Abstract
Piglet lethality is one of the major concerns in pig breeding programs. Deletion of a 212-kb region within the Bardet-Biedl syndrome 9 (BBS9) gene has been linked to a reduction in the number of piglets born alive per litter. The BBS9 mutant gene carrier-by-carrier mating scheme could result in mummification of piglets carrying 2 copies of the BBS9 mutant allele, which ultimately affects the reproductive performance of the sow. Our aim was to develop a simple, rapid, and cost-efficient method that could be applied in a BBS9 mutant gene carrier screening program in low- and middle-income countries within basic laboratory settings. Here, we report an optimized multiplex PCR assay that we have established successfully for detection of a 212-kb deletion within the BBS9 genomic sequence. We genotyped 420 animals from Yorkshire, Duroc, and Landrace purebred populations in Vietnam. We found that while the BBS9 mutant allele was not identified in Duroc pigs, the frequency of BBS9 carriers was 10% in both Yorkshire and Landrace populations. We subsequently validated our results using Sanger sequencing. Our multiplex PCR method could be utilized as a BBS9 screening test in pig breeding programs.
The global demand for agricultural products is projected to increase by 60% by 2050 to meet the needs of an estimated 9.7 billion people. 7 Currently, pork is one of the most consumed types of animal protein, accounting for 33% of world per capita consumption. 6 In Southeast Asian countries, pork is an important component of daily meals due to its high quality. According to the Food and Agriculture Organization, the demand for pork products is calculated to grow at 13.1% annually. 3 This clearly puts pressure on pig farming to be successful.
In pig production, the number of weaned piglets per sow is one of the main determinants of profitability. As a result, since ~2000, research has shifted toward increasing the number of piglets born per litter. However, genetic improvements of litter sizes are also associated with the number of piglets not born live, including mummifications, which are the result of fetal death after ossification. Mummified piglets are among the leading causes of fertility loss, and mummification severely hampers the reproductive performance of sows. The rates of mummified piglets are reported as 3–8%, which could be attributed to several factors, including housing, nutrition, infections, immunity, and management. 9 A 2018 study using next-generation sequencing found that genetic variation also contributes to the mummified piglet phenomenon. 2 A structural variant was identified that was significantly correlated with the number of mummified piglets per litter. A recessive lethal mutation involving a 212-kb deletion within the Bardet-Biedl syndrome 9 (BBS9) genomic sequence leads to a remarkable reduction of gene expression of the bone morphogenetic protein-binding endothelial regulator gene, an important gene required for normal fetal development. 2 While increased numbers of mummified piglets occur per litter when homozygotes harbor 2 copies of the BBS9 mutant allele (del/del genotype), heterozygotes with 1 copy of the BBS9 mutant allele (+/del genotype) have an increased growth rate, which is an agriculturally desirable trait in the pig breeding industry. Our results suggest that despite the lethal effect of the BBS9 mutant allele, we need to maintain its frequency in pig herds.
There are several documented PCR-based detection approaches, including whole-genome sequencing, real-time PCR, multiplex ligation-dependent probe amplification (MLPA), and DNA microarray assays, that are utilized to detect structural variants. 10 The preference for the various techniques depends on the sensitivity and specificity of the methods, the budget of the laboratory, the training of the technical staff, and the time required for the workflow. However, given the need to acquire specialized equipment and supplies, and the quantity of samples to be processed, the high cost of these methods constitutes their greatest disadvantage.
Multiplex PCR is a variant of conventional PCR in which several sets of primers are utilized to detect 2 or more loci in a single reaction. Therefore, multiplex PCR has been applied successfully in numerous genetic tests including screening of large deletions, an important genomic structural variation. 4 As a result, for laboratories in low- and middle-income countries, the use of multiplex PCR is recommended, when considering cost, ease of use, and performance. We aimed to develop an efficient, rapid, and cost-effective multiplex PCR method to detect BBS9 carriers (+/del genotype) animals in commercial pig herds in southern Vietnam.
