Abstract
Tetratrichomonas gallinarum and Trichomonas gallinae are pathogenic avian parasites that infect a wide range of bird species. The pathologic potential of T. gallinarum is controversial, whereas T. gallinae causes disease in many avian species. Infections are often asymptomatic in doves and pigeons; thus, columbids are presumed to represent the natural hosts for trichomonads. The detection of T. gallinarum and T. gallinae is based on direct microscopic observation or a conventional PCR assay. Microscopy is not very sensitive, and identification of the trichomonads at the genus or species level is not possible. Conventional PCR assays have been developed primarily for phylogenetic studies, which detect a wide range of Trichomonas spp. but do not allow their differentiation. We developed a duplex real-time PCR (rtPCR) assay for the simultaneous detection and differentiation of T. gallinarum and T. gallinae. We found that the rtPCR assay detected 102 plasmid DNA copies of T. gallinarum and as few as 101 plasmid DNA copies of T. gallinae.
Keywords
Parabasalia are a group of protists comprised of parasites and commensals of vertebrate and invertebrate hosts.1,22,25 The family Trichomonadidae (order Trichomonadida) contains 2 species found commonly in birds: Tetratrichomonas gallinarum (genus Tetratrichomonas) and Trichomonas gallinae (genus Trichomonas). The latter is closely related to Trichomonas vaginalis, the most prevalent sexually transmitted non-viral human pathogen worldwide. 39 T. vaginalis or its ancestor likely originated in birds, more precisely in columbids, which are probably the natural hosts for trichomonads. 16 This assumption is supported by the high prevalence of asymptomatic infections in doves and pigeons. 31 In addition to Columbiformes, T. gallinae has been found in other avian orders, including Falconiformes, 35 Accipitriformes,5,8 Strigiformes, 26 Galliformes,24,30 Struthioniformes, 33 Passeriformes, 4 Psittaciformes, 28 and Anseriformes. 15 Following the first confirmed epizootic occurrence of T. gallinae in wild finches (Chloris chloris, Fringilla coelebs) in the United Kingdom in 2005, 29 the disease has spread rapidly across continental Europe.7,13,21,32,38,45 Since then, a drastic decline in various finch populations has been observed.
Transmission of T. gallinae among pigeons occurs primarily through direct contact, such as during courtship or feeding of pigeon pulli with crop milk. 42 Raptors can become infected through ingestion of prey carrying T. gallinae. 5 Another possible source of infection is drinking water contaminated by infected animals. 18 T. gallinae is found predominantly in the upper digestive tract where it can cause signs ranging from mild inflammation of the mucosa to caseous areas that can block the esophageal lumen. 41 The disease (generally known as trichomoniasis, “frounce” in raptors, and “canker” in doves and pigeons) can spread systemically and affect the liver, lungs, heart, pancreas, air sacs, and pericardium. 42
In contrast, the pathogenic potential of T. gallinarum is still under debate because the studies conducted on this subject are difficult to compare. Several authors investigated naturally infected chickens and turkeys; others performed experimental infections. But even in the latter studies, the conditions varied to such an extent (origin, amount, application of the inoculum, target species [e.g., laying hens, SPF chickens, turkeys]) that conclusive assessments of the results were not possible.2,12,17,27 Although the pathogenic potential of T. gallinarum is unclear, studies have elucidated the effects of infection, which lead to macroscopic lesions in the ceca as well as granular necrotic areas in the liver. T. gallinarum is found in the large intestine of Galliformes mainly, and Anseriformes.15,19,37 Usually other protozoa, such as Histomonas meleagridis and Blastocystis spp., are present as well. Transmission of T. gallinarum can occur via the fecal–oral route; trophozoites are excreted from infected birds and form pseudocysts within a few hours at ambient temperature. After ingestion, trophozoites develop in the cecum.2,12
To date, no systematic studies exist on the occurrence of coinfections by T. gallinarum and T. gallinae of the same individual, but both protozoa have been detected in common eiders (Somateria mollissima), supporting the possibility of coinfections. 15
In routine testing, trichomonads are detected either microscopically (possibly with prior enrichment) or by a PCR assay. Most PCR primers were developed for phylogenetic analyses and thus were designed to recognize a broad range of Trichomonas spp.6,9 –11 Specific PCRs were developed for the detection of T. gallinarum targeting the 18S rRNA sequence and for T. gallinae based on the Fe-hydrogenase gene or the ITS sequences.14,20,34 Here we describe a new, sensitive and specific, duplex real-time PCR (rtPCR) for the simultaneous detection and differentiation of T. gallinarum and T. gallinae.
