Abstract
Formalin-fixed, paraffin-embedded tissues from European rabbits (Oryctolagus cuniculus) that succumbed to rabbit hemorrhagic disease virus 2 (RHDV2; Lagovirus GI.2) during the 2019 outbreak in Washington, USA, were utilized for in situ hybridization via RNAscope (ACDBio). This detection method was both sensitive and specific, with no staining in tissues from RHDV- (Lagovirus GI.1) and RHDV2-negative rabbits, and only slight background staining of RHDV-positive rabbits; RHDV2-positive tissues had bright-red cytoplasmic staining. Although much of the viral mRNA detection was consistent with previously described antigen detection via immunohistochemistry of the liver, lungs, and spleen, there was also significant glomerular staining in the kidneys, and endothelial staining within blood vessels of almost all organs. We validated the RNAscope technique for detection of RHDV2 mRNA in formalin-fixed, paraffin-embedded tissues, with increased sensitivity from previous techniques, and identified additional affected cell types that may contribute to the understanding of pathogenesis.
Rabbit hemorrhagic disease virus 2 (RHDV2; Caliciviridae, Lagovirus, Lagovirus GI.2) causes severe hepatic necrosis, disseminated intravascular coagulation, and death. Like other caliciviruses, it is a nonenveloped, positive-sense, single-stranded RNA virus with an icosahedral structure. It was first discovered in France, in 2010, as antigenically and phylogenetically distinct from rabbit hemorrhagic disease virus (RHDV1; Lagovirus GI.1).5,6 It has since spread throughout Europe, Asia, Oceania, and Africa, and is now causing a high-mortality outbreak in North America.4,7,9,16 Unlike RHDV1, which only affects European rabbits (Oryctolagus cuniculus), RHDV2 has caused mortality in Italian hares (syn. Corsican hare; Lepus corsicanus), Sardinian Cape hares (Lepus capensis mediterraneus), European brown hares (Lepus europaeus), mountain hares (Lepus timidus), desert cottontails (Sylvilagus audubonii), black-tailed jackrabbits (Lepus californicus), antelope jackrabbits (Lepus alleni), and mountain cottontails (Sylvilagus nuttallii).2,13,14,16
Although there have been numerous studies describing the pathogenesis of RHDV1, studies on the pathogenesis of RHDV2 are relatively few.8,11,12 To date, neither virus has been grown in tissue culture, hence identification in situ remains the best means of evaluating pathogenesis. There are many methods for detecting RHDV through serum antibodies (hemagglutination inhibition, ELISA), serum or tissue antigen (hemagglutination, western blot, antigen ELISA, lateral flow immunoassay, Staphylococcus protein A co-agglutination test), viral RNA within tissues or bodily secretions (reverse-transcription PCR [RT-PCR], real-time RT-PCR [RT-rtPCR]), simultaneous viral RNA and antibody detection (Luminex xTAG and xMAP), and viral particles from tissue (negative staining electron microscopy), but some of these methods require destruction of the tissue and do not specify infected cell type. Each of these methods has its utility; however, for the most definitive detection, RT-rtPCR with TaqMan probes remains the assay of choice. For in situ detection of RHDV, traditional in situ hybridization (ISH) for RNA detection with digoxygenin-labeled probes has been developed for RHDV, but not RHDV2. In general, ISH techniques are more sensitive and specific than immunohistochemistry (IHC), but high cost and complicated methods have limited their use. 1
To date, IHC for RHDV2 relies on a pan-lagovirus antibody that is not specific for RHDV2. A study utilizing IHC demonstrated reliable detection of viral capsid antigen in the liver and spleen, whereas detection of antigen in bone marrow, lung, and kidneys was less consistent; viral antigen was found in the liver of 4 of 4 rabbits, spleen of 3 of 4, lung of 3 of 4 (small amounts), bone marrow of 1 of 2, and the kidney of 2 of 4 (small amounts within glomeruli and adjacent vessels). 13 When comparing IHC results to RT-rtPCR, the detection threshold for IHC was ~107 viral copies per mg tissue. Viral RNA was detected via RT-rtPCR throughout the gastrointestinal tract, heart, thymus, brain, and lymph nodes; however, IHC was inadequate to detect viral antigen in these tissues. 12 To better understand the distribution of RHDV2 within tissues, particularly those with viral loads that are significant but not detected via IHC, we performed ISH of viral mRNA for the capsid protein (VP60). We aimed to validate a highly sensitive technique to detect RHDV2 viral mRNA and to study the distribution of viral mRNA in different tissues.
