1 and -2 have been published elsewhere.13,18 Animals examined in the context of the monitoring program were further tested for canine distemper virus (CDV) by a real-time reverse transcription (RT)-PCR published earlier. 11 All oligonucleotides were synthesized by TIB Molbiol Syntheselabor (Berlin, Germany).
Oligonucleotide sequences used in the current study.*
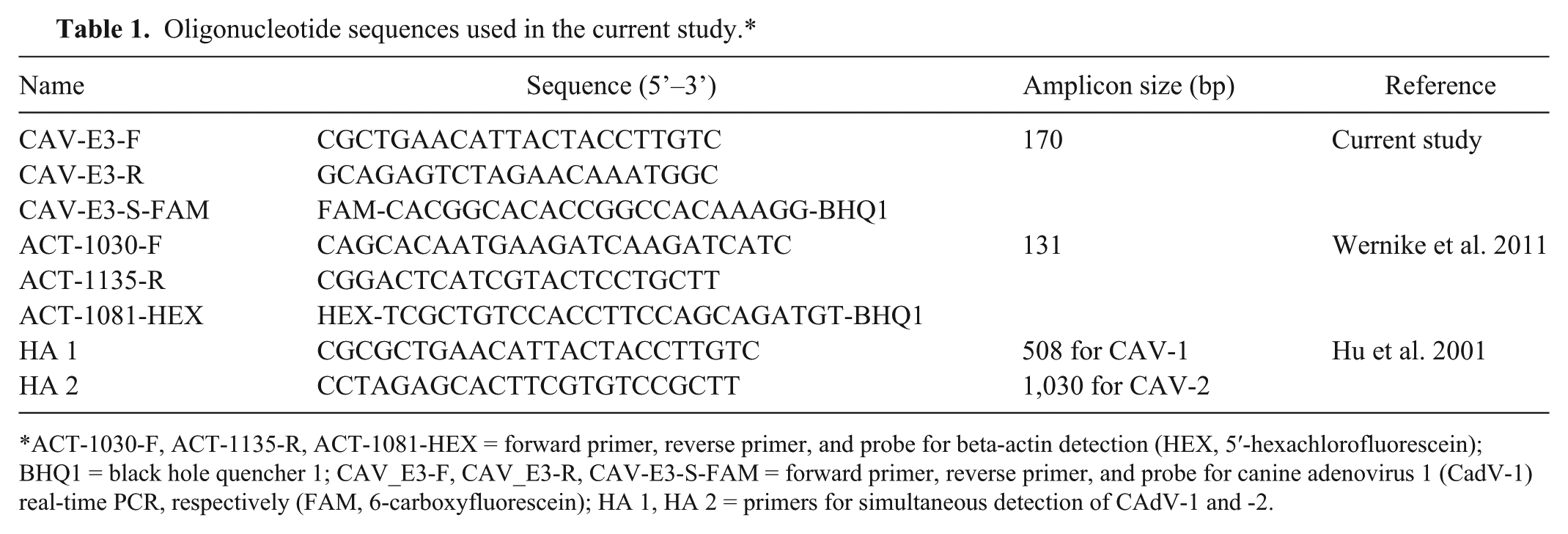

ACT-1030-F, ACT-1135-R, ACT-1081-HEX = forward primer, reverse primer, and probe for beta-actin detection (HEX, 5′-hexachlorofluorescein); BHQ1 = black hole quencher 1; CAV_E3-F, CAV_E3-R, CAV-E3-S-FAM = forward primer, reverse primer, and probe for canine adenovirus 1 (CadV-1) real-time PCR, respectively (FAM, 6-carboxyfluorescein); HA 1, HA 2 = primers for simultaneous detection of CAdV-1 and -2.
