Abstract
Immunization of calves with Anaplasma centrale is used to prevent acute anaplasmosis caused by A. marginale. Natural and vaccine-acquired immunity is detected through serologic tests based primarily on A. marginale recombinant major surface protein 5 (MSP5m) because it has 91% identity with MSP5 from A. centrale (MSP5c). We developed a displacement, double-antigen, sandwich ELISA (ddasELISA) to detect antibodies against A. marginale or A. centrale. For ddasELISA validation, we analyzed serum samples positive for antibodies against Anaplasma spp. from cattle naturally infected with A. marginale (n = 300) or vaccinated with A. centrale (n = 255). Species-specific nested PCR (nPCR) assays were used to confirm infection. The optical density (OD) values obtained from antibodies directed at unique epitopes of A. marginale (ODAm) or A. centrale (ODAc) were used in the formula ODAm/ODAc. If the derived ratio was >0.38, the serum sample was considered positive for antibodies against A. marginale, with 98.9% sensitivity and 98.0% specificity. In a field evaluation, we analyzed 702 Anaplasma spp. antibody–positive serum samples from 34 herds by ddasELISA and nPCR; 571 were classified by ddasELISA as A. marginale–infected or A. centrale–vaccinated, with 84% agreement (κ = 0.70) between ddasELISA and nPCR. Our results indicate that ddasELISA could be used as a cost-effective alternative to molecular techniques to confirm infection with A. marginale in countries in which prevention is based on vaccination with A. centrale.
Keywords
Anaplasma marginale, an intraerythrocytic bacterium transmitted to susceptible cattle biologically by ticks or mechanically by biting flies and fomites, is the main causative organism of bovine anaplasmosis.2,7,18 Acute anaplasmosis affects mostly adult cattle and is characterized by severe anemia (from destruction of erythrocytes), abortion, weight loss, reduced milk production, and death. Cattle recovered from acute disease remain as carriers that serve as reservoirs for transmission of this pathogen to other animals. 4 Immunization with live organisms of A. centrale, a species closely related to A. marginale, is used to prevent acute anaplasmosis in several regions worldwide. Vaccination with A. centrale causes only mild clinical signs and does not prevent infection with A. marginale, but reduces disease severity and prevents death. 1
Molecular methods, mainly PCR, are used to accurately differentiate cattle infected with A. marginale from those vaccinated with A. centrale because the available serologic tests do not differentiate between the 2 species.6,11 Commercial ELISA kits are based on the A. marginale recombinant major surface protein 5 (MSP5m), an immunodominant antigen that is highly conserved within and between Anaplasma spp. Thus, these ELISAs also detect antibodies generated by A. centrale 10 and A. ovis. 8 Species-specific epitopes in MSP5m and MSP5 of A. centrale (MSP5c) have been demonstrated using a competitive ELISA (cELISA) 13 and a double-antigen, sandwich ELISA (dasELISA). 16 In countries in which anaplasmosis is prevented through vaccination with A. centrale, there is a need for methods to differentiate A. marginale carriers from animals immunized with A. centrale. The inclusion of a competitive displacement step in a dasELISA would enhance the specificity of this assay for A. marginale or A. centrale antibody detection.
Our aim was to develop a displacement dasELISA (ddasELISA) to identify antibodies against A. marginale or A. centrale, detecting epitopes that are not shared between MSP5m and MSP5c. We based the assay on 2 ELISA determinations performed on samples that were seropositive for Anaplasma spp., one to detect antibodies against specific epitopes of MSP5m and the other to detect antibodies against specific epitopes of MSP5c. The ratio between MSP5m- and MSP5c-specific antibodies would allow us to differentiate between A. marginale–infected and A. centrale–vaccinated cattle.
