Abstract
Porcine astrovirus (PAstV) has been associated experimentally with diarrhea in piglets, but much more knowledge is needed about this virus. PAstV has high genetic variability, and 5 genotypes have been identified, namely PAstV1–5. To obtain information on the epidemiology of PAstV, we established a multiplex PAstV PCR assay to detect and differentiate the 5 PAstV genotypes simultaneously. The assay utilized specific primers for each genotype, producing fragments of 307, 353, 205, 253, and 467 bp, representing PAstV1–5, respectively. Our multiplex PCR assay amplified all 5 DNA fragments from single or mixed viral genomes without cross-reactions with other PAstV genotypes or other viruses in pigs. The limit of detection of the multiplex PCR assay was 5 × 102 copies/μL for PAstV1 and PAstV4, and 5 × 103 copies/μL for PAstV2, PAstV3, and PAstV5. We examined 76 pig fecal specimens with our multiplex PCR assay. PAstV was detected in 36 of 76 (47.4%) samples; ≥2 PAstVs were found in 20 of 76 (26.3%) samples. The multiplex PCR assay results were essentially the same as the results using a monoplex PAstV PCR assay, with a coincidence rate of >96%. Our multiplex PCR method provides a simple, sensitive, and specific detection tool for PAstV detection and epidemiologic surveys.
Astrovirus (AstV; Astroviridae) is a spherical, single-stranded, positive-sense RNA virus that is characterized by a typical star structure. 8 AstVs are widely distributed and can infect a multitude of host species, including humans, pigs, dolphins, and other animals.4,13 Pigs were recognized as an AstV host in the 1980s when porcine AstV (PAstV) was reproduced successfully in pig embryonic kidney cells. 2 Early findings indicated that, under experimental conditions, the virus could cause mild diarrhea in piglets. 8 The PAstV genome has 3 consecutive open reading frames (ORFs): 1a, 1b, and 2. ORF1a and ORF1b, situated at the 5′-end, encode nonstructural polyproteins, including a protease and a RNA-dependent RNA polymerase; ORF2 at the 3′-end encodes the capsid protein. 17 Based on genetic differences in the complete capsid sequence, 5 genotypes of PAstV (PAstV1–5) have been identified, with different prevalences worldwide.25,29
Traditionally, AstVs are associated with intestinal disorders in most mammals, including diarrhea and vomiting that usually lasts 2–3 d after infection, especially in immunocompromised individuals.16,29 Only a small percentage of AstVs were pathogenic in infected hosts; most animals were infected subclinically.5,16 Given the difficulty of in vitro isolation of AstVs to test infection, only PAstV1 and PAstV5 have been isolated from cell culture, and have been shown to be associated with intestinal pathogenicity, including damage to the small intestinal mucosa resulting in mild diarrhea and growth retardation.6,17
PAstVs have also been found outside the intestinal tract. For example, PAstV2 has been demonstrated in the brains of neonatal piglets with congenital tremor, 1 and PAstV3 was described in CNS tissues of piglets and sows.22,23 As a rather prevalent genotype, PAstV4 has been detected in nasal swabs and feces from pigs. 29 A novel PAstV4 strain, with ~53% ORF2 amino acid sequence identity to previously characterized PAstV4 isolates, has been identified in nasal swabs from pigs with acute respiratory diseases, and it has been suggested that it may cause septicemia and systemic transmission. 19 These findings suggest that the virus has more complex potential pathogenicity. Therefore, the establishment of an efficient and simple multiplex detection method for PAstVs may be helpful to understand the prevalence of the virus and to enable formulation of accurate prevention and control strategies.