Samples were collected only for routine monitoring purposes in the breeding programs and not specifically for our project. Sample collection and data recording were conducted according to the Vietnamese law on animal protection and welfare. In our study, a total of 320 12-mo-old animals from 3 commercial purebred pig breeds (110 Yorkshire, 100 Duroc, 110 Landrace) were selected from several pig farms in Binh Duong Province, Vietnam. All chosen pigs were healthy purebreds. Animals were reared in groups of 12–14 individuals under the closed-house system, with an ambient temperature of 28–30°C and relative humidity of 60–70%. Blood samples were aseptically collected from the jugular vein of the animals as described previously. 8 Ten stillborn Yorkshire piglets were collected from different farms and stored at −20°C until use.
Genomic DNA was extracted (GeneJet whole blood genomic DNA purification kit; Thermo Fisher) according to the manufacturer’s instructions. The integrity of extracted DNA was analyzed by gel electrophoresis. The DNA concentration was determined with a spectrophotometer (Nanodrop 2000; Thermo Scientific). Samples with a 260/280 ratio between 1.8 and 2 were diluted to a final concentration of 50 ng/µL and were used for PCR amplification. The PCR analysis was carried out by a gel electrophoresis procedure. The electrophoresis conditions were 150 V for 30 min using 0.5× Tris–borate–EDTA (TBE) buffer along with a 100-bp molecular weight marker to determine the size of the amplified fragments. Gel images were captured (UVP GelDoc-It2; Fisher Scientific).
Based on the location of the deletion (position 39,817,373 to 40,029,300) within the BBS9 genomic sequence (NCBI reference sequence NC_010460.4), 2 sets of primers were designed using Primer3 software (https://primer3.ut.ee/; Table 1). The specificity of the designed primers was verified using BLAST (https://www.ncbi.nlm.nih.gov/tools/primer-blast/index.cgi).
BBS9 gene multiplex PCR primer sequences.
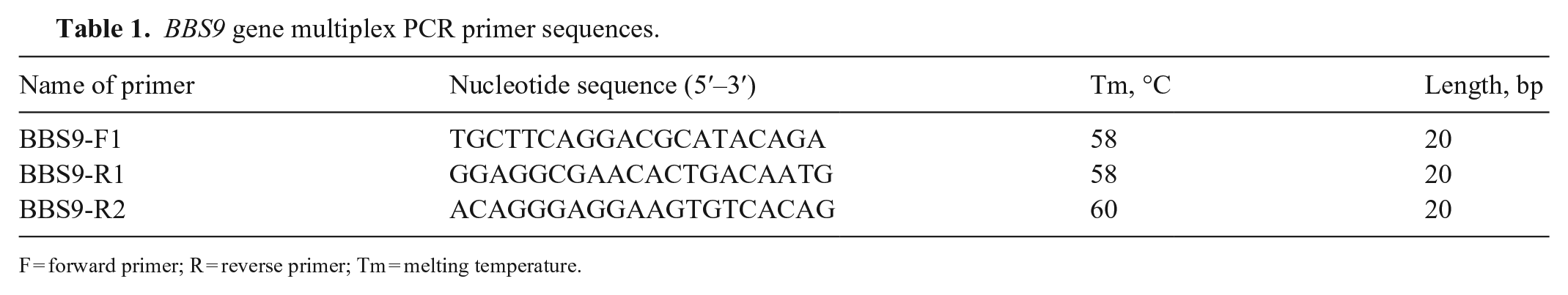
F = forward primer; R = reverse primer; Tm = melting temperature.
Each primer set (Phu Sa, Vietnam) had one forward primer in common. The forward primer BBS9-F1 was located 500 bp upstream of the deletion (position 39,816,902 to 39,816,922); reverse primer BBS9-R1 was designed to match the sequence in the 212-kb deletion (position 40,029,029 to 40,029,048); the other reverse primer BBS9-R2 was placed 500 bp downstream from the 212-kb deletion (position 40,029,772 to 40,029,753; Fig. 1). Primers were diluted with sterile water and stored at −20°C until use.