Recombinant antigens for dasELISA and ddasELISA were produced. 13 Briefly, genomic DNA (gDNA) was obtained from 1 mL of blood from A. marginale– or A. centrale–infected cattle by the phenol–chloroform method. This gDNA was used as a template for PCR amplification of the DNA region encoding residues 28–210 (MSP5 full length without transmembrane region) with a 6-histidine tag at the C-terminus of the protein MSP5m (tMSP5m) or MSP5c (tMSP5c). The proteins were expressed in Escherichia coli BL21 RIL (DE3) pLysS cells (Novagen) and purified by pseudoaffinity with nickel–nitrilotriacetic acid agarose (Qiagen) from the soluble fraction. 13 The purity of recovered recombinant proteins evaluated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) was >95%. Molar concentration in pure samples was calculated by absorbance at 280 nm using a molar extinction coefficient (ε280nm) equal to 8,940 M-1cm-1 for tMSP5m or 10,430 M-1cm-1 for tMSP5c. 16 Purified tMSP5m and tMSP5c were biotinylated to obtain tMSP5m-biotin and tMSP5c-biotin, as described. 16
dasELISA was performed as described 16 ; ddasELISA was performed as follows. Microplate wells 1–6 and 7–12 were coated overnight at 4°C with 100 µL of 0.8 µg/mL of tMSP5m or tMSP5c in PBS (50 mM sodium phosphate, 150 mM NaCl, pH 7.2), respectively. After 2 washes with PBS, blocking buffer (PBS + 10% fat-free dried milk, 300 µL/well) was added, and the plates were incubated at 25°C for 1 h. After washing 3 times with PBS containing 0.05% Tween 20 (PBST), the undiluted samples and controls (100 µL/well) were added in parallel in the wells 1,7; 2,8; 3,9; 4,10; 5,11; and 6,12; the plates were incubated at 25°C for 1 h in a shaking incubator at 100 rpm. The plates were washed 5 times with PBST, and 100 µL of tMSP5m-biotin (1 µg/mL) plus tMSP5c (10 µg/mL), or tMSP5c-biotin (1 µg/mL) plus tMSP5m (10 µg/mL) in PBST + 10% fat-free dried milk, were added to wells 1–6 or 7–12, respectively. After 1-h incubation (25°C, 100 rpm), the plates were washed 5 times with PBST, and bound protein–biotin was detected by incubating the plate with 100 μL/well of streptavidin–HRP (Jackson ImmunoResearch), diluted 1:500 in PBST + 10% fat-free dried milk, for 1 h (25°C, 100 rpm). After the final wash (5 times with PBST), the chromogenic substrate 1 mM 2,2′-Azino-bis [3-ethylbenzothiazoline-6-sulfonic acid]-diammonium salt (ABTS; MilliporeSigma) was added, diluted in 0.05 M sodium citrate, pH 4.5, 0.015% v/v H2O2 (100 μL/well). Then the plates were incubated in the dark for 30 min (25°C, 100 rpm). The optical density (OD) was measured at 405 nm with an ELISA plate reader (Labsystems Multiskan FC; Microlat). Samples with results of OD405nm <0.2 in wells analyzed with both tMSP5m and tMSP5c antigens were not included in the analysis because the antibodies present in the samples were displaced by the 2 antigens (tMSP5m and tMSP5c). Results were expressed as the ratio between antibodies specific for tMSP5m and for tMSP5c (ODAm/ODAc), using the following formula: ODAm/ODAc ratio = (sample × OD in well 1A)/(sample × OD in well 7 A); exemplified for sample x, analyzed in wells 1 A and 7 A.
Positive DNA and serum controls specific for A. marginale (CAm+) or A. centrale (CAc+) were obtained from blood samples of 2 calves, each experimentally infected with 1 of the 2 species. Blood samples were collected aseptically with and without 5% citrate as anticoagulant. Serum and whole blood samples were distributed in aliquots and kept at −20°C until use. All procedures were approved by the Animal Care Committee of the Faculty of Veterinary Sciences, National University of Littoral (protocol 243/15).
For validation assays, we used serum and blood samples from 300 cows from herds not vaccinated with A. centrale that were born and raised in a highly endemic area of anaplasmosis in Argentina, and 255 8–10-mo-old calves from herds free of A. marginale infection that had been vaccinated with 107 A. centrale–parasitized RBCs 2–4 mo earlier. The status of Anaplasma infection was determined by nested PCR (nPCR) assays specifically validated for the 2 Anaplasma spp., 9 using 10 ng of gDNA, purified from 1 mL of blood by the phenol–chloroform method, per reaction. Antibodies against Anaplasma spp. were detected by dasELISA in 294 and 255 samples from the infected and vaccinated cattle, respectively. Results were expressed as percent positivity (%P) according to the following formula: %P = 100% (sample OD/positive control OD).