Before the development of molecular technology, virus isolation and electron microscopy (EM) were widely used in the diagnosis of viral gastroenteritis.20,24 However, the isolation and propagation of AstV in cell culture is complicated8,19; EM is less sensitive, and the virus is usually detected only during the peak period of viral spread. These previously important methods are no longer in use for routine detection of viruses. Other methods, such as the fluorescent antibody test, have the advantages of simple and rapid operation, but cannot identify different genotypes of AstV, and detection of the virus at low concentrations is not accurate. 10 Considering the number of samples currently required for testing, cost, and actual benefits, these methods have gradually been replaced by molecular biological testing methods. Multiplex PCR assays, which can amplify multiple targets through system optimization in one reaction, with subsequent amplified target fragment differentiation based on size by electrophoresis, have become common for detection of AstVs.7,20 The application of multiplex PCR in the genotyping of PAstV has been reported, but only for PAstV2–4 genotypes.30,34 To detect all 5 genotypes of PAstV simultaneously, 2 multiplex reverse-transcription real-time PCR panels or nested PCR assays were established and showed high efficiency and precision even at low viral titers.21,27,29 However, both methods require 2 rounds of PCR, which is time-consuming and laborious. Moreover, the cost of fluorescent quantitative PCR is relatively high, and the contamination probability of nested PCR increases greatly in the second amplification, leading to false-positive results. 26
We established a simple, specific, and sensitive multiplex PCR assay based on agarose gel electrophoresis separation to detect and distinguish 5 genotypes of PAstV simultaneously. After validation, we used our multiplex PAstV assay to test for PAstVs in 76 pig fecal specimens obtained from 6 farms with no history of clinical signs. We compared our results with those of monoplex PAstV PCR tests to determine the sensitivity and specificity of our multiplex PCR assay.
Materials and methods
Sources of viruses
Pig fecal specimens positive for each of the 5 genotypes of PAstV had been collected previously from field samples. To test specificity of our new multiplex PCR assay, we also checked for non-target viruses or virus-positive samples: porcine epidemic diarrhea virus (PEDV) and transmissible gastroenteritis virus (TGEV; Alphacoronavirus 1) bivalent vaccine strain (Harbin Weike Biotechnology), classical swine fever virus (CSFV; Pestivirus C) vaccine strain (Qianyuanhao Biological), and positive samples containing porcine circovirus 2 (PCV2), porcine reproductive and respiratory syndrome virus (PRRSV; Betaarterivirus suid), porcine rotavirus A (RVA; Rotavirus A), porcine rotavirus C (RVC; Rotavirus C), and porcine bocavirus (PBoV; Bocaparvovirus). All of these viruses or samples were already available in our laboratory.
Field specimens and pretreatment
During the period of January 2013 to March 2014, we collected 76 fecal specimens from 6 pig farms in Zhejiang province in accordance with international animal welfare standards. Specimen pretreatment was performed. 28 Specifically, 150 mg of fecal specimens were re-suspended in 1,500 μL of PBS, vortexed for 2 min, and then centrifuged at 14,463 × g for 10 min to remove impurities. Collected supernatant was immediately used for nucleic acid extraction or stored at −80°C.
Design of primers