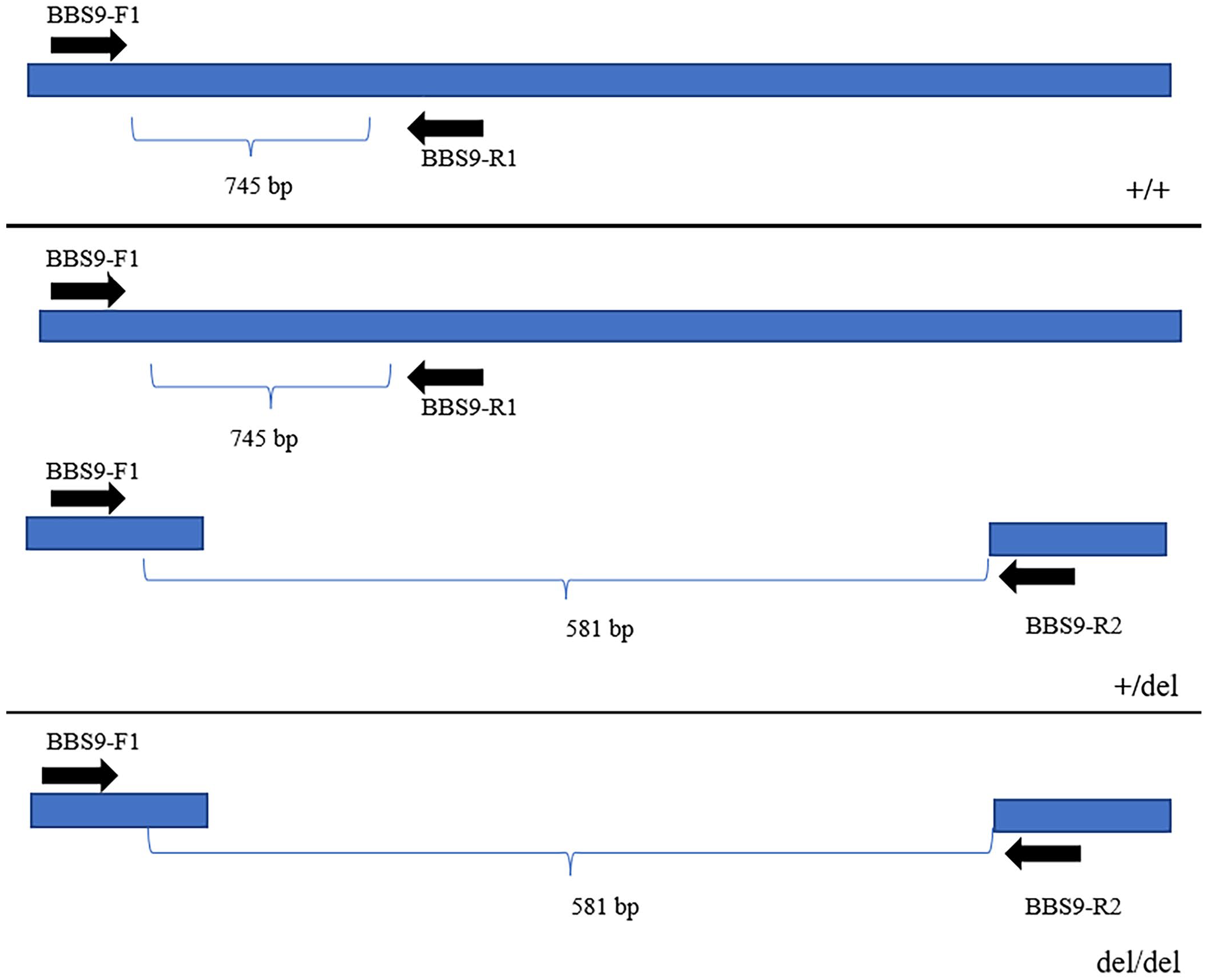
Schematic drawing of the location of PCR primers. Two PCR primer pairs using the same forward primer to amplify homozygous wild-type (+/+), heterozygous (+/del), and homozygous mutant (del/del) are shown. The forward primer is located upstream of the deletion boundary.
