Abstract
We made 2 Z-based in situ hybridization (ISH) probes for the detection of rabbit hemorrhagic disease virus 2 (RHDV2; Lagovirus GI.2) nucleic acid in formalin-fixed, paraffin-embedded tissues from European rabbits (Oryctolagus cuniculus) that had died during an outbreak of RHD in Washington, USA. One probe system was made for detection of negative-sense RNA (i.e., the replicative intermediate RNA for the virus), and the other probe system was constructed for detection of genomic and mRNA of the virus (viral mRNA). Tissue sets were tested separately, and the viral mRNA probe system highlighted much broader tissue distribution than that of the replicative intermediate RNA probe system. The latter was limited to liver, lung, kidney, spleen, myocardium, and occasional endothelial staining, whereas signal for the viral mRNA was seen in many more tissues. The difference in distribution suggests that innate phagocytic activity of various cell types may cause overestimation of viral replication sites when utilizing ISH of single-stranded, positive-sense viruses.
Z-based in situ hybridization (ISH) methods, commonly known as RNAscope, have proven to be effective tools for in situ detection of viral RNA, cancer biomarkers, and endogenous gene expression.1,4,7 Developed in 2011, this novel ISH method allows for the detection of RNA that may be in shorter segments than can traditionally be found with ISH on formalin-fixed, paraffin-embedded (FFPE) tissues. 9 Traditional ISH methods such as riboprobe require probe lengths often >500 nucleotides, and short probes can lack sufficient amplification potential for detection of low-abundance RNA molecules. 10 The utilization of up to 20 pairs of Z-shaped probes, which when matched to their counterparts span 36–50 bases, markedly reduces the negative impact of RNA-crosslinking from formalin fixation. Additionally, each double-Z probe pair has up to 400 potential reaction sites with the label probe, allowing a single strand of RNA to have up to 8,000 label probe reaction sites. The capacity for visual detection of a single viral genome makes Z-based ISH an ideal tool for viral pathogenesis studies. 9
Viral pathogenesis studies often rely on cell culture to elucidate specific cell types supportive of replication in vivo. To date, rabbit hemorrhagic disease virus 2 (RHDV2; Caliciviridae, Lagovirus GI.2), a pathogenic calicivirus of lagomorphs, has not been isolated successfully in cell culture, requiring all pathogenesis studies to be conducted on tissues of the host animal, rabbits. Consequently, the common modalities currently available to determine tissue tropism for RHDV2 utilize FFPE tissues with techniques including immunohistochemistry (IHC), traditional riboprobe-based ISH, or RNAscope.3,5 Our laboratory recently utilized this RNAscope technique to determine the tissue distribution of RHDV2, which yielded abundant signal for positive-sense RNA in various organ systems. Tissue distribution was very wide. 6 To discern which tissues were the primary viral target, and which tissues might just be participating phagocytically in limited viral replication, we compared positive- and negative-sense RNAscope ISH.
As described previously, 6 RNAscope ISH detection of RHDV2 positive-sense viral RNA was validated utilizing FFPE sections of RHDV2-positive tissues from 3 rabbits from the 2019 outbreak in Washington, USA (tissues kindly provided by the Washington Animal Disease Diagnostic Laboratory, Washington State University, Pullman, WA, USA). Fresh liver from each rabbit was confirmed positive for the virus via RT-PCR at the Foreign Animal Disease Diagnostic Laboratory (FADDL), USDA-APHIS. In our current study, tissues evaluated via ISH for viral mRNA and negative-sense replicative intermediate RNA included: liver, gallbladder, spleen, tongue, trachea, esophagus, thyroid, lung, diaphragm, heart, stomach, pancreas, duodenum, jejunum, ileum, cecum, colon, adrenal glands, kidney, urinary bladder, reproductive tract, and brain.