The x̄ ± SD results were 94.0 ± 12.9 %P and 91.8 ± 15.0 %P for infected and vaccinated cattle, respectively (Fig. 1A). Six positive samples for A. marginale by nPCR were negative by dasELISA. This result is expected during early primary infections, given that under experimental conditions, the seroconversion by cELISA is detected 16–27 d after inoculation, whereas molecular techniques are able to detect A. marginale DNA from 2 d post-inoculation.3,5,6
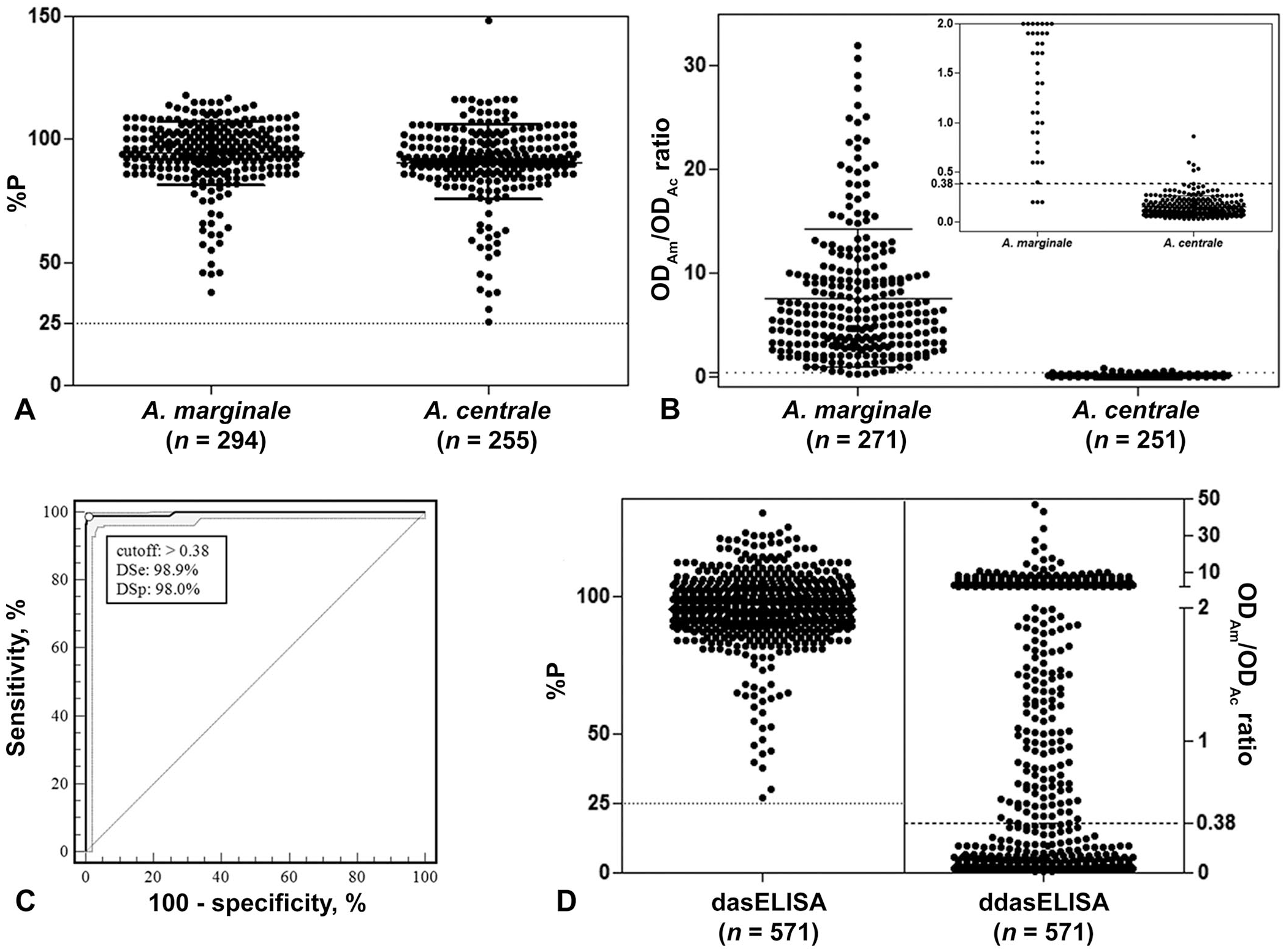
Validation of a displacement, double-antigen, sandwich ELISA (ddasELISA; A–C) and its field evaluation (D).