The genomic sequences of PAstV1–5 were downloaded from NCBI and aligned using DNAStar Lasergene v.7.1. According to the alignments, primers specific for each of the 5 genotypes of PAstV were designed by using Primer Premier v.5.0 (Premier Biosoft) and verified by BLAST (Table 1; Suppl. Figs. 1–5). The primers were synthesized by Azenta Life Sciences.
Primers used in our multiplex porcine astrovirus (PAstV) PCR assay.
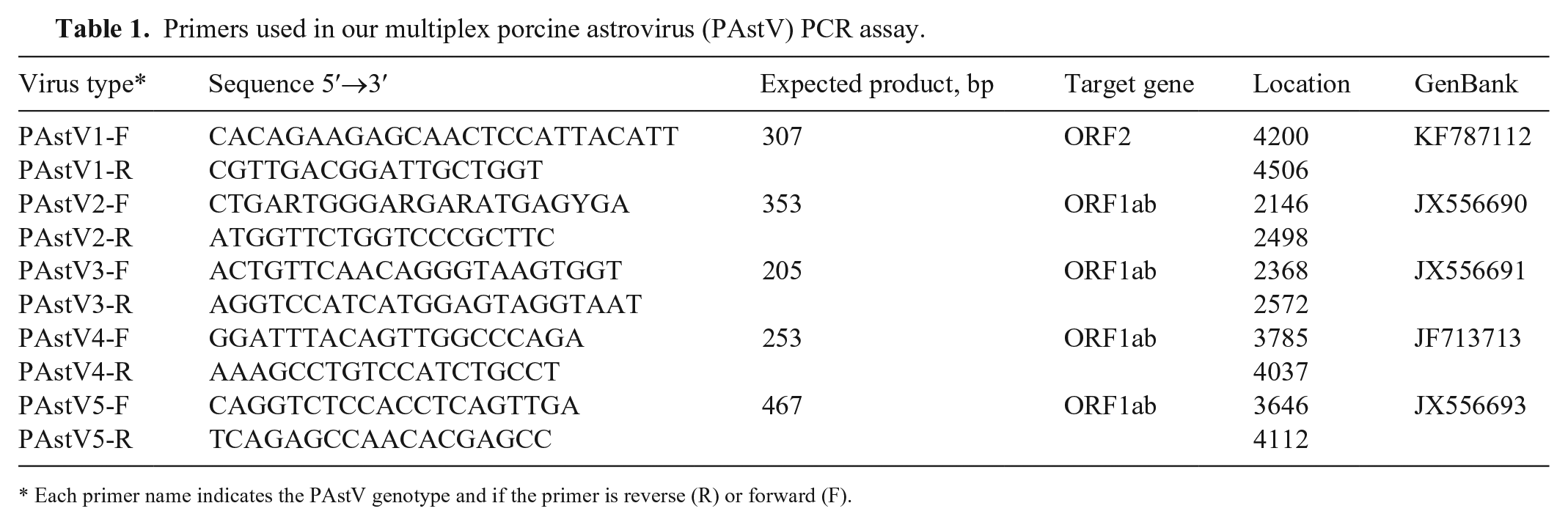
Each primer name indicates the PAstV genotype and if the primer is reverse (R) or forward (F).
Extraction of viral RNA
The genomic RNA or DNA of the viruses was extracted from the supernatant (Viral DNA/RNA miniprep kit; Axygen), according to the manufacturer’s instructions. Total RNA or DNA was dissolved with buffer TE and stored at −80°C for subsequent use.
Reverse transcription
The above RNA or DNA was used for reverse transcription (HiScript 1st strand cDNA synthesis kit; Vazyme), following the manufacturer’s instructions. The prepared cDNA or DNA was stored at −80°C for subsequent analysis.
Monoplex PCR assay and recombinant plasmid preparation
The monoplex PCR assay for PAstV1–5 was established, including 1 μL of virus cDNA or DNA, 1× master mix, 0.5 μL of each primer (10 μM; Table 1), and ddH2O to a total volume of 20 μL. Cycling reactions were performed as follows: initial pre-denaturation at 95°C for 3 min, 35 cycles of denaturation at 95°C for 30 s, annealing at 55°C for 30 s, and extension at 72°C for 30 s, followed by a final extension at 72°C for 5 min. The 5-μL amplified product was analyzed by electrophoresis on 1% agarose gel. The PCR products of each genotype were then purified and ligated with pUCm-T vector (Axygen) to construct recombinant plasmids. The plasmids were sequenced for identity confirmation by Genewiz.
After measuring the concentration and purity of each standard plasmid by Nanodrop (Thermo Fisher), the plasmid copy number was calculated according to the size of amplicons using a calculation formula. 28 The PAstV1–5 plasmids were 10-fold serially diluted to make standard plasmids with concentrations of 5.0 × 107 to 5.0 × 100 copies/μL.
Optimization and establishment of the multiplex PAstV PCR method
Using 5 PAstV plasmid standards as templates, we established our multiplex PAstV PCR assay by optimizing primer concentration, annealing temperature, the concentration of the master mix, and PCR cycling. The optimized multiplex reaction mixtures included 12 μL of 2× master mix, 0.15 μL of PAstV1 primers (10 μM), 0.45 μL of PAstV2 primers (10 μM), 0.1 μL of PAstV4 primers (10 μM), 0.4 μL of PAstV3 and PAstV5 primers (10 μM), 1 μL of viral standard plasmid or cDNA or DNA, and ddH2O to a total volume of 20 μL. Thermocycling conditions consisted of 1 cycle at 95°C for 3 min, then 35 cycles at 95°C for 30 s, 55°C for 30 s, 72°C for 30 s, and finally 72°C for 5 min. Negative controls were run with each assay. Amplification products were visualized and analyzed on 2% agarose gels.
Specificity and sensitivity assessments of the monoplex and multiplex PAstV PCR assays
The specificity of the monoplex and multiplex PAstV PCR assays was tested using different viruses, including representative members of all 5 genotypes of PAstV, PCV2, PEDV, TGEV, PBoV, PRRSV, RVA, RVC, and CSFV. The prepared diluents, including each PastV genotype or equimolar mixtures of the 5 PastV plasmids, ranging from 5.0 × 107 copies/μL to 5.0 × 100 copies/μL, were used as templates to determine the LODs for the monoplex and multiplex PCR assays.