Materials and methods
References and clinical samples
DNA of T. gallinae (T. gallinae/Racing Pigeon/Austria/8855-C3/06) and T. gallinarum (T. gallinarum/Turkey/Austria/2721-C7/03) and clinical sample 7 (liver of a not-further-specified bird) were kindly provided by Prof. M. Hess. 3 T. vaginalis DNA was gifted by Unilabs, Zürich. Clinical samples 1–6 (liver and cecum) and 40 (crop) were collected during routine autopsies of birds (Department of Poultry and Rabbit Diseases, Zurich, Switzerland). Other samples were kindly provided by S. Borel (8–38; esophagus, formalin-fixed paraffin-embedded; FFPE), by P. Mattmann (39), and by S. Dirren (41–46; saliva swabs; Table 1). Liver and cecum samples were taken in 2014 and 2020 and stored at −20°C. Esophagus samples were taken between 2012 and 2019, FFPE, and stored at room temperature. Saliva swabs were sampled in 2019 and 2020 and were stored at 4°C. To serve as controls for the validation of our new duplex rtPCR, clinical samples were characterized with current standard methods. Samples 1–7 were examined for the presence of T. gallinarum by conventional gel-based PCR (conPCR). 14 Samples 8–38 were examined microscopically and found positive for Trichomonas spp. Samples 39–46 were found suspicious for trichomonosis at autopsy (caseous lesions in the esophagus). Samples 8–46 were further examined with 2 T. gallinae–specific conPCRs as published elsewhere.20,34
Clinical samples analyzed with the new TriTet real-time PCR duplex assay for the detection of Trichomonas spp.
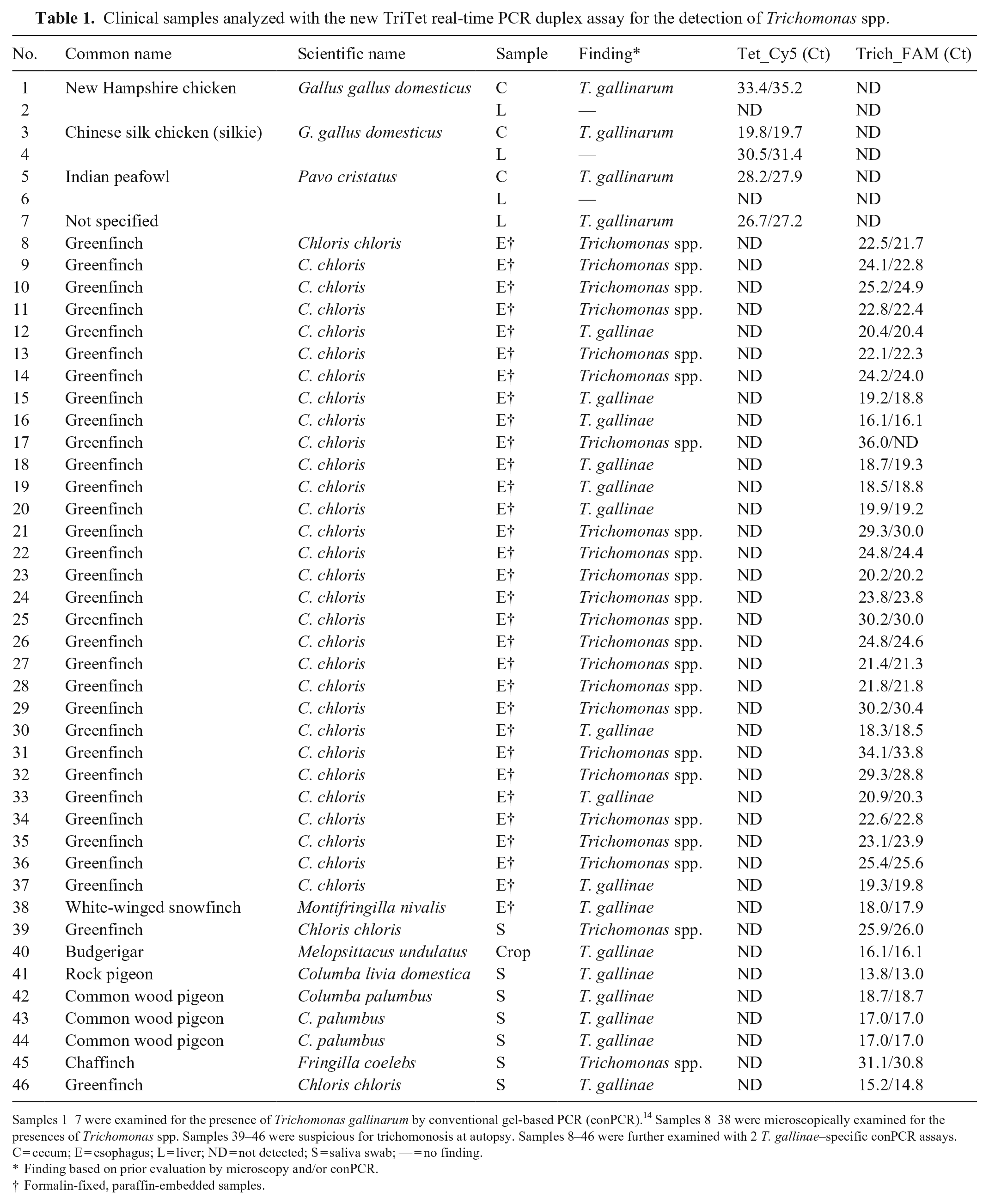
Samples 1–7 were examined for the presence of Trichomonas gallinarum by conventional gel-based PCR (conPCR). 14 Samples 8–38 were microscopically examined for the presences of Trichomonas spp. Samples 39–46 were suspicious for trichomonosis at autopsy. Samples 8–46 were further examined with 2 T. gallinae–specific conPCR assays.