Formalin-fixed, paraffin-embedded (FFPE) sections of RHDV2-positive rabbit tissues were kindly provided by the Washington Animal Disease Diagnostic Laboratory (Washington State University, Pullman, WA, USA). Four O. cuniculus that succumbed to RHDV2 during the 2019 outbreak in the state of Washington 15 were autopsied and confirmed positive via RT-PCR of fresh liver at the Foreign Animal Disease Diagnostic Laboratory (FADDL), USDA-APHIS. Sections of brain, tongue, trachea, esophagus, thyroid glands, lungs, diaphragm, heart, stomach, pancreas, duodenum, jejunum, ileum, cecum, colon, rectum, spleen, liver, gallbladder, adrenal glands, kidney, urinary bladder, reproductive tract, skin, lymph nodes, eyes, and skeletal muscle were routinely fixed in 10% neutral-buffered formalin and embedded in paraffin wax. As control specimens, FFPE liver sections from experimentally infected O. cuniculus confirmed as negative, RHDV1 positive, or RHDV2 positive via RT-PCR were kindly provided by FADDL. Two negative controls (mock-inoculated with PBS), 1 RHDV-positive rabbit, and 3 RHDV2-positive rabbits were utilized for assay validation. H&E staining was performed on 4-µm sections of each prepared block to evaluate histopathologic changes associated with RHDV2 (Fig. 1).
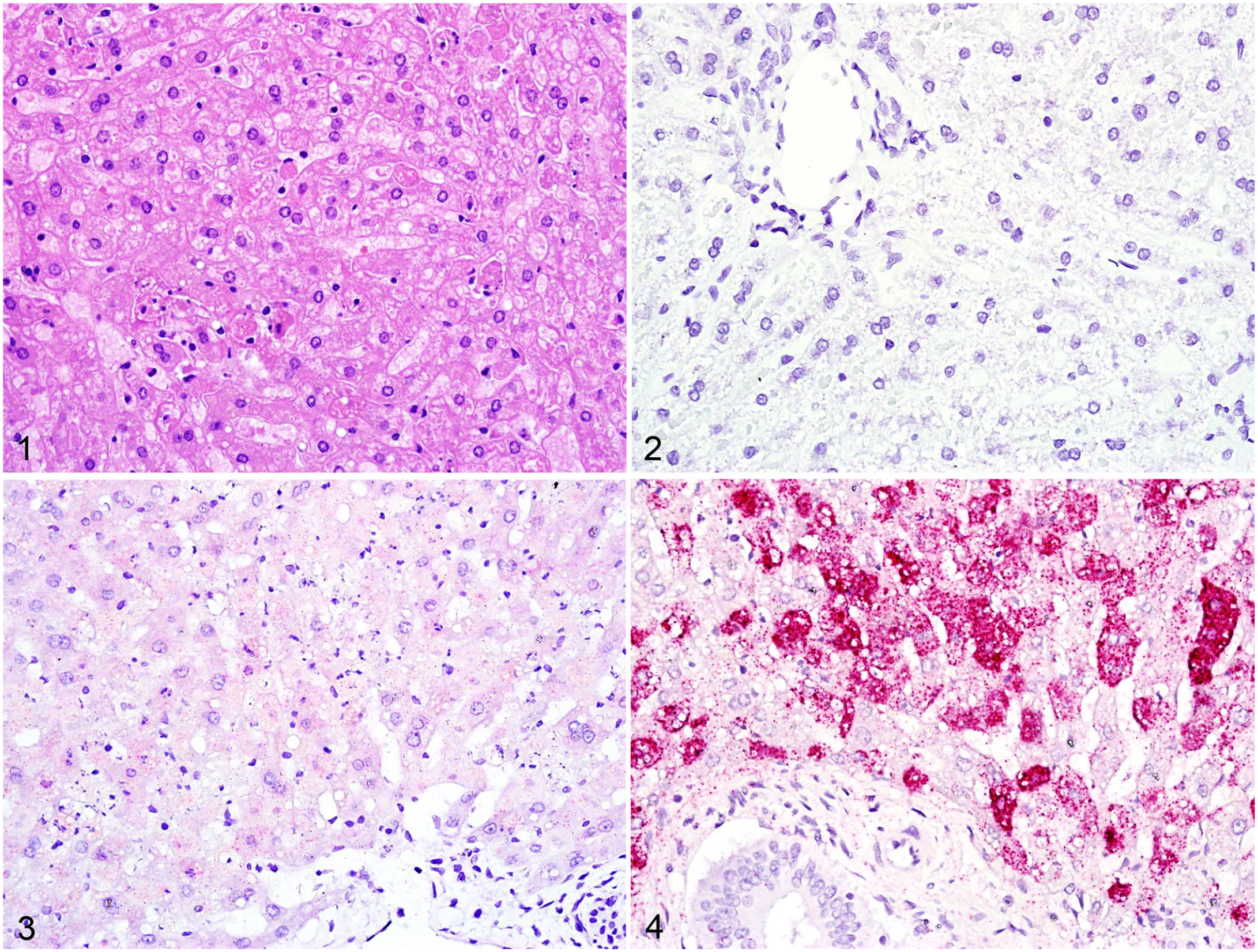
Rabbit liver sections with H&E staining or RNAscope in situ hybridization.