Application of the multiplex PastV PCR assay to fecal specimens from field pigs
We tested all 76 fecal specimens from pigs without obvious clinical signs for PastVs by our multiplex PCR and by all 5 monoplex PCR assays. Products with positive results were sent to Genewiz for sequencing to confirm our test results.
Results
Specificity and sensitivity of the monoplex PCR and multiplex PastV PCR assays
Clearly visible and single bands of the expected sizes were obtained with the monoplex PCR assay for each genotype (Fig. 1). When various combinations of genotypes were present as templates in the multiplex PCR system, all of the respective amplification products were produced as expected (Fig. 2). No specific amplification products were generated in the negative controls, including ddH2O and non-targeted viruses or genotypes. Hence, the monoplex PCR and multiplex PastV PCR assays both had high specificity to the target genotypes.
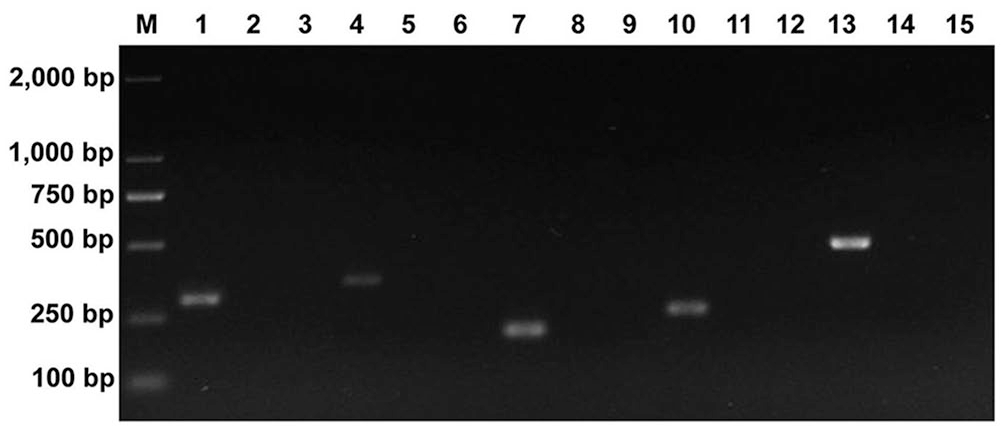
Specificity of the monoplex porcine astrovirus (PAstV) PCR assays under different template combinations. PAstV1 assay used in lanes 1–3. PAstV2 assay used in lanes 4–6; PAstV3 assay used in lanes 7–9; PAstV4 assay used in lanes 10–12; PAstV5 assay used in lanes 13–15. Lanes: 1 = PAstV1; 4 = PAstV2; 7 = PAstV3; 10 = PAstV4; 13 = PAstV5. Lanes: 2 = PAstV2, PAstV3, PAstV4, PAstV5; 5 = PAstV1, PAstV3, PAstV4, PAstV5; 8 = PAstV1, PAstV2, PAstV4, PAstV5; 11 = PAstV1, PAstV2, PAstV3, PAstV5; 14 = PAstV1, PAstV2, PAstV3, PAstV4. Non-target virus lanes 3, 6, 9, 12, and 15 are mixtures of PEDV, TGEV, PCV2, CSFV, PRRSV, RVA, RVC, and PBoV. M = 2,000-bp DNA ladder.