C = cecum; E = esophagus; L = liver; ND = not detected; S = saliva swab; — = no finding.
Finding based on prior evaluation by microscopy and/or conPCR.
Formalin-fixed, paraffin-embedded samples.
DNA extraction
DNA was extracted from organ samples and swabs (NucleoSpin tissue kit; Macherey-Nagel) and from FFPE samples (ReliaPrep FFPE gDNA miniprep system; Promega) according to the manufacturers’ instructions. The extracted DNA was stored at −20°C until further use.
Primer and probe design
For primer and probe design, sequences of the small subunit (SSU) 18S rRNA gene of T. gallinarum isolate GB 1404/11 (JN619423.1), T. gallinae isolate MR30 (KX353945.1), and T. vaginalis isolate 73 (JX943586.1) were aligned using Clustal Omega. 40 This generally well-conserved gene enabled the manual design of 2 forward and 1 reverse primers, allowing the amplification of T. gallinarum, T. gallinae, and T. vaginalis (Fig. 1; Table 2). However, the chosen sequence also contained stretches with variable regions, enabling the differentiation of T. gallinarum and T. gallinae by specific probes (Tet_Cy5 and Trich_FAM, respectively). To allow for simultaneous usage of both probes, different reporter fluorophores were chosen for each of the probes (Fig. 1; Table 2). Primers and probes were synthesized by Microsynth. To confirm the specificity of the probes, BLAST was applied with default settings (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Probe Tet_Cy5 showed 100% nucleotide identity with the SSU rRNA gene of the yellow-necked dry-wood termite (Kalotermes flavicollis), and probe Trich_FAM revealed one potential off-target, which is an uncharacterized protein of Metarhizium album, a fungal pathogen of leaf- and plant-hoppers of rice.