For the RHDV2 riboprobe, FFPE sections of RHDV2-positive liver were prepared following the manufacturer’s protocol for RNA extraction (RNeasy FFPE kit [Buffer PKD, DNase booster buffer, Buffer RBC, RNeasy MinElute spin column, Buffer RPE]; Qiagen). Briefly, four 5-μm thick sections from the tissue block were combined in a microcentrifuge tube, deparaffinized, incubated with digestion buffer (Buffer PKD), centrifuged, incubated with proteinase K, the resulting digested material transferred to a new tube, incubated on ice, centrifuged, and the supernatant saved. The supernatant was incubated with DNase booster buffer, then Buffer RBC and ethanol to optimize binding conditions, and then centrifuged through a RNeasy MinElute spin column; the flow-through was discarded. Buffer RPE was spun through the column twice to wash away remaining contaminants, the collection tube was replaced, and then 14 μL of RNase-free water was spun through the column to elute the extracted RNA. RNA content for each sample was then tested via spectrophotometer (Thermo Fisher). A first-strand cDNA template was synthesized from 1 μg of sample RNA (Superscript III reverse transcriptase, random primers; Invitrogen). Using the cDNA template and primers from RHDV2 VP60 mRNA (Integrative DNA Technology, GenBank KP862933.1), formation of 385-, 242-, and 128-bp fragments were each attempted, with only the 128-bp fragment successful. Given that this was an insufficient length for riboprobe formation, this portion of the experiment was terminated. All procedures were performed using RNase-free equipment and RNase decontamination of workstations.
For RNAscope (ACDBio) ISH, 2 probes were generated based on the VP60 capsid protein gene, using GenBank sequence KP862933.1 (ACDBio, probe V-RHDV-vp60). All RNAscope procedures were performed following the manufacturer’s protocol for the chromogenic, manual, 2.5HD duplex assay for dual staining, and the 2.5HD detection kit (red) for single staining. In short, 4-µm slices of FFPE tissue were mounted, deparaffinized, rehydrated, pretreated with RNAscope 1× target retrieval reagent, then RNAscope protease plus hybridized with the VP60 RHDV2 probes, and detected with the channel 2 alkaline phosphatase enzymatic step with red chromogen. For dual staining, Oc-IL6 (rabbit-specific interleukin-6 mRNA) hybridization was performed and detected with channel 1 using horseradish peroxidase and green chromogen.
In situ detection via RNAscope was accomplished in all RHDV2-positive liver sections. The RHDV1- and 2-negative control liver samples had no detectable viral RNA and lacked any background staining (Fig. 2). There was indistinct background staining in the RHDV1-positive liver sample (Fig. 3), likely as a result of sequence overlap for the VP60 mRNA, but no punctate bright-red staining that was characteristic of all RHDV2-positive liver samples.
There was strong cytoplasmic staining of viral RNA in RHDV2-positive liver samples, particularly within hepatocytes and Kupffer cells (Fig. 4). Biliary epithelium, vascular smooth muscle, and stromal fibroblasts had no consistently detectable viral RNA. In less-affected lobules, viral RNA remained in periportal hepatocytes, with little centrilobular staining; within affected hepatocytes, staining was evenly distributed throughout the cytoplasm, and nuclei were largely unstained. In more heavily affected or autolyzed samples, staining was consistent throughout lobules but concentrated just below the plasma membrane within individual hepatocytes.