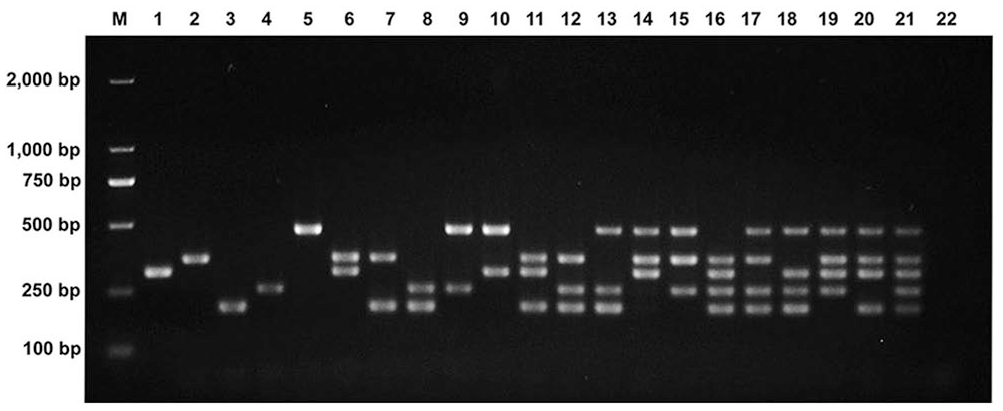
Specificity of the multiplex porcine astrovirus (PAstV) PCR assay under different template combinations. Lanes: 1 = PAstV1; 2 = PAstV2; 3 = PAstV3; 4 = PAstV4; 5 = PAstV5; 6 = PAstV1, PAstV2; 7 = PAstV2, PAstV3; 8 = PAstV3, PAstV4; 9 = PAstV4, PAstV5; 10 = PAstV5, PAstV1; 11 = PAstV1, PAstV2, PAstV3; 12 = PAstV2, PAstV3, PAstV4; 13 = PAstV3, PAstV4, PAstV5; 14 = PAstV1, PAstV2, PAstV5; 15 = PAstV2, PAstV4, PAstV5; 16 = PAstV1, PAstV2, PAstV3, PAstV4; 17 = PAstV2, PAstV3, PAstV4, PAstV5; 18 = PAstV1, PAstV3, PAstV4, PAstV5; 19 = PAstV1, PAstV2, PAstV4, PAstV5; 20 = PAstV1, PAstV2, PAstV3, PAstV5; 21 = PAstV1, PAstV2, PAstV3, PAstV4, PAstV5; 22 = negative control. M = 2,000-bp DNA ladder.
For monoplex PCR assays, the LODs of PastV1 and PastV2 were 5 × 100 copy/μL and 5 × 102 copies/μL, respectively, and 5 × 101 copies/μL for PastV3–5 (Fig. 3A–E). When the 5 PastV genotypes were present, the LODs of the multiplex PCR were 5 × 102 copies/μL for PAstV1 and PastV4, and 5 × 103 copies/μL for PAstV2, PAstV3, and PAstV5 (Fig. 3F).
