Abstract
We identified a novel herpesvirus in 2 deceased captive blue penguins (Eudyptula minor). Moderate-to-severe myocardiocyte atrophy and necrosis, and eosinophilic intranuclear inclusion bodies (INIBs), were seen in myocardiocytes in one bird; reticuloendothelial (RE) cell INIBs and multifocal RE cell necrosis were seen in both birds. The histologic findings were suggestive of viral infection. A herpesvirus PCR assay was positive in myocardial tissue from the bird with myocardial degeneration and in splenic tissue from both birds. Sequencing and phylogenetic analysis showed that the virus, accessioned as spheniscid alphaherpesvirus 2 (SpAHV2), groups within the Alphaherpesvirinae subfamily and forms a unique branch point in a subclade containing members of the Mardivirus, Simplexvirus, and Varicellovirus genera. Herpesvirus screening of tissues from 8 additional blue penguin postmortem examination cases (7 spleen, 1 liver) and combined conjunctival-choanal-cloacal swab samples from 13 live penguins revealed 5 additional dead and 7 live penguins that were positive for SpAHV2. The presence of SpAHV2 in healthy live animals and lack of significant herpesvirus-associated lesions as the cause of death in 6 of 7 SpAHV2-positive dead penguins suggests that this virus may be an endemic in blue penguins, and that recrudescence may cause disease and death.
Herpesviruses are large DNA viruses of 125–290 kbp.4,13 Family Orthoherpesviridae is divided into 3 subfamilies: Alpha-, Beta-, and Gammaherpesvirinae; Alphaherpesvirinae is the only subfamily in avian hosts. Many herpesviruses are host-adapted, having co-evolved with a specific host species. 2 Herpesviruses are known for latency and lifelong infections. However, recrudescence of infection in immunosuppressed natural hosts, acute infection in naïve animals, or transmission to non-definitive hosts can lead to morbidity and, in some cases, fatal consequences.2,18
Multiple herpesviruses have caused severe disease and death in birds.9,12,18 Early reports of herpesviral-like disease in penguins did not include virus characterization.1,10,16 Magellanic penguin herpesvirus 1 (MagHV1), also known as penguin herpesvirus 1 in GenBank, 14 is an Iltovirus first identified in Magellanic penguins (Spheniscus magellanicus) from a marine animal rehabilitation center on the southern coast of Brazil in 2011. Infection caused death in 58.3% of penguins. 14 Other penguin herpesviruses fall in a clade containing Mardivirus, Simplexvirus, and Varicellovirus. Magellanic penguin herpesvirus 2 (MagHV2), listed in GenBank as penguin herpesvirus 2, was found in subclinical Magellanic penguins from 4 colonies in Argentina. 14 Viruses with 93.6% and 94.9% homology to MagHV2 in the polymerase gene, named spheniscid alphaherpesvirus 1 (SpAHV1), have been reported in cases of fatal necrotizing diphtheritic oropharyngitis-laryngotracheitis and enteritis in captive Humboldt (S. humboldti) and African (S. demersus) penguins, respectively. 17 Although both studies describe similar viruses, the sequence and host species and range differences may be more consistent with related but distinct herpesvirus species of congenerics. Lastly, a captive blue penguin chick with fatal encephalitis housed in an exhibit with African penguins and southern rockhopper penguins (Eudyptes chrysocome) had a herpesvirus 100% homologous to the African penguin sequences (SpAHV1). 24
Blue penguins (Spheniscidae, Eudyptula minor), also known as little penguins, little blue penguins, fairy penguins, and their Māori name kororā, live and breed along the coast of New Zealand. 3 Although listed as a species of Least Concern by the International Union for the Conservation of Nature (IUCN) in 2020, populations have been decreasing and are considered at risk due to a number of threats. 3
We describe here a novel alphaherpesvirus associated with myocardial degeneration and necrosis and splenic reticuloendothelial (RE) cell hyperplasia and necrosis in Eudyptula minor in a zoo collection. On 2021 Feb 9, a 9-y-old female blue penguin at the Bronx Zoo (Bronx, NY, USA) was found dead in its nest (case 1). On 2021 Feb 27, an 18-y-old female blue penguin died after 1 d of leg swelling and difficulty ambulating (case 2).