TriTet assay real-time PCR primers and probes. Multiple sequence alignment was performed of the SSU 18S rRNA partial gene sequences of Trichomonas gallinarum isolate GB 1404/11 (JN619423.1), T. gallinae isolate MR30 (KX353945.1), and T. vaginalis isolate 73 (JX943586.1).
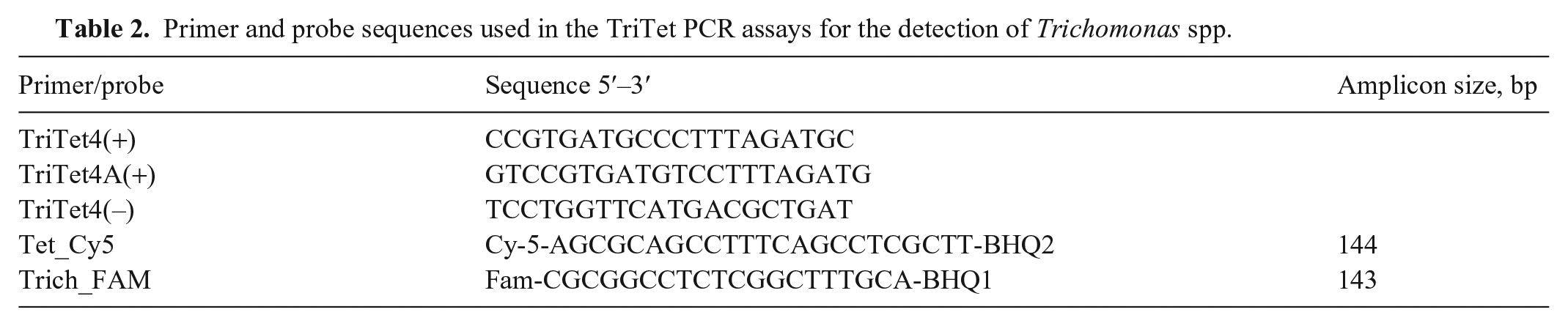
Primer and probe sequences used in the TriTet PCR assays for the detection of Trichomonas spp.

Cloning of recombinant reference plasmids
Using primers TriTet18S(+) (5′-GGATAGCAGCAGCAACTCTG-3′) and HisTriTet18S(–) (5′-CACGTTGAATACGTCCCTG-3′), a 1,302-bp sequence of the SSU 18S rRNA gene of T. gallinarum, T. gallinae, and T. vaginalis was amplified and cloned into the pCR4-TOPO vector (Invitrogen). The resulting plasmids were designated pTet_2721, pTrich_8855, and pTVag, respectively. Using primers Grahn_F1T (5′-GTAGGTGAACCTGCCGTTG-3′) and cf_r1 (5′-TTCAGTTCAGCGGGTCTTC-3′), 379-bp (T. gallinarum), 364-bp (T. gallinae), and 363-bp (T. vaginalis) sequences covering the respective ITS1 regions were amplified and cloned into the pCR4-TOPO vector. The resulting plasmids were designated pTet_ITS, pTrich_ITS, and pTVag_ITS, respectively. Successful cloning was verified by sequencing (Microsynth).
PCR assays
Comparative conPCR
Defined copy numbers of reference plasmids encoding the respective target sequences of T. gallinarum, T. gallinae, and T. vaginalis were used as templates for the conPCR (TriTet conPCR) using the newly designed primers (Table 2) compared to primer sets published previously.11,14 The new primers as well as primers published previously (Tgf/r conPCR), 14 both targeting the SSU 18S rRNA gene, amplify a 144-bp and a 526-bp fragment, respectively. Other primers published previously (ITS conPCR) 11 amplify a 333–379-bp fragment covering the ITS1 region of various Trichomonas spp.