For the detection of viral replicative intermediate RNA, a probe matching a portion of the VP60 capsid protein mRNA probe (V-RHDV-vp60-sense-C2) was generated from GenBank KP862933.1 using a chromogenic, manual, 2.5HD duplex detection kit (ACD, Bio-Techne); manufacturer’s protocols were followed. Briefly, 4-µm sections of FFPE tissues were mounted, deparaffinized with xylene and alcohol, and rehydrated within the HybEZ oven (ACD, Bio-Techne). After rehydration, a 3% hydrogen peroxide solution was applied to quench endogenous peroxidase activity, 1× target retrieval reagent was applied to reduce cross-linking from fixation, then RNAscope protease plus was applied to digest proteins that may block target sequences. Hybridization was then performed with either the sense or the anti-sense-RHDV2-VP60 probe, and detection with the alkaline phosphatase enzymatic step, utilizing red chromogen.
As expected, for both the viral mRNA probe and the probe to detect replicative intermediate RNA, there was strong, bright-red cytoplasmic signal in hepatocytes for all rabbits (Figs. 1, 2). For both probes, biliary epithelium, vascular smooth muscle, and stromal fibroblasts did not have discernable signal. Heavily autolyzed sections of liver generally had minimal signal. In the spleens of all rabbits, there was strong signal for viral mRNA within cells morphologically consistent with macrophages throughout both lymphoid and sinus regions (Fig. 3), with moderate signal for replicative intermediate RNA in the same areas and cells (Fig. 4).
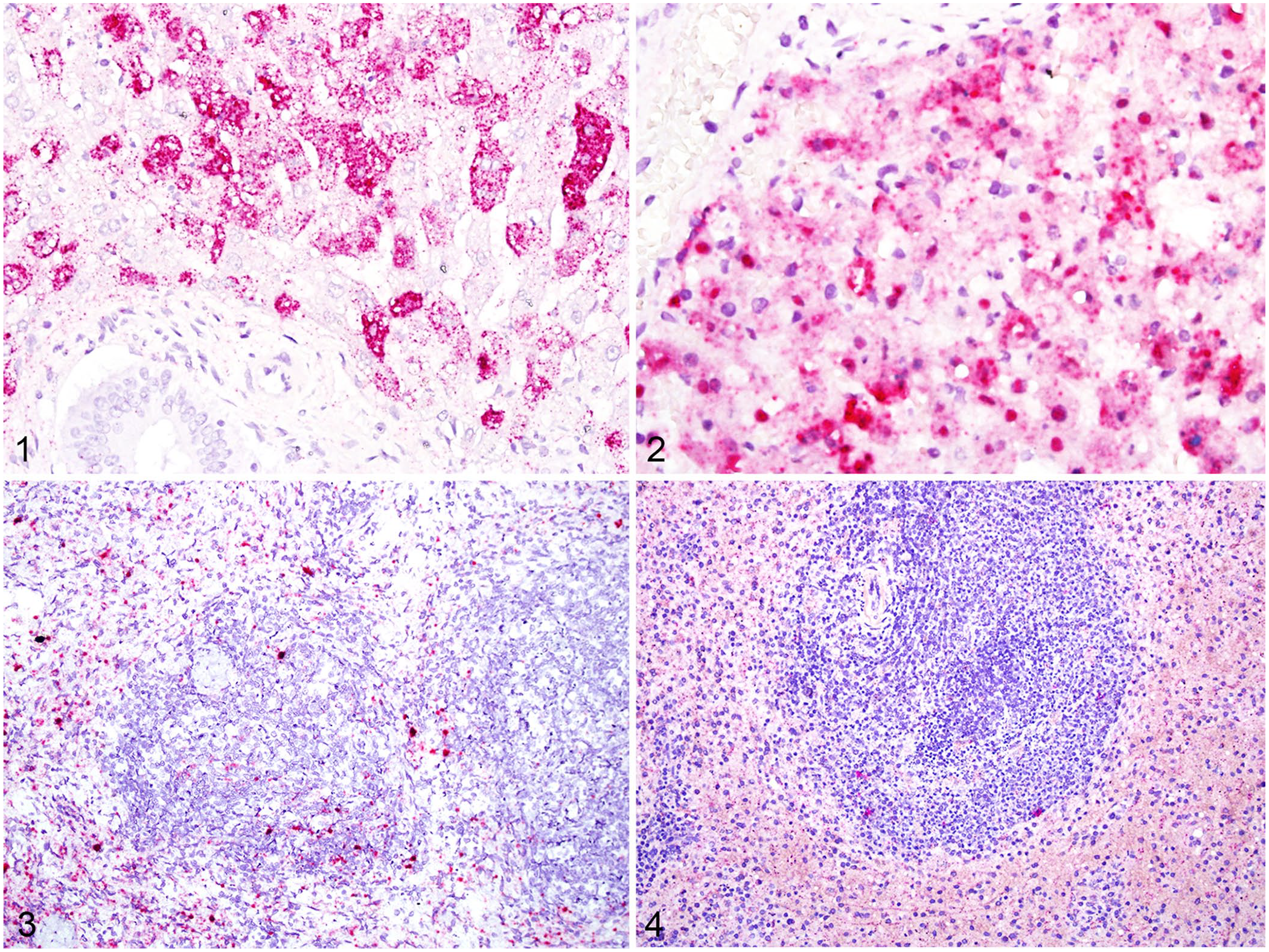
RNA detection of viral mRNA and replicative intermediate RNA in liver and spleen by RNAscope in situ hybridization.