For in vitro evaluation of candidate primers and probes, cell culture supernatant was used, containing CAdV-1, CAdV-2, bovine adenovirus 2 and 3 (BoAdV-2, -3), porcine adenovirus 1, 2, and 3 (PoAdV-1, -2, -3), and pigeon adenovirus 1 (PiAdV-1), respectively (Supplementary Table 1). For each type of adenovirus listed above, 1 strain was available. Strains of CAdV-1, BoAdV-2 and -3, PoAdV-1, -2, and -3, and PiAdV-1 were provided from the collection of virus strains of the Friedrich-Loeffler-Institute (Greifswald, Germany). A field isolate of CAdV-2 from a Hessian dog was kindly provided by Dr. Werner Herbst from the Institute of Veterinary Hygiene and Infectious Diseases of the Justus-Liebig-Universität (Gießen, Germany). The isolates of CAdV-1 and -2 were propagated on Vero cells in our laboratory (cell line obtained from the Collection of Cell Lines in Veterinary Medicine at the Friedrich-Loeffler-Institute, Federal Research Institute of Animal Health, Insel Riems, Germany).
The titers of the resulting virus stocks were determined to be 2.4 × 108 (CAdV-1) and 1.8 × 105 (CAdV-2) 50% tissue culture infectious dose (TCID50)/mL, respectively. Serial 10-fold dilutions of CAdV-2 stock solution were prepared in sterile cell culture minimum essential medium, and used as positive controls in the combined CAdV-1/CAdV-2 PCR in single replicates. Serial 10-fold dilutions of the CAdV-1 cell culture strain were prepared in tissue homogenate of a CAdV-negative fox to simulate CAdV-1 detection from tissue samples, and used as positive controls for setup and optimization of the CAdV-1 rtPCR. Template DNA from each dilution was tested in triplicate to calculate PCR efficiency for detection of CAdV-1 alone or in combination with beta-actin detection (Mx3005P QPCR cycler and MxPro manager software, Agilent, Santa Clara, CA). Single replicates of these dilutions were tested in the combined CAdV-1/CAdV-2 PCR in order to compare the detection limit of conventional and rtPCR. DNA extracted from cell culture supernatant containing BoAdV-2 and -3, PoAdV-1, -2, and -3, and PiAdV-1 was tested with the duplex CAdV-1/beta-actin rtPCR in duplicate.
The QuantiTect multiplex PCR NoROX kit (Qiagen) was used for the setup of the duplex rtPCR for combined detection of CAdV-1 and beta-actin. The final protocol for the assay was based on 25-µL total reaction volume. Per reaction, the master mix consisted of 4.25 µL of nuclease-free water, 12.5 µL of 2× QuantiTect multiplex PCR NO ROX mix (Qiagen), 1.25 µL of a CAdV-1–specific primer–probe mix, and 2 µL of a beta-actin–specific primer–probe mix. The CAdV-1–specific primer–probe mix consisted of 1× Tris-EDTA (TE) buffer (pH 8) containing 4 µM CAV_E3-F and CAV_E3-R each and 2 µM CAV_E3-S-FAM. The beta-actin–specific primer–probe mix consisted of 1× TE buffer (pH 8) containing 2.5 µM ACT2-1030-F and ACT-1135-R each and 1.25 µM ACT-1081-HEX. Finally, 5 µL of template DNA were added. The rtPCR was carried out applying a thermal profile with an initial activation step of 15 min at 95°C followed by 45 cycles consisting of a denaturing step (60 s at 94°C) and a combined annealing and elongation step (60 s at 60°C; Mx3005P QPCR cycler and MxPro manager software, Agilent).
For simultaneous detection of CAdV-1 and -2, reactions of 25-µL total volume each were prepared containing 13.1 µL of nuclease-free water, 0.8 µL of dimethyl sulfoxide (AppliChem, Darmstadt, Germany), 2 µL of 10× polymerase buffer (DreamTaq, Thermo Fisher Scientific, Schwerte, Germany) including 20 mM MgCl2, 0.8 µL of each primer (final concentration 0.5 µM), 0.5 µL of DNA polymerase (final concentration 2.5 U per reaction; DreamTaq, Thermo Fisher Scientific), 2 µL of dNTP mix (final concentration 100 µM per nucleotide), and 5 µL of template DNA. PCR was carried out applying the following temperature profile: initial activation for 3 min at 95°C followed by 35 cycles of 30 s at 95°C, 30 s at 58°C, and 60 s at 72°C. Additionally, a final elongation step of 10 min at 72°C was included (Cycler T3000, Biometra, Göttingen, Germany). PCR products were analyzed via electrophoresis in 1.5% agarose gel.