Postmortem examination findings included moderate splenomegaly (case 1); a normally formed soft-shelled egg was present in the distal oviduct of each bird. Tissues collected from both cases (skin, skeletal muscle, eye, heart, lung, spleen, liver, kidney, tongue, trachea, esophagus, proventriculus, ventriculus, small intestine, large intestine, ceca, pancreas, oviduct, ovary, thyroid glands, adrenal glands, uropygial gland, peripheral nerve, brain) were fixed in 10% neutral buffered formalin and processed using routine methods. 5-µm tissue sections were stained with H&E for histologic examination. Select tissues (liver, spleen, brain) were collected and archived at –80°C.
Histologic findings included moderate splenic RE cell hyperplasia with multifocal RE cell necrosis and lymphocytolysis in both cases (Fig. 1A, 1B), and moderate-to-severe, multifocal-to-regionally extensive myocardiocyte atrophy, degeneration, and necrosis in one bird (case 2; Fig. 1C, 1D). Small-to-moderate numbers of myocardiocyte nuclei (case 2) contained eosinophilic intranuclear inclusion bodies (INIBs; Fig. 1D). Similar INIBs were seen in splenic RE cells of both birds (Fig. 1B). Degenerative cardiac changes were the cause of death in case 2. A clear cause of death was not identified in case 1.
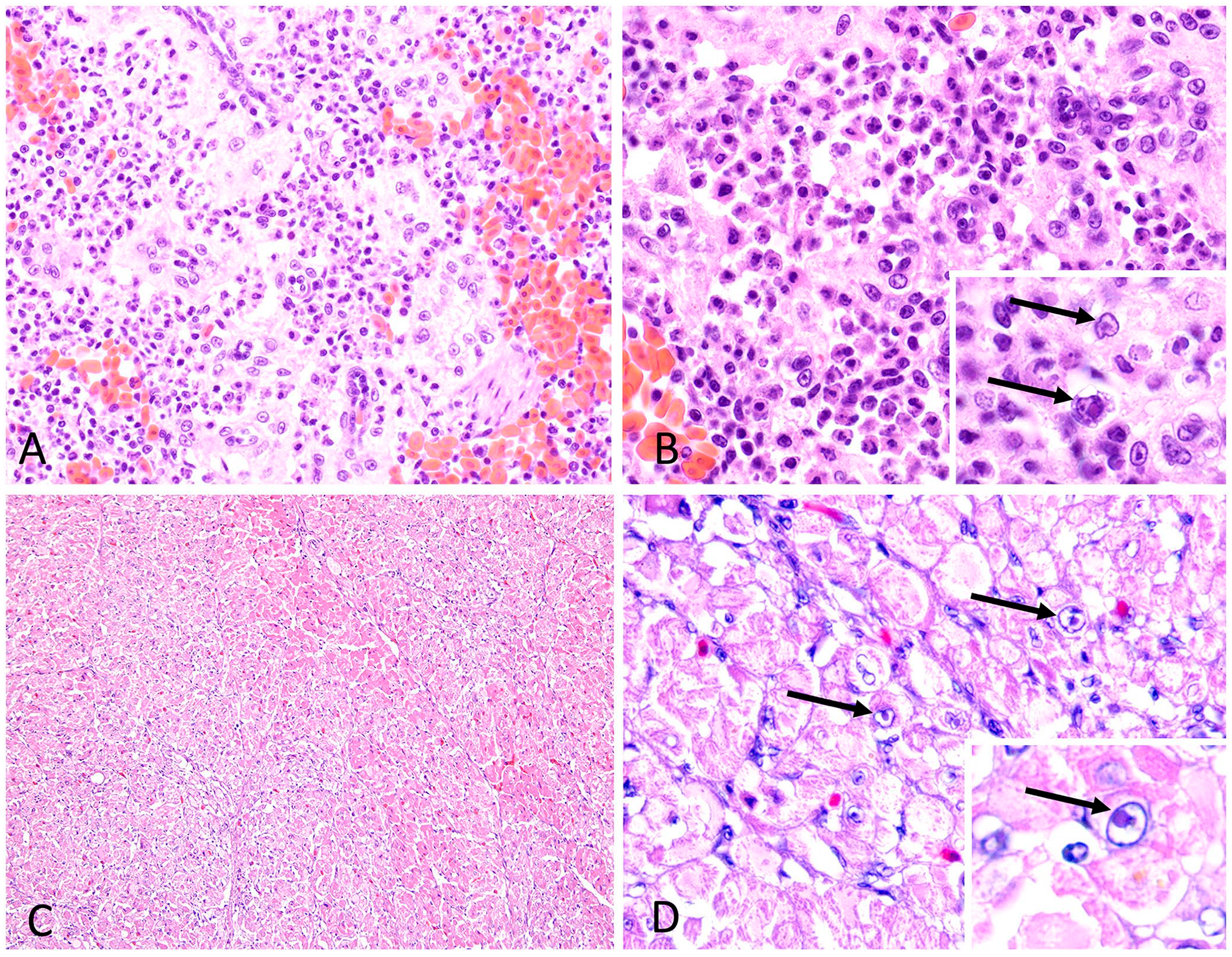
Notable histologic findings included:
Frozen spleen from each bird was tested for herpesviruses, paramyxoviruses, and hemosporidian parasites (genera Haemoproteus, Leucocytozoon, Plasmodium) by PCR (see below). Polyomavirus and adenovirus testing was also performed on spleen from case 1. Other testing included viral culture (liver, both cases; Cornell University Animal Health Diagnostic Center [CU-AHDC], Ithaca, NY, USA) and vitamin E, vitamin A, and mineral analysis (case 2; Michigan State University Veterinary Diagnostic Laboratory, E. Lansing, MI, USA).
Total nucleic acid was extracted from tissue samples and formalin-fixed paraffin-embedded (FFPE) tissue samples as described previously. 18 PCR amplification was performed (One-Step RT-PCR kit for RNA viruses, Qiagen; or AmpliTaq 360 master mix, Thermo Fisher) for DNA viruses and hemosporidian parasites. Tissue and FFPE samples were tested using the following conventional PCR targets as described previously: herpesvirus DNA polymerase gene, Mod-DFA-F1: GAYTTYGCNAGYYTNTAYCCR; ILK-F2, TCCTGGACAAGCAGCARNYSGCNMTNAA; KG1-R: GTCTTGCTCACCAGNTCNACNCCYTT; TGV-F: TGTAACTCGGTGTAYGGNTTYACNGGNGT; IYG-R: CACAGAGTCCGTRTCNCCRTADAT15,25; paramyxovirus L gene (large polymerase)19,23; Haemoproteus, Leucocytozoon, and Plasmodium cytochrome b gene7,21; adenovirus DNA polymerase gene19,26; and polyomavirus VP1 gene. 8 Positive, negative, and inhibition controls were included in all assays. Amplified PCR products of appropriate molecular weight were enzymatically purified or gel-purified (ExoSAP-IT express and PureLink quick gel extraction kit; Thermo Fisher). Purified PCR products were directly sequenced in the forward and reverse directions (Eton Bioscience) and trimmed of the primers; a consensus sequence was generated using Geneious software (Prime v.2019.2.3; Biomatters).