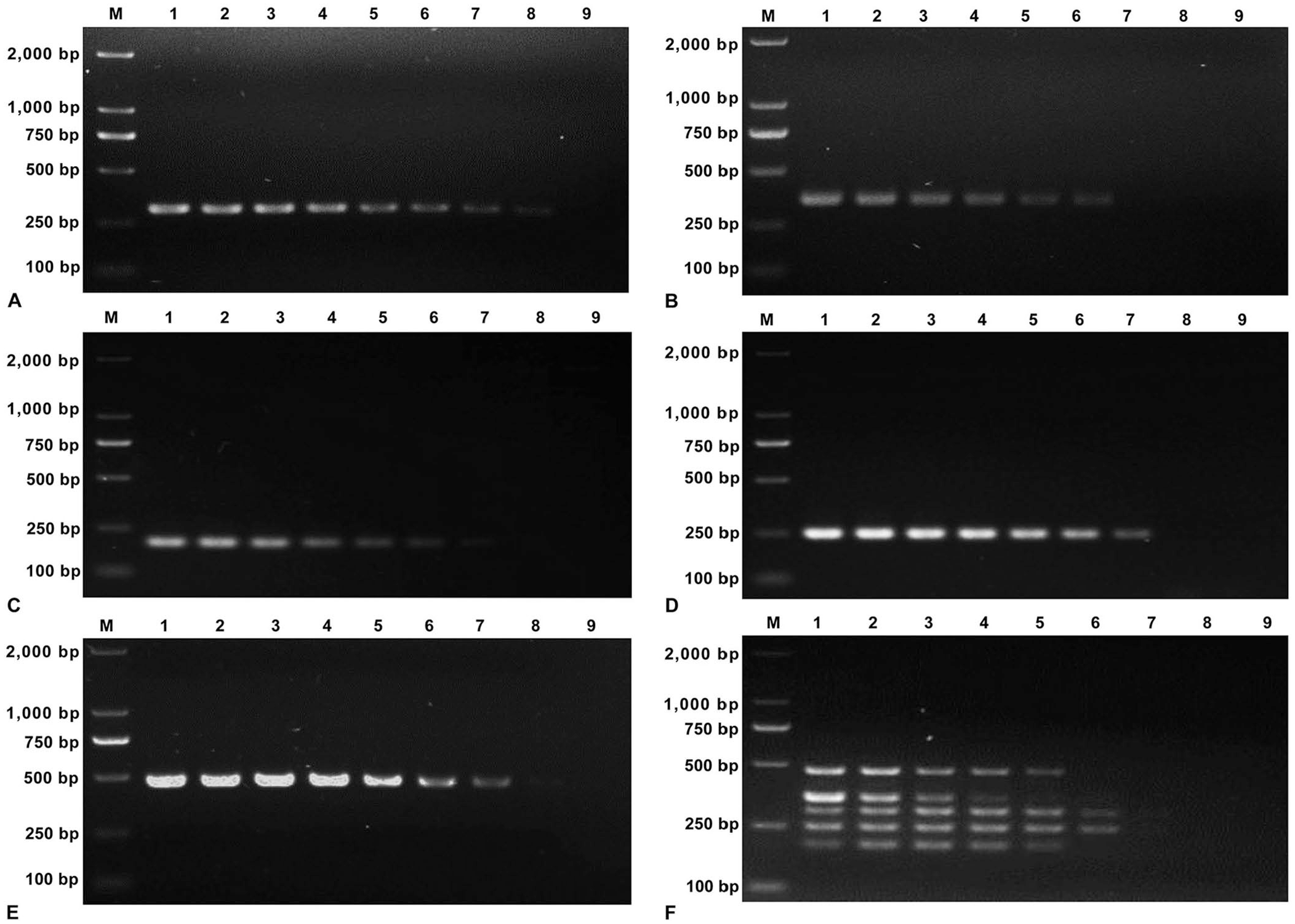
Sensitivity of the monoplex and multiplex porcine astrovirus (PAstV) PCR assays for the 5 PAstV genotypes.
Evaluation of the multiplex PAstV PCR assay using field specimens
We found 36 of 76 (47.4%) specimens to be positive by the multiplex assay, including 4 PAstV1 (5.3%), 1 PAstV2 (1.3%), 9 PAstV4 (11.8%), and 2 PAstV5 (2.6%; Table 2). PAstV3 was not detected in any of these samples. In addition, 20 of 76 (26.3%) samples were coinfected with ≥2 genotypes. To further verify the specificity of the multiplex system, some PCR products obtained from positive samples were sequenced; BLAST and phylogenetic analysis confirmed that the PAstV genotype PCR-positive samples based on amplicon size were positive for the expected genotypes (data not shown).