BLAST analysis of CAdV-1–specific primer and probe sequences gave no indication of nonspecific amplification of unwanted targets (data not shown). Moreover, the oligonucleotides matched all published sequences of the CAdV-1-E3 region, except 3 (JX908001.1, JX908001.1, S38212.1; Fig. 1), with S38212.1 being the sequence of a CAdV-2 isolate in fact (Supplementary Fig. 1). For accession JX908001.1, detection in the rtPCR can only be expected with reduced sensitivity; 4 mismatches in positions 19–22 of the forward primer were predicted, caused by base substitutions in the viral sequence (TGTC → AAAA); perfect matches were achieved with sequences of probe and reverse primer (Fig. 1). For accession EF090910.1 (an Indian CAdV-1 strain with unknown animal origin), amplification by the oligonucleotides presented herein cannot be expected; merely partial complementarity with the probe sequence was noted. Furthermore, the forward primer binding site could not be verified because of insufficient length of the published sequence. Solely, the reverse primer sequence was found to be identical to its binding site (Fig. 1). Comparison to other sequences of CadV-1-E3 demonstrates that the first 45 nucleotides of this sequence significantly differ from other CAdV-1 strains, which prevents matching with the probe sequence. However, this seems to be a unique feature of this specific strain, as other Indian strains, for which sequences have been published (EF057101.1, GQ340423.1.), do not contain those mutations.

Alignment of canine adenovirus 1 (CadV-1)–specific real-time PCR oligonucleotides with CAdV-1 sequences from GenBank. The binding sites are highlighted by large frames. Variable bases are marked with small frames. Variable bases within primer or probe binding sites are marked by small, gray frames.
Some E3 sequences for which the forward primer binding site is not included in the published sequences (GQ340423.1, KF676980.1, KF676977.1, JX416840.1, JX416839.1, JX416838.1, KC577558.1, KJ451612.1, KP670422.1, KP670423.1, KP670424.1) were generated with the help of a PCR published previously, 13 and the forward primer of that PCR is almost identical to the one of our rtPCR (Table 1; Fig. 1). In general, comparison of available CAdV-1-E3 sequences demonstrates a high genetic stability for this genomic region during the past 20 y, which has also been noted in earlier studies. 2
The efficiency of CAdV-1 detection was determined to be 103% and 90% for the singleplex and duplex (including beta-actin detection) real-time assay, respectively. Absence of cross-reaction with bovine, porcine, or avian adenoviruses was confirmed by testing DNA extracts in vitro. Most importantly, an isolate of CAdV-2 with a titer of 1.8 × 105 TCID50/mL was not amplified by the CAdV-1–specific primers, which confirmed the results of in silico analysis (Supplementary Table 1). The limit of detection for both the singleplex and the duplex assay was calculated to be 3 TCID50/mL; fluorescence signals for 3 of 3 replicates were detected for dilutions 10-1 to 10-8 of cell culture supernatant containing CAdV-1 at a titer of 2.4 × 108 TCID50/mL (Supplementary Table 1). With the conventional, gel-based PCR assay for simultaneous detection of CAdV-1 and -2, positive results for CAdV-1 were only achieved for dilutions 10-1 to 10-6, with a very low amount of PCR product detectable in dilution 10-6. This corresponds to a calculated detection limit of 240 TCID50/mL (Fig. 2). In comparison, the assay gave positive results for CAdV-2 up to a dilution of 10-4, corresponding to a calculated detection limit of 18 TCID50/mL.