Many tissues had signal using both probes, but with marked differences in intensity when comparing the signal for viral mRNA versus the signal for replicative intermediate RNA. There was extensive signal for viral mRNA throughout the interstitium of the lungs, which could be either endothelium or pulmonary intravascular macrophages (Fig. 5). However, there was markedly less signal for viral replicative intermediate RNA with a similar distribution (Fig. 6). The vascular endothelium within the myocardium had prominent cytoplasmic signal for viral mRNA, but occasionally spilling into adjacent cardiomyocytes. Signal for viral replicative intermediate RNA was less intense but had similar distribution to positive-sense RNA. Although there was significant signal for viral mRNA in the glomeruli of 3 rabbits (Fig. 7), there was moderate signal for replicative intermediate RNA within glomeruli for 2 rabbits, and only occasional punctate signal for the third. Tissues that contained endothelial cytoplasmic signal for both viral mRNA and replicative intermediate RNA included the cerebrum (Fig. 8), cerebellum, glandular stomach, duodenum, jejunum, ileum, and colon.
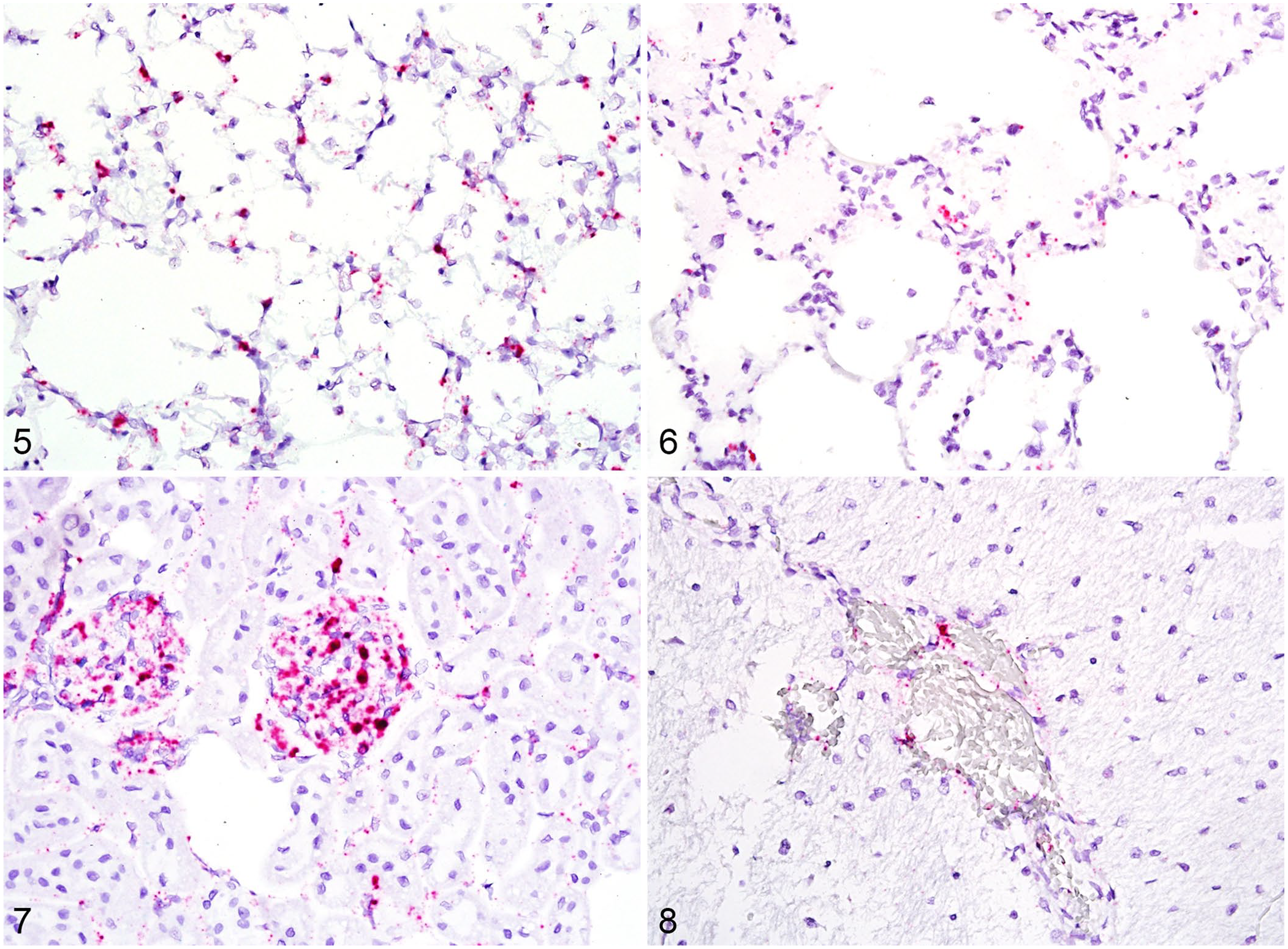
RNA detection of viral mRNA and replicative intermediate RNA in lung, viral mRNA in kidney, and replicative intermediate RNA in the cerebrum.
The following tissues had some signal for viral mRNA but complete absence of signal for replicative intermediate RNA: adrenal gland zonae glomerulosa and fasciculata, thyroid, pancreas, and thymus. The colon of one rabbit had moderate cytoplasmic signal for viral mRNA within enterocytes, and an absence of signal for replicative intermediate RNA.