Each reaction contained 12.5 µL of REDTaq ReadyMix (MilliporeSigma), 1 µL of each primer (10 µM), 1 µL of DNA, and ddH2O in a total volume of 25 µL. The amplification conditions were as follows: 1 × 94°C for 15 min, 40 cycles of 94°C for 30 s, 57°C for 30 s, 72°C for 1 min, and a final extension at 72°C for 10 min. PCRs were performed (AllInOneCycler; Bioneer), and amplification products were subjected to capillary electrophoresis (QIAxcel advanced system; Qiagen).
TriTet rtPCR
Reactions were performed using the newly designed primers with either an individual probe or both probes combined (Table 2). The 12.5-µL reaction contained 6.25 µL of Path-ID qPCR master mix (Applied Biosystems), 0.5 µL of each primer (10 µM) and 0.375 µL of each probe (10 µM), 1 µL of reference plasmid DNA or 2.5 µL of DNA extracted from clinical samples in DEPC-treated H2O. The rtPCR assays were performed (QuantStudio 5 rtPCR system; Applied Biosystems) with the following conditions: 1 × 50°C for 2 min, 1 × 95°C for 10 min, 40 cycles of 95°C for 15 s, and 60°C for 1 min.
Determination of assay sensitivity and specificity
The sensitivity and specificity of the assays were determined by performing the rtPCR using either individual probes or as a duplex assay (both probes together) with defined copy numbers of reference plasmid DNA (10-fold serial dilutions corresponding to 100–106 copies per reaction). Three independent experiments were performed in triplicate. Standard curves were generated with the mean Ct values of all experiments and regression analysis was performed using Excel v.16.59 (Microsoft).
Results
Conventional PCR and comparison with published primers
Using defined copy numbers of plasmid DNA encoding the respective target sequence of T. gallinarum, T. gallinae, and T. vaginalis resulted in the detection of all tested Trichomonas samples with as few as 102 copies per reaction (Table 3).
Analytical sensitivity of the conventional gel-based PCR for the detection of Trichomonas spp.
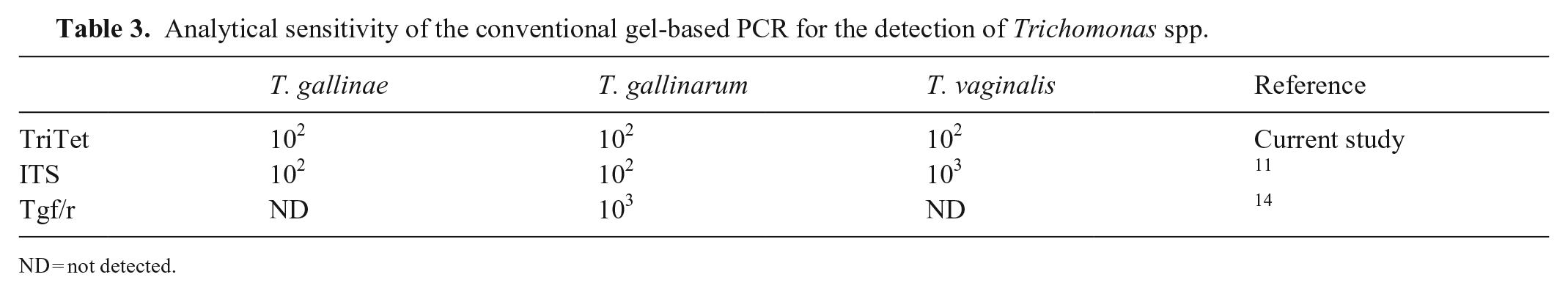

ND = not detected.
The TriTet conPCR described here had the highest analytical sensitivity, with 102 copies being the lowest copy number detected for all 3 tested samples (Table 3). The ITS conPCR was equally sensitive in detecting T. gallinarum and T. gallinae samples, but the lowest detected copy number of the T. vaginalis sample was 103. The Tgf/r conPCR had the lowest analytical sensitivity, with as few as 103 copies detected for the T. gallinae sample. T. gallinarum and T. vaginalis samples were not detected (Table 3).