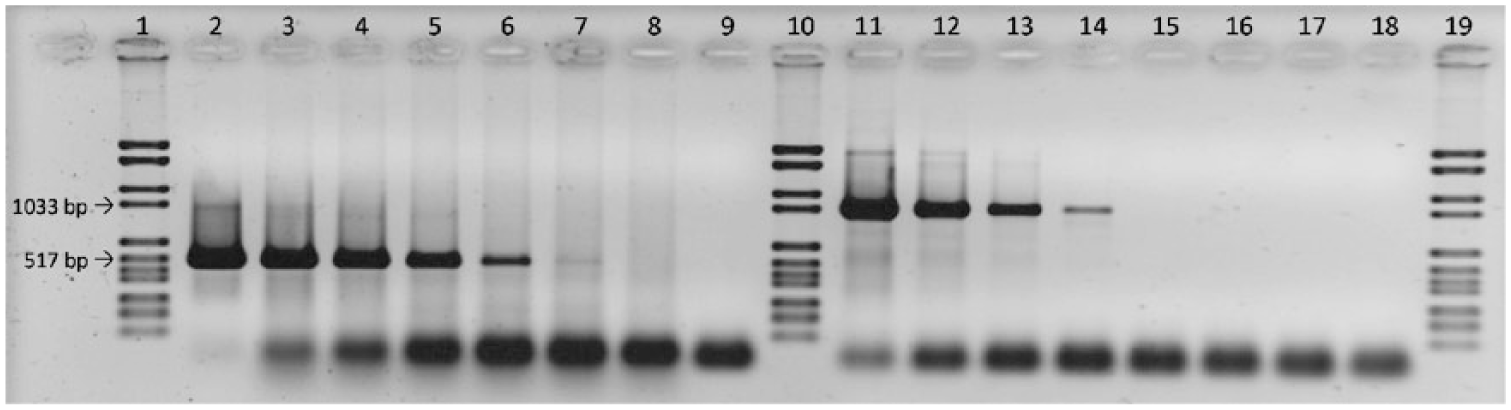
Comparative PCR analysis of 2 serial 10-fold dilutions of DNA prepared from canine adenovirus 1 (CAdV-1)– and CAdV-2–infected tissue culture supernatant, respectively. A 508-bp fragment is amplified for CAdV-1 and a 1,030-bp fragment for CAdV-2. Arrows indicate the size of the respective marker bands. Lanes 1, 10, and 19: marker; lanes 9 and 18: no template control; lanes 2–8: tissue culture supernatant containing CAdV-1, dilutions 10-1 to 10-7; lanes 11–17: tissue culture supernatant containing CAdV-2, dilutions 10-1 to 10-7.
None of the 48 raccoons tested was found positive for CAdV-1, but 22 (46%) tested positive for CDV. CAdV-1 DNA could be detected in 11 of 97 foxes (11%; Table 2). A total of 34 foxes (35%) tested positive for CDV, among which were 2 of the CAdV-1–positive animals.
Findings in wild foxes in the current study.*
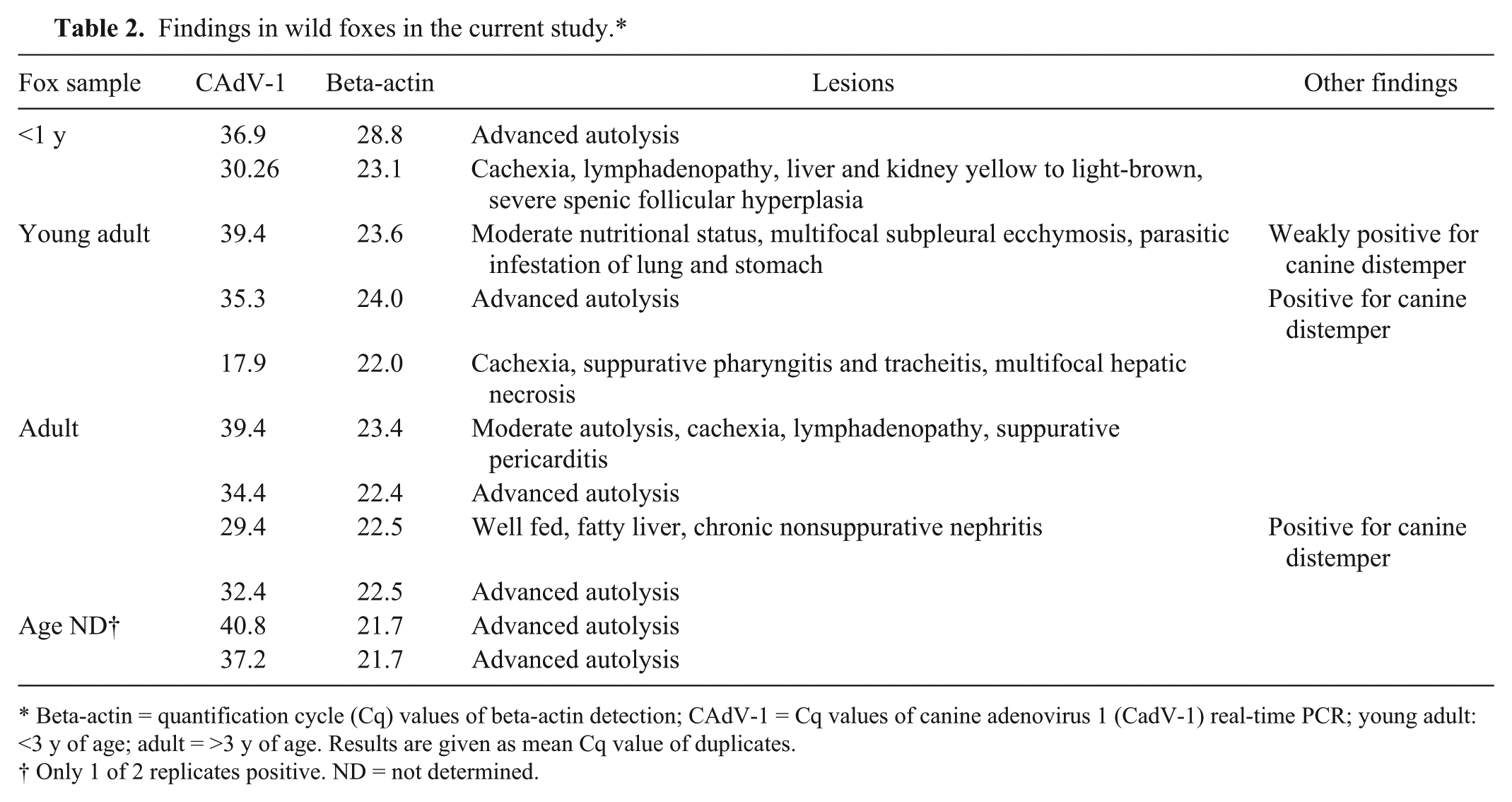
Beta-actin = quantification cycle (Cq) values of beta-actin detection; CAdV-1 = Cq values of canine adenovirus 1 (CadV-1) real-time PCR; young adult: <3 y of age; adult = >3 y of age. Results are given as mean Cq value of duplicates.
Only 1 of 2 replicates positive. ND = not determined.
Autopsy was informative in 5 of the foxes that tested CAdV-1 positive; findings were generalized lymphadenopathy, yellow to light-brown liver and kidney (suspicion of necrosis), splenic follicular hyperplasia, multifocal subpleural ecchymoses, chronic nonsuppurative nephritis, or multifocal hepatic necrosis (Table 2). Two of the 5 foxes reacted only weakly in the rtPCR (quantification cycle [Cq]: 39.4); the remaining 3 animals were clearly positive, with one of them carrying a very high viral load (Cq: 17.9). For this latter fox, histologic examination of liver also confirmed the PCR result; hepatocellular necrosis and characteristic intranuclear inclusion bodies could be found (Fig. 3). For 6 of the CAdV-1–positive foxes, postmortem examination was inconclusive because of advanced autolysis of the carcasses; for 4 of these 6 foxes, rtPCR results unambiguously indicated presence of viral DNA (mean Cq: 32.4, 34.4, 35.3, and 36.9, respectively). For the remaining 2 animals, only 1 of 2 replicates each gave a signal during rtPCR (Cq: 40.8 and 37.2, respectively).