Detection of 5 porcine astrovirus (PAstV) genotypes in pig field fecal samples tested by monoplex and multiplex PAstV PCR assays.
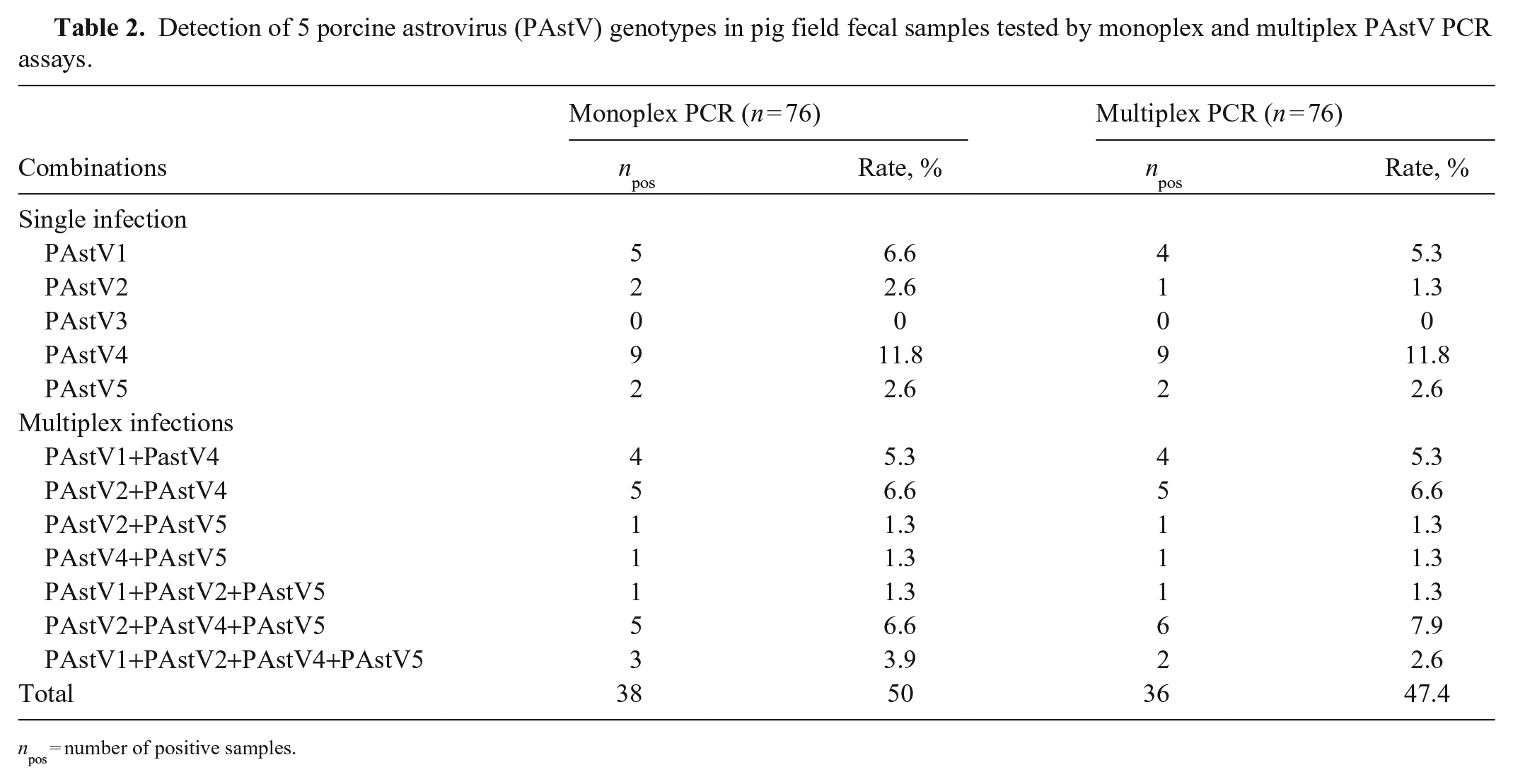
npos = number of positive samples.
According to the comparison of the genotype detection results under the multiplex and single systems (Table 3), the diagnostic specificity and sensitivity of the multiplex PCR were 98–100% and 84–100%, respectively. The coincidence rate between the multiplex and single systems was 96–100%, confirming that our multiplex PCR method is reliable and efficient.
Performance evaluation of our multiplex porcine astrovirus (PAstV) PCR assay using monoplex PAstV PCR assays as a reference.
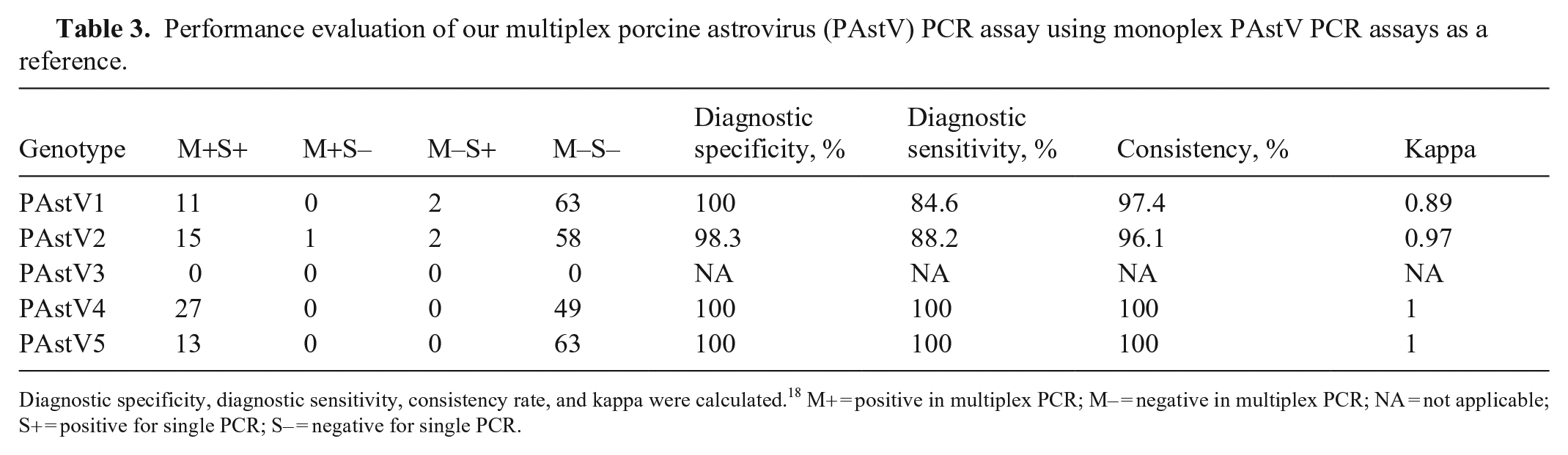
Diagnostic specificity, diagnostic sensitivity, consistency rate, and kappa were calculated. 18 M+ = positive in multiplex PCR; M– = negative in multiplex PCR; NA = not applicable; S+ = positive for single PCR; S– = negative for single PCR.