Splenic tissues from cases 1 and 2 were PCR-positive for herpesvirus and negative for all other PCR tests. FFPE cardiac tissue from case 2 was also positive for herpesvirus; 178 bp of trimmed DNA sequences were obtained from each positive sample; all were 100% identical to each other. BLASTN (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Nucleotides/) analysis of the consensus sequences from these cases to the GenBank nucleotide database showed that the sequence was 71.4% identical to ciconiid herpesvirus 1 (GenBank MN017363, highest query coverage 86%). Viral cultures of liver tissue were negative in both cases (CU-AHDC), and vitamin and mineral results (case 2) were within normal RIs. RNAscope and BaseScope in situ hybridization (Advanced Cellular Diagnostics) were also attempted using 2 different custom probes on FFPE heart and spleen from case 2. Although rare amplification was detected using one of the custom herpesviral probes (RNAscope protocol), there was substantial off-target signal seen with 2 different negative control probes (bacterial gene DapB, scrambled). These assays were therefore considered uninterpretable.
Subsequent to these positive results, we reviewed blue penguin pathology records and sampled the resident collection of blue penguins to investigate the presence of this novel herpesvirus. Archived frozen tissue samples (spleen or liver) from 8 blue penguins that died from 2016–2023 were available. All were tested for herpesvirus, and 4 were tested for paramyxovirus. Samples from 5 of the 8 were PCR-positive for herpesvirus. Deaths in these cases occurred on 2018 Feb 5 (case 3, female), 2017 Mar 6 (case 4, male), 2022 Dec 16 (case 5, male), 2023 Mar 26 (case 6, female), and 2023 Jun 26 (case 7, male). Paramyxovirus PCR testing was negative for cases 3 and 4, and testing was not performed in cases 5–7. DNA sequence analysis of the herpesvirus PCR product from the 5 herpesvirus-positive autopsy cases showed that all sequences were 100% identical to the sequences from cases 1 and 2. Herpesvirus-associated disease was not the cause of death in any of these cases.
Combined choanal-cloacal-conjunctival swab samples were collected from all 13 live blue penguins in our collection. Total nucleic acid extraction was performed using 150 µL of PrepMan (Thermo Fisher) according to the manufacturer’s protocol, and swab extracts were diluted 1:10 in RNase/DNase-free water prior to PCR (described above). Swab samples from 7 of 13 birds were positive for herpesvirus. Two of the swabs tested were from birds that later died (cases 5, 7). Antemortem and postmortem samples were positive in case 5; the antemortem sample was negative and postmortem sample was positive in case 7. All sequences were 100% identical to each other and to cases 1–7. Three samples had DNA sequences that contained one random ambiguous nucleotide in each sequence.
To obtain additional DNA polymerase gene sequence for phylogenetic analysis, additional PCR testing was performed on a herpesvirus-positive swab sample using primers that targeted a 404-bp sequence upstream and overlapped with the TGV and IYG target region. This overlapping region was amplified with the forward primer SIIQ-F 5 (5′-AGYATHATHCARGCNCAY-3′) and a reverse blue penguin herpesvirus–specific primer developed using the original 178-bp sequence: LBP-Hpol-R (5′-GTGGCCCACCTACTGTGTAT-3′). To obtain additional downstream sequence, we amplified a 354-bp target from the swab sample using the forward LBP-Hpol-F (5′-ATACACAGTAGGTGGGCCAC-3′, reverse complement of LBP-Hpol-R) and primer KG1. 25 A 635-bp contiguous DNA sequence of herpesviral DNA polymerase gene was subsequently created by aligning the original trimmed 178-bp amplicon with the longer trimmed amplicons (383-bp and 272-bp) from the blue penguin swab sample. BLASTN analysis of the longer nucleotide sequence against GenBank showed that SpAHV1 was the closest match (68.3% identity) when filtered for the highest query coverage (86.7%). Based on the standards set by the International Committee on the Taxonomy of Viruses, 11 we named the herpesvirus in the blue penguins spheniscid alphaherpesvirus 2 (SpAHV2). The contiguous sequence has been accessioned in GenBank (PP501783).