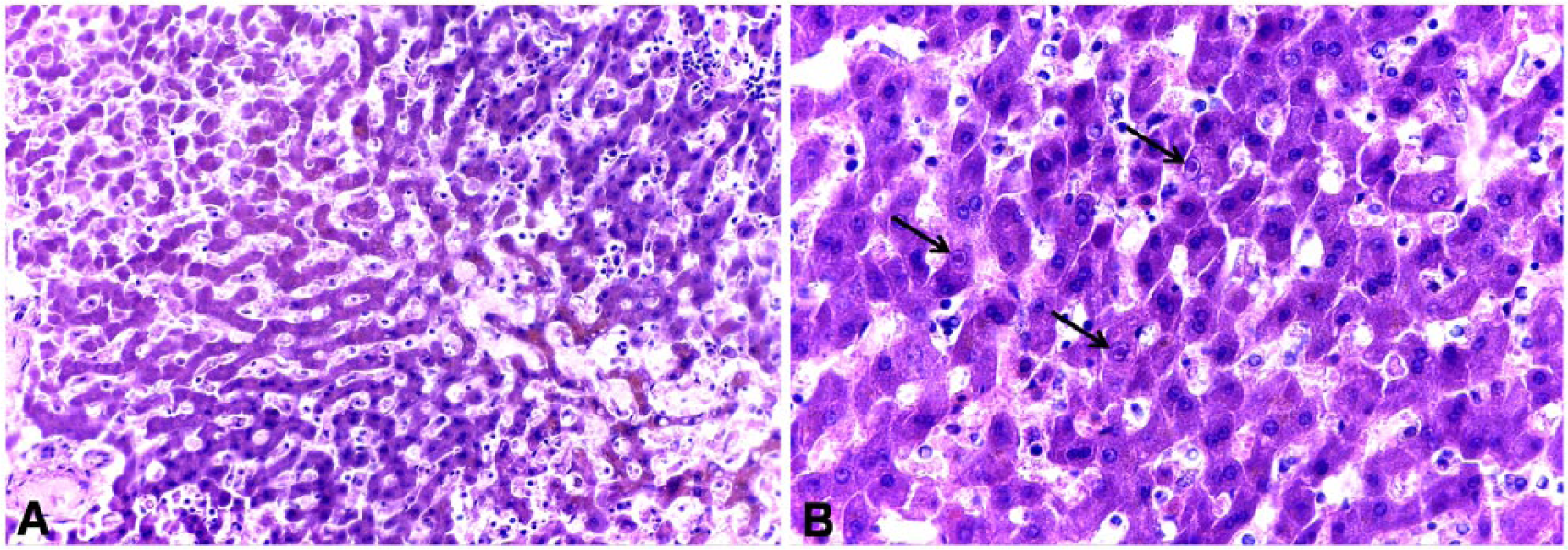
Histologic lesions in the liver of a canine adenovirus 1 (CAdV-1)–infected fox.
With this newly developed rtPCR assay, sensitivity of CAdV-1 detection is enhanced by a factor of 100 compared to the conventional gel-based PCR. Additionally, rtPCR offers the possibility of objectively quantifying genome loads in samples with the help of plasmid standards for CAdV-1. Linearity, efficiency, and sensitivity of our real-time assay are equivalent to another TaqMan rtPCR for detection of CAdV-1 and -2, which was published recently and is based on the sequence of the CAdV-hexon gene. 10 The E3-based rtPCR described herein is an alternative method for epidemiologic screenings, if detection of the less pathogenic CadV-2 is negligible.
The results of our wildlife monitoring demonstrate the presence of CAdV-1 in the Hessian fox population. However, correlation of pathologic findings with the detection of genomic material of the pathogen has proven to be difficult. This is influenced by several factors. First, postmortem examination is impaired by the degree of decomposition of the carcasses in many cases. Second, wild animals may show signs of multiple diseases or nonspecific signs of disease (lymphadenopathy, splenic follicular hyperplasia). Moreover, most of the PCR-positive animals in our study seemed to carry rather low viral loads. This would be compatible with chronic or subclinical infection that would in turn not be recognizable during macroscopic postmortem examination. Also, the degree of autolysis of the carcasses or variability of viral sequences could add to the observation of low viral loads in samples. Nonetheless, some of the pathologic changes observed in CAdV-1–infected foxes in our study (subpleural ecchymoses, hepatic and renal necrosis) are compatible with a diagnosis of infectious canine hepatitis and, in one case with a very high viral load, PCR detection was confirmed by histopathology (Fig. 3).
Overall, it cannot be excluded that certain field strains might be missed by our assay. To enable adoption, additional sequence data from currently circulating virus strains would be necessary—ideally on a global scale. Nonetheless, applicability of the newly designed oligonucleotide sequences for detection purposes can be judged as satisfactory.
Footnotes
Acknowledgements
We thank Johanna Schmidt, Björn Hebel, Simon Schneider, and Marie-Luise Sonneborn at the Hessian State Laboratory for their excellent technical assistance.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The Hessian State Laboratory (Hessisches Landeslabor) is supported by the Hessian Ministry for the Environment, Climate Protection, Agriculture and Consumer Protection (HMUKLV). The funder had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