For standardization, a synthetic gene mimicking the deletion from BBS9 was manufactured (Integrated DNA Technologies) and synthesized at a concentration of 50 ng/µL. We used 10 ng of synthetic gene in our PCR assay. The F1-R1 combination primers amplify a region of 745 bp, corresponding to the BBS9 + allele (wild-type allele). The F1-R2 combination primers amplify a region of 581 bp, corresponding to the del allele (mutant allele). After optimization, the multiplex PCR was performed in a 25-µL mixture containing 100 ng of DNA, 0.2 µL of 200 µM dNTP, 0.75 µL of 50 mM MgCl2, 0.5 µL of Taq polymerase, 0.5 µL of 10 pM primers, and 10 µL of 1× PCR buffer (Suppl. Fig. 1). The reaction conditions for multiplex PCR were as follows: pre-incubation at 94°C for 5 min; 35 cycles including denaturation at 94°C for 30 s, annealing at 61°C for 30 s, extension at 72°C for 60 s; 1 cycle of final extension at 72°C for 7 min. The PCR was carried out in a thermal cycler (Gene Amp PCR system; Thermo Fisher). After thermal cycling, 5 µL of the amplified products were analyzed by horizontal electrophoresis on 1% agarose gels.
To validate the accuracy of our developed multiplex PCR method, amplified products associated with BBS9 genotypes +/+ (745 bp), +/del (745 bp and 581 bp), and del/del (581 bp) were purified by ethanol precipitation and were sequenced by the Sanger method in both directions (Genlab, Vietnam). The primers used for sequencing were used in the multiplex PCR.
We performed a screening test by utilizing multiplex PCR on 320 animals. The 212-kb deletion from the BBS9 gene was not detected in Duroc swine; the frequency of +/del individuals in both Yorkshire and Landrace pigs was 10% (Table 2). Because of the poor quality of the DNA, we were unable to extract DNA from 10 mummified Yorkshire piglets, as described by others. 2
Genotype and allele frequency of BBS9 gene in 3 commercial pig breeds: Yorkshire, Landrace, and Duroc.

Despite increasing applications of multiple target detection, multiplex PCR detection is still challenging because of potential complex interactions among multiple primer sets. When developing a multiplex PCR assay, the most crucial step is primer design, which usually defines the specificity and sensitivity of the assay. Because of the difference in melting temperatures of 3 primers, we carried out tests of optimization to determine the optimal annealing temperature of each primer. We found that the optimal annealing temperature was 61°C. As expected, the dominant homozygote BBS9 +/+ and the recessive homozygote del/del were observed as a single band of 745 bp and a single band of 581 bp, respectively. In contrast, the heterozygote +/del was seen with both bands. The 212-kb deletion from the BBS9 gene was confirmed by Sanger sequencing from PCR products of 2 dominant homozygotes BBS9 +/+ and 2 heterozygotes +/del (Suppl. Fig. 1). Hence, the design of oligonucleotides and optimization of PCR conditions were suitable for distinguishing 3 BBS9 genotypes (Fig. 2).