Spleen was only available for 2 rabbits, and viral RNA was detected in abundance within splenic macrophages of both the white and red pulp, although generally more heavily concentrated within the marginal zone of white pulp follicles (Fig. 5). In all rabbits, viral RNA was detected in alveolar macrophages, and, in pulmonary tissue with microthrombi and individual cell necrosis, it was also detected in venous endothelium and alveolar capillaries (Fig. 6). Viral RNA was detected throughout the myocardium in all rabbits, with bright-red cytoplasmic staining in both cardiomyocytes and vascular endothelial cells (Fig. 7).
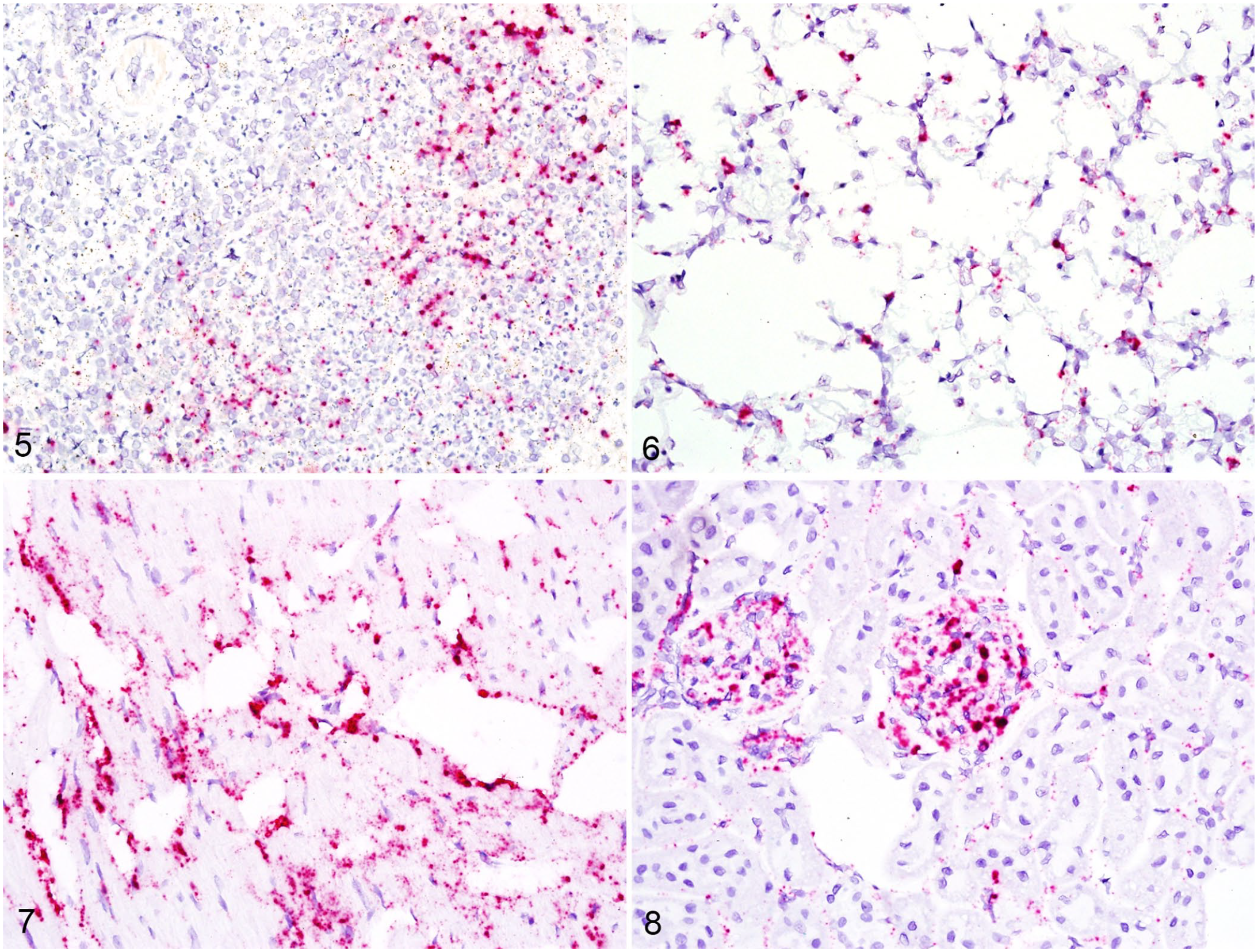
RNA detection in spleen, lung, heart, and kidney by RNAscope in situ hybridization.