The 635-bp nucleotide sequence was translated into a 211 amino-acid sequence. The amino-acid sequence was aligned with 54 representative alpha-, beta-, and gammaherpesvirus amino-acid sequences from GenBank using the Geneious global alignment with free end gaps tool (BLOSUM62 cost matrix, with guide tree option and up to 99 iterations). The resulting alignment was color mapped using Geneious software, and a BLOSUM62 scoring matrix with a default setting of 1 (Prime v.2019.2; Biomatters). The alignment was trimmed to 152 amino acids to accommodate shorter sequences (Fig. 2). Ciconiid herpesvirus 1 (white stork), Fregata magnificens herpesvirus (magnificent frigate bird), and SpAHV1 were among the closest in identity to SpAHV2 based on distance matrix analysis and GenBank searches.
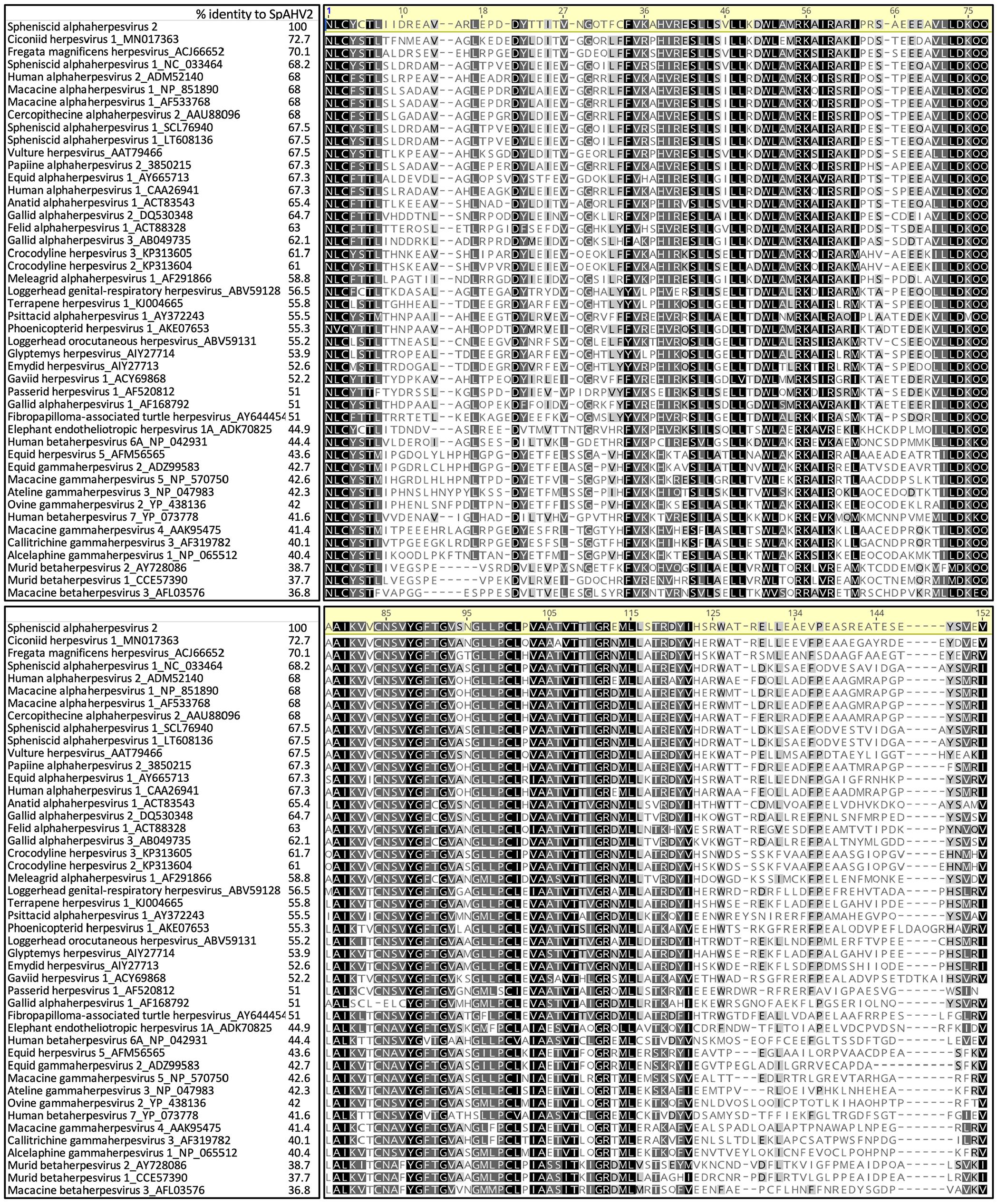
Protein alignment of a 152 amino-acid sequence of the DNA polymerase gene. Amino acids that are 100% similar throughout the alignment are shown in black, those with 80–100% similarity are shown in dark gray, those with 60–80% similarity are shown in light gray, and those with <60% similarity are shown in white using a BLOSUM identity score matrix (Geneious Prime). GenBank accessions precede the names of each herpesviral species.