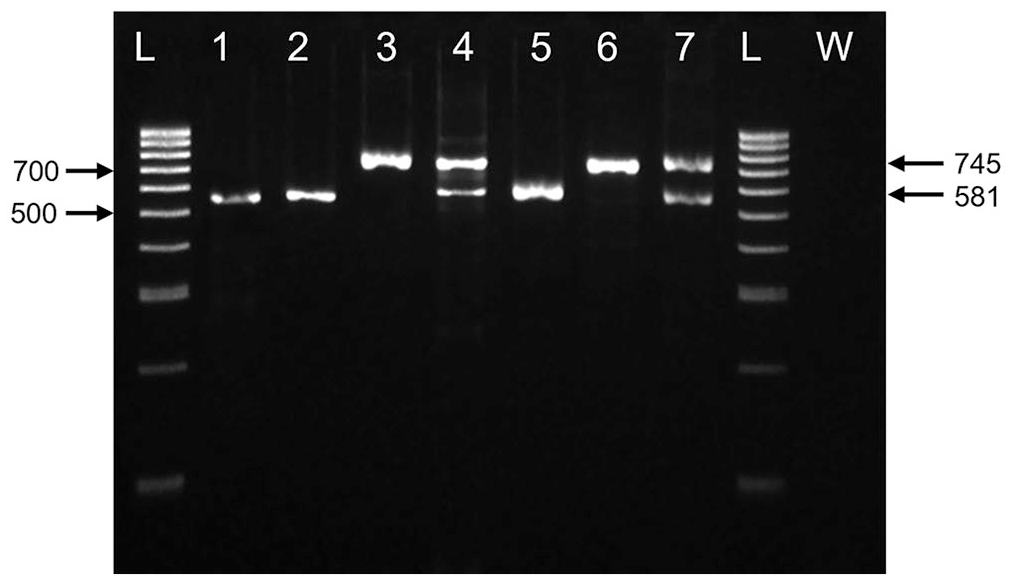
PCR amplification of the structural variant of the BBS9 gene. From left to right: agarose gel picture of 3 del/del genotypes (lanes 1, 2, 5), 2 carrier genotypes (lanes 4, 7), 2 wild-type genotypes (lanes 3, 6). L = 100–1,000-bp molecular marker; W = water sample.
Our findings suggest that the 212-kb deletion was present in Yorkshire and Landrace swine (Table 2). This observation is not in agreement with a study that indicated that this mutation was present only in Yorkshire swine 3 ; however, that study was of pigs in the Netherlands whereas, in our study, pig stock had been imported from Canada, France, and Denmark. Additionally, we found that the frequency of the carrier state was lower than in the study in the Netherlands (10% vs. 11.8%). A limitation of our study is the smaller sample size (423) than in the previous study (22,300). 3 More samples will need to be analyzed to obtain a more accurate estimate of the frequency of BBS9 carriers in commercial pigs in southern Vietnam.
The multiplex PCR method has been documented in the detection of large genomic deletions in cattle and pigs.4,5 For instance, the multiplex PCR was successfully applied in screening for a 50-kb deletion within the RSPO2 gene in Holstein-Friesian cattle. 1 In pigs, numerous large deletions were discovered and genotyped; however, the conventional PCR method used in that study could not differentiate the deletion carrier from the wild-type when the deletion was >10 kb. 4 Our multiplex PCR method can distinguish the 212-kb deletion in the BBS9 gene and provide an efficient genomic screening approach for pig breeders to select the appropriate mating scheme. Compared to other time-consuming and laborious methods, including WGS, and rtPCR plus MLPA, a multiplex PCR assay can circumvent these limitations due to its ability to amplify multiple alleles in a single reaction with minimal economic setup and professional training. In short, our method not only provides a reliable, cost-effective, and practical tool for BBS9 carrier detection but also promotes a monitoring system for piglet lethality in commercial pig operations.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387241282082 – Supplemental material for Novel genotyping assay for a 212-kb deletion from the BBS9 gene, and frequency of the allele in pig populations in Vietnam
Supplemental material, sj-pdf-1-vdi-10.1177_10406387241282082 for Novel genotyping assay for a 212-kb deletion from the BBS9 gene, and frequency of the allele in pig populations in Vietnam by Nguyen H. Tinh, Nguyen V. Hop, Pham T. Phuong, Trinh L. H. Tam, Nguyen B. Quoc, Trinh H. Son and Anh P. N. Bui in Journal of Veterinary Diagnostic Investigation
Footnotes
Data-sharing statement
Raw data were generated at the Animal Genetics Laboratory, Faculty of Biotechnology, Ho Chi Minh City Open University, Vietnam. Derived data supporting the findings of our study are available from the corresponding author on request.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our research was funded by the Ho Chi Minh City Open University, Ho Chi Minh City, Vietnam grant E2022.01.2 and the Ministry of Agriculture and Rural Development grant 5285/QD-BNN-KHCN.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