Discussion
The sensitivity of multiplex PCR assays usually decreases as the number of intended targets in the system increases.31,32 We found that the LOD of PAstV1–5 by the multiplex PCR assay was 5 × 102 to 5 × 103 copies/μL, which was 10–100-fold higher than that of the single PCR assay. This improvement may be attributed to factors that influence the efficiency of multiplex PCR assay, such as competition between reactions and appropriate annealing temperature. 7 A multiplex RT-PCR assay has been reported to detect the 5 PAstV genotypes, 15 but the sensitivity of the assay was lower, with a LOD of 10 pg/μL (~5 × 105 copies/μL), compared with the enhanced sensitivity of our multiplex PCR method. Similarly, 2 sets of multiplex real-time fluorescence quantitative PCR panels were established for these 5 genotypes with a lower LOD of single copies 30 ; however, our multiplex PAstV PCR assay has a significant advantage of low cost.
Our multiplex PCR and monoplex PCR assays on 76 field specimens had a consistency of 96–100% for the 5 genotypes, indicating the usefulness of our multiplex PAstV PCR in clinical investigations. In previous reports, the prevalence of PAstV was 6–90%, and was usually higher in North America and Europe, including Canada (79.2%), the United States (64%), and Italy (67.4%)16,29,34; it was lower in Asia: Thailand (6.5%), South Korea (19.4%), and India (17.6% and 31.8% in 2018 and 2021, respectively).9,11,12,14 The high overall positive rate of PAstV (47.4%) in our study, 56.4% in Guangxi, 21 and 46.3% in Hunan, 30 indicates that PAstV can be found frequently in pigs in China. Our findings of the positive rates of PAstV1, PAstV2, PAstV4, and PAstV5 of 11 of 76 (14.5%), 16 of 76 (21.1%), 27 of 76 (35.5%), and 13 of 76 (17.1%), respectively, are similar to reports indicating that PAstV2 and PAstV4 were the genotypes detected most frequently in domestic pigs, regardless of whether 2 genotypes14,17 or 5 genotypes3,21,29 were prevalent in the respective pig populations. Although the positive rate of PAstV detection is high in many pig populations, infection by PAstVs is usually latent and can be easily missed under production conditions.15,21 Our results also indicate that producers and diagnostic units should be vigilant about the prevalence of PAstVs.
We detected ≥2 PAstV genotypes in 20 of 76 (26.3%) field samples. The simultaneous presence of multiple PAstV genotypes in a single individual may increase the opportunity for recombination events and genetic diversity, leading to more complex pathogenesis outside the intestinal tract.21,27,33 Therefore, genotyping of PAstVs is necessary in recombination and genetic analyses.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387221145329 – Supplemental material for Multiplex gel-based PCR assay for the simultaneous detection of 5 genotypes of porcine astroviruses
Supplemental material, sj-pdf-1-vdi-10.1177_10406387221145329 for Multiplex gel-based PCR assay for the simultaneous detection of 5 genotypes of porcine astroviruses by Qiuya Zhang, Qin Liu, Tanja Opriessnig, Dan Wen, Keda Gu and Yonghou Jiang in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was supported by the Science and Technology Bureau of the Zhejiang Province of China (2018C37051) and the Natural Science Foundation in Zhejiang (LGC21C180002).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