Anaplasma spp. antibody–positive samples of the 2 groups (n = 549) were analyzed by ddasELISA. Of those, 4.9% (n = 27; 17 A. marginale–infected and 10 A. centrale–vaccinated) samples had an OD405nm of <0.2 for both tMSP5m and tMSP5c antigens. These 27 samples were not included in the analysis; the OD405nm values of <0.2 are close to the background noise of the method and cannot provide reliable results for the ODAm/ODAc ratio. For ddasELISA, the ODAm/ODAc ratio of 522 samples had x̄ ± SD results of 7.72 ± 6.70 for cattle infected with A. marginale and 0.14 ± 0.11 for those vaccinated with A. centrale (Fig. 1B). The optimal cutoff point and the diagnostic sensitivity (DSe) and diagnostic specificity (DSp) were established considering samples with results of A. marginale nPCR–positive/A. centrale nPCR–negative as A. marginale positive, and samples with results of A. marginale nPCR–negative/A. centrale nPCR–positive as A. marginale negative, via receiver operating characteristic (ROC) analysis (MedCalc v.13.0; MedCalc Software). The cutoff point for ddasELISA was >0.38 with 98.9% (95% CI: 96.8–99.8%) DSe and 98.0% (95% CI: 94.4–99.4%) DSp for species-specific A. marginale antibody detection (Fig. 1C). Thus, the bovine samples with a ODAm/ODAc ratio >0.38 were classified as being infected with A. marginale, and those with a ratio ≤ 0.38 were classified as being vaccinated with A. centrale. Eight samples were misclassified, of which 5 (62.5%) were vaccinated with A. centrale and 3 (37.5%) were infected with A. marginale.
For the field evaluation, 702 serum and blood samples from cattle from 34 herds received at the Immunology and Parasitology Laboratory of the Institute of Agricultural Technology (INTA-Rafaela, Argentina) and positive for Anaplasma spp. antibodies by dasELISA were analyzed by ddasELISA and nPCR. 9 A total of 131 (19%) samples with the ODAm and ODAc of <0.2 were excluded from further analysis. The nPCR results showed that, of the excluded samples, 27 were A. marginale– and A. centrale–negative, 68 were A. marginale–positive and A. centrale–negative, 30 were A. marginale–negative and A. centrale–positive, and 6 were A. marginale– and A. centrale–positive.
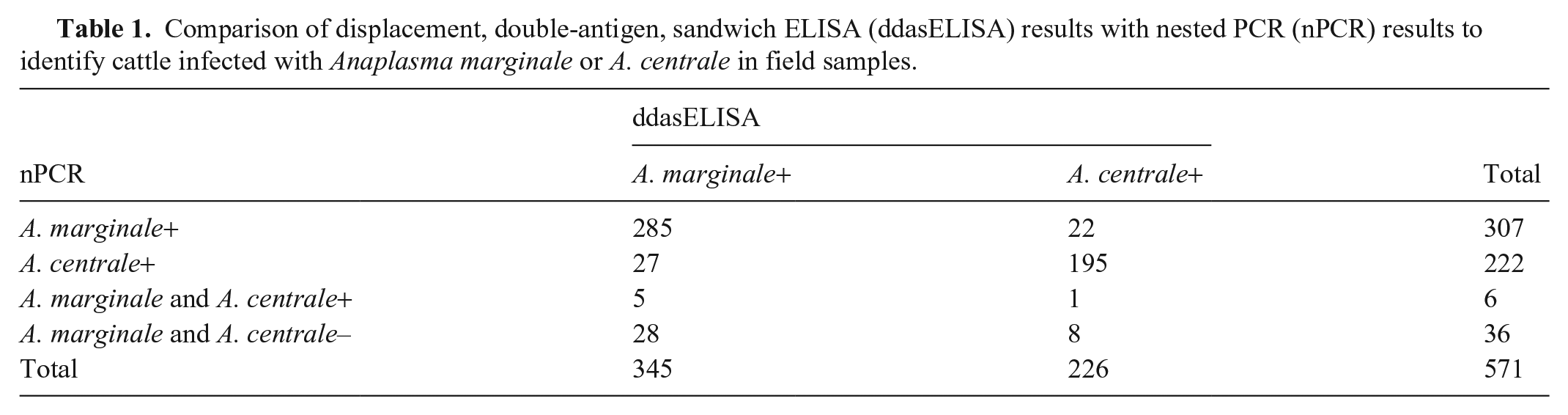
For the 571 field samples with an OD ≥0.2 in at least 1 well, the agreement between the ddasELISA and nPCR results was 84% and the kappa (κ) coefficient (MedCalc v.13.0) was 0.70 (95% CI: 0.635–0.754; Fig. 1D). Of the 313 A. marginale nPCR–positive samples and of the 228 A. centrale nPCR–positive samples, 93% (n = 290) and 86% (n = 196) were consistent with the ddasELISA results (Table 1). Thirty-six nPCR-negative samples tested positive for antibodies against A. marginale (n = 28) or A. centrale (n = 8) by ddasELISA.