Bayesian phylogenetic analysis was used to determine the relationship of the blue penguin herpesvirus amino-acid sequence to other Orthoherpesviridae, as described previously. 18 Trees were finalized and labeled (FigTree v.1.3.1, 2006–2009, http://tree.bio.ed.ac.uk/software/figtree/). SpAHV2 is a member of the subfamily Alphaherpesvirinae and forms a unique lineage with strong Bayesian support values (posterior probability 100%) that groups within the Mardivirus/Simplexvirus/Varicellovirus clade and unclassified avian herpesviruses including ciconiid herpesvirus 1, Fregata magnificens herpesvirus, vulture herpesvirus, and SpAHV1 (Fig. 3). These avian herpesviruses are closest to the origin of the mammalian alphaherpesviruses, and further study may provide insights into a host jump first infecting mammals.
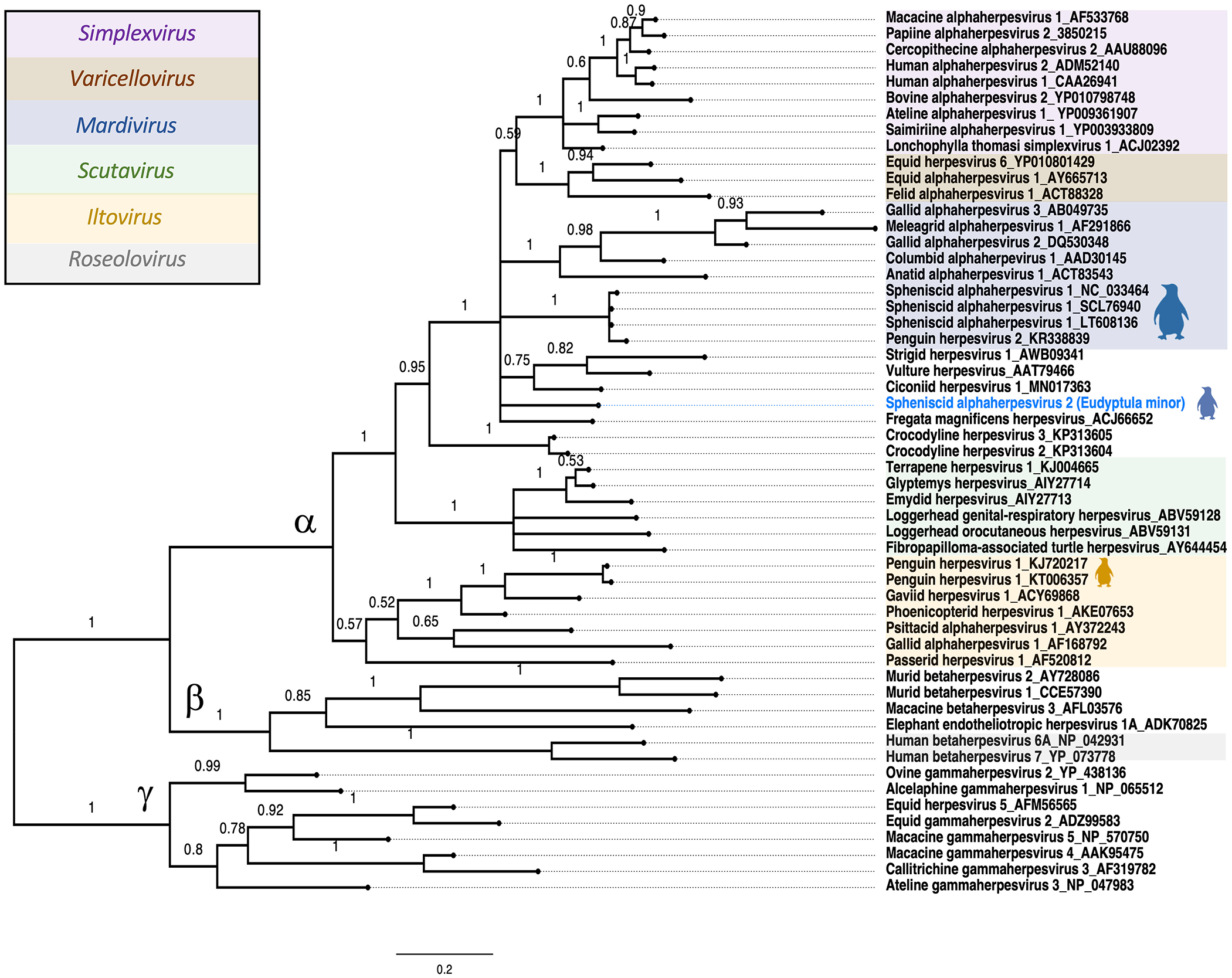
Bayesian phylogenetic tree of the herpesviral DNA polymerase amino-acid sequences from representative Alphaherpesvirinae, Betaherpesvirinae, and Gammaherpesvirinae subfamilies obtained from GenBank. Bayesian posterior probabilities of branching demonstrate the robustness of the individual clades. Spheniscid alphaherpesvirus 2 from blue penguins (Eudyptula minor) is shown in blue, along with other penguin herpesviruses noted by the silhouette icons.
Retrospective sample testing confirmed SpAHV2 in our collection as early as 2016. SpAHV2 was associated with myocardial degeneration and atrophy in an 18-y-old female penguin (case 2), and splenic RE cell necrosis in 2 cases (cases 1, 2), but was not associated with histologic lesions, death, or clinical disease in other subclinical SpAHV2-positive penguins. Herpesviruses are known to co-evolve with specific hosts, and viral latency is a key feature and mode of viral persistence. 20 Detection of the virus in apparently healthy animals is suggestive of host-adaptation or co-evolution in blue penguins. Predisposing factors for herpesviral recrudescence in human and animal infections include immunosuppression, microbial infections, trauma (e.g., surgery, tissue injury, nerve damage), systemic stress, fever, hormonal imbalances, age-related factors, and stressors related to reproductive activity.6,22 Acute infection or recrudescence of latent herpesviral infection may have been factors in the death of case 2 and development or virus-related hyperplasia and cell necrosis in case 1 in our study. Future studies in blue penguin populations and other penguin species could provide insight into the potential pathogenicity of this virus and its association with morbidity and mortality, potential host range, and the relationship to other penguin viruses.
Footnotes
Acknowledgements
We thank the staff of the Wildlife Conservation Society Ornithology Department and Zoological Health Program clinical and pathology staff, as well as Gretchen Anchor at Zoological Pathology Program University of Illinois for performing RNAscope and BaseScope.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