Within the kidneys of all rabbits, viral RNA was detected within glomerular tufts and resident macrophages (Fig. 8). In one rabbit with medullary hemorrhage, there was also detectable viral RNA in nearby venous endothelial cells. Two rabbits had adrenal gland included with the tissue set, and the cortex had significant positive staining for viral RNA, particularly in the zona glomerulosa and zona fasciculata. The pancreas and thyroid had no detectable histologic lesions, but viral RNA was plentiful within the stroma in both rabbits for which tissues were included.
Tissues in which viral RNA was detected only within endothelium or tissue macrophages included cerebrum, cerebellum, esophagus, glandular stomach, duodenum, jejunum, ileum, colon, oviduct, uterus, skeletal muscle, and haired skin. Bone marrow was not available for analysis from any of the provided rabbit autopsy tissues. Our results with RNAscope were consistent with positive immunohistochemical staining with pan-lagovirus antibody reported in the liver, spleen, and lungs. 12 Viral RNA was more consistently detected in renal glomeruli compared to antigen detection via IHC, and was additionally detected in the adrenal cortex, myocardium, thyroid, and pancreatic stroma, as well as the endothelium of the meninges, cerebrum, cerebellum, gastrointestinal tract, reproductive tract, skeletal muscle, and skin. The presence of viral RNA in the heart, kidney, gastrointestinal tract, and brain has been reported with PCR detection; however, IHC did not detect viral antigen in tissues with <10 6 viral copies per mg. 12 It is unknown if those tissues with lower viral copy numbers contain actively replicating virus; developing a probe for the negative-sense replicative intermediate may answer that question in the future. Although endothelial cells do have limited phagocytic activity that could explain in situ detection in these areas via RNAscope, the consistent detection of viral RNA within glomerular tufts, adrenal cortex, and large numbers of endothelial cells are more indicative of true infection.
The RNAscope in situ technique proved both sensitive and specific for detection of RHDV2 and was consistently able to detect both the isolate from the Washington State outbreak and the natural recombinant isolate from FADDL, which both differed from the isolate utilized to generate the probe. Although there was minor background staining of the RHDV-positive tissue compared to the control, there was a clear distinction from RHDV2-positive tissues, even those that were reported to have lower viral copy numbers. The background staining noted with RHDV is likely the result of high sequence homology of the VP60 capsid protein to RHDV2, and the short double-Z probes that require only 50–56 bases to form appropriate pairing for amplification. Although this is generally advantageous, given that up to 20 probes can bind a single strand of mRNA each to be amplified up to 1,000 times, for samples with high sequence similarity to the intended target, there may be occasional binding of short exact-match regions. 17 Our attempt to isolate RNA for traditional ISH via riboprobe was unsuccessful because the RNA strands were not long enough to form an effective probe. This failure could be the result of autolysis or prolonged tissue fixation, and further exploration with fresh or more recently fixed tissues may be needed. 10 Successful riboprobe formation for RHDV relied on fresh liver homogenates and cDNA plasmid to accomplish the ideal probe length. 3
In addition to validating the RNAscope method for detection of RHDV2 in situ, we demonstrated viral RNA within the vascular endothelium of multiple organs. Given that endothelial cells have limited phagocytic activity, viral replication may not necessarily occur within these cells, and further study is needed to detect the replicative intermediate negative-sense RNA. Given the role of endothelial cells in disseminated intravascular coagulation, it is worth considering that infection of endothelial cells may contribute to the rapidly fatal course of RHDV2. This highly sensitive and specific method of in situ detection may significantly contribute to further pathogenesis studies to better understand the course of RHDV2 infection.
Footnotes
Acknowledgements
We thank Dr. Fawzi Mohamed, FADDL, USDA-APHIS, for his assistance with control, and RHDV- and RHDV2-positive FFPE blocks, without which our study would not have been possible.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was fully funded through the United States Department of Agriculture Animal Plant Health Inspection Service’s National Bio- and Agro-defense Facility Scientist Training Program.