Comparison of displacement, double-antigen, sandwich ELISA (ddasELISA) results with nested PCR (nPCR) results to identify cattle infected with Anaplasma marginale or A. centrale in field samples.

Discordant results have been observed between molecular and serologic methods in several epidemiologic studies, with concordances of 54–95% reported between the techniques.5,9,12,17 The authors suggest that discrepancies between positive results by serology (cELISA), but negative by molecular methods might be the result of recently resolved Anaplasma infections, with persistence of serum antibodies or a cross-reactivity with other related organisms in the region. The detection of antibodies by our new technique should reduce the possibility of cross-reactivity among Anaplasma spp. through the displacement of antibodies that recognize conserved regions of the MSP5 proteins. Therefore, in our study, cattle with nPCR-negative and dasELISA-positive results could be explained by the elimination or reduction of parasitemia, either naturally (cycles of parasitemia) or through the administration of antibiotics.14,15 We did not explore these aspects because we did not know if the cattle had been treated with antibiotics nor did we know the time elapsed between infection and sampling.
The expression of results as an ODAm/ODAc ratio leads to the classification of samples into only 2 categories, whether they contain specific antibodies against A. marginale or A. centrale, which could be a limitation in detecting coinfections by ddasELISA. Our nPCR identified 12 cattle with A. marginale and A. centrale coinfections; 6 were excluded from the analysis and 5 and 1 were classified by ddasELISA as infected with A. marginale and vaccinated with A. centrale, respectively. Therefore, when there was coinfection, 50% of the results could not be processed because none of the wells yielded OD405nm ≥0.2, indicating a prevalence of antibodies directed to epitopes conserved between MSP5m and MSP5c that were displaced by 2 antigens in the ddasELISA. In cases in which the samples could be classified, the ratios were close to the cutoff point (0.27, 0.39, 0.53, 0.57, 0.79, 2.81). Another strategy for conducting the ddasELISA or for expression of results should be evaluated for the detection of both Anaplasma spp. in the presence of coinfection. The small number of cattle with coinfections detected did not allow us to evaluate this aspect.
ddasELISA could be used as a cost-effective alternative to molecular techniques to confirm infection with A. marginale in countries in which prevention is based on vaccination with A. centrale. The new assay would also be useful when a laboratory receives only serum samples (instead of whole blood) and a species determination by PCR cannot be performed. Furthermore, the assay could be used in disease control programs to monitor the prevalence of A. marginale in a region or a herd. Although it was possible to classify only 81% of the samples with antibodies against Anaplasma spp. and the detection of coinfections was not possible, to our knowledge, ddasELISA is the first reported test capable of detecting species-specific antibodies against Anaplasma and could be a starting point for the future development of other ELISAs based on MSP5 peptides.
Footnotes
Acknowledgements
We thank Paola Amherdt for technical assistance in sample processing.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was supported by INTA, Asociación Cooperadora INTA Rafaela (TCP 426100), ASATEC (IO-2019-130), and Agencia Nacional de Promoción Científica y Tecnológica (PICT2013-0369, PICT2019-01498).