Our results indicate a distinct difference in tissue distribution for detectable viral particles and viral replicative intermediate RNA, suggesting that detection of viral mRNA does not necessarily correlate with sites of viral replication, and care should be taken with interpretation of ISH for targets for viral replication. Given that the virus is known for its tropism for hepatocytes, it is unsurprising to find both viral mRNA and replicative intermediate RNA within the liver. In all of the other tissues examined, endothelium and macrophages often had viral mRNA and also some replicative intermediate RNA. Phagocytic activity could account for the abilities to take up the virus actively and allow some replication. 8 Although replicative intermediate RNA was demonstrated in the spleen, lung, myocardium, kidney, and endothelium of various tissues, the signal was significantly reduced compared to viral mRNA. This suggests that these tissues may support limited viral replication but are not primary sites as is the liver. The presence of viral mRNA but lack of replicative intermediate RNA within gastrointestinal epithelium may be the result of pinocytosis of intraluminal viral particles shed through the biliary system. Although the mechanism for RHDV2 shedding in feces has not been determined specifically, biliary excretion seems likely given that this is a common mechanism for other hepatotropic viruses. 2
Overall, RNAscope is a highly specific, highly sensitive tool that can further our understanding of viral pathogenesis and has utility as a replacement for IHC, which is less reliable for detection. This may prove useful for preliminary diagnosis wherein RHDV2 is suspected and PCR must be performed at a separate laboratory. However, our results indicate that sites of viral replication do not necessarily correlate with detection of viral mRNA, and overestimation of viral replication sites is possible when only utilizing a single probe set.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was fully funded through the United States Department of Agriculture Animal Plant Health Inspection Service’s National Bio- and Agro-defense Facility Scientist Training Program.