Analytical sensitivity and specificity of the TriTet duplex rtPCR assay
When using the newly designed probes separately in the TriTet singleplex assay, probe Trich_FAM detected as few as 101 copies of the T. gallinae reference plasmid. Probe Tet-Cy5 recognized as few as 102 copies of the T. gallinarum reference plasmid (Fig. 2; Table 4). The efficiency (E), which is the rate of amplicon generation, was 91.4% and 97.5% when using probes Trich_FAM and Tet_Cy5, respectively. The coefficient of determination (r2), reflecting the linearity of the standard curve, was 0.997 (Trich_FAM) and 0.987 (Tet_Cy5; Table 4).
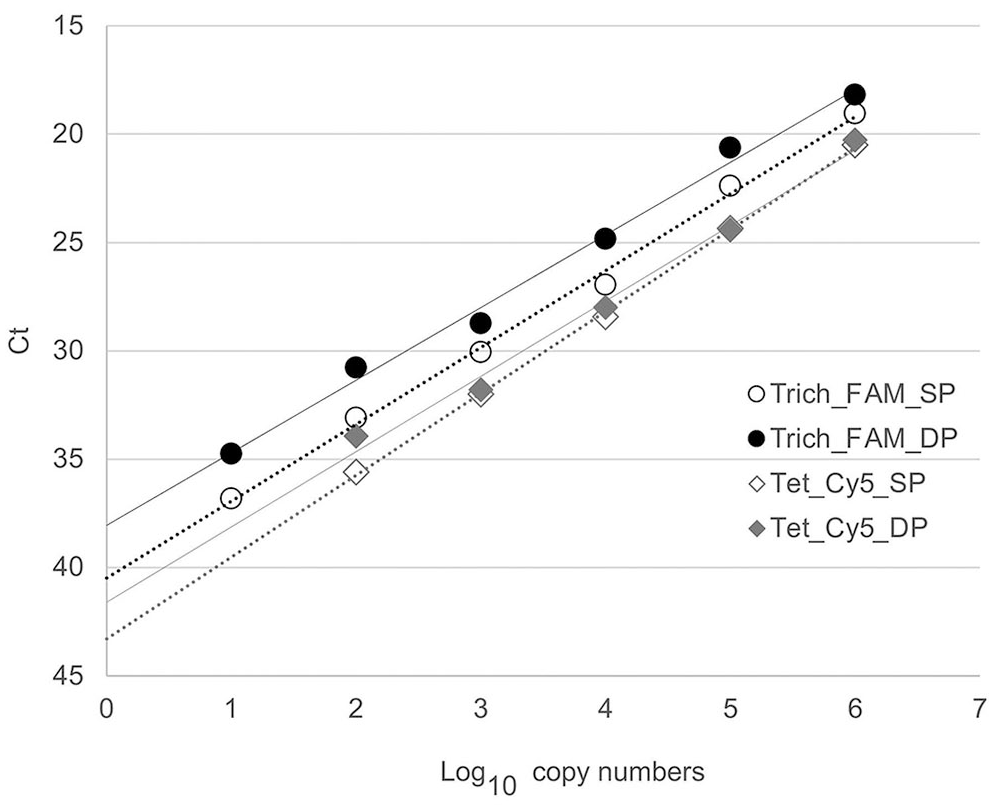
Standard curves of the TriTet real-time PCR assay based on the amplification of 10-fold serial dilutions corresponding to 100–106 copies of the positive control plasmids pTet_2721 and pTrich_8855. The Trichomonas gallinae–specific probe Trich_FAM and the T. gallinarum–specific probe Tet_Cy5 were used either separately in a singleplex (SP) assay or combined in a duplex (DP) assay.
Analytical sensitivity of the TriTet real-time PCR assay for the detection of Trichomonas spp. and regression analyses of standard curves.
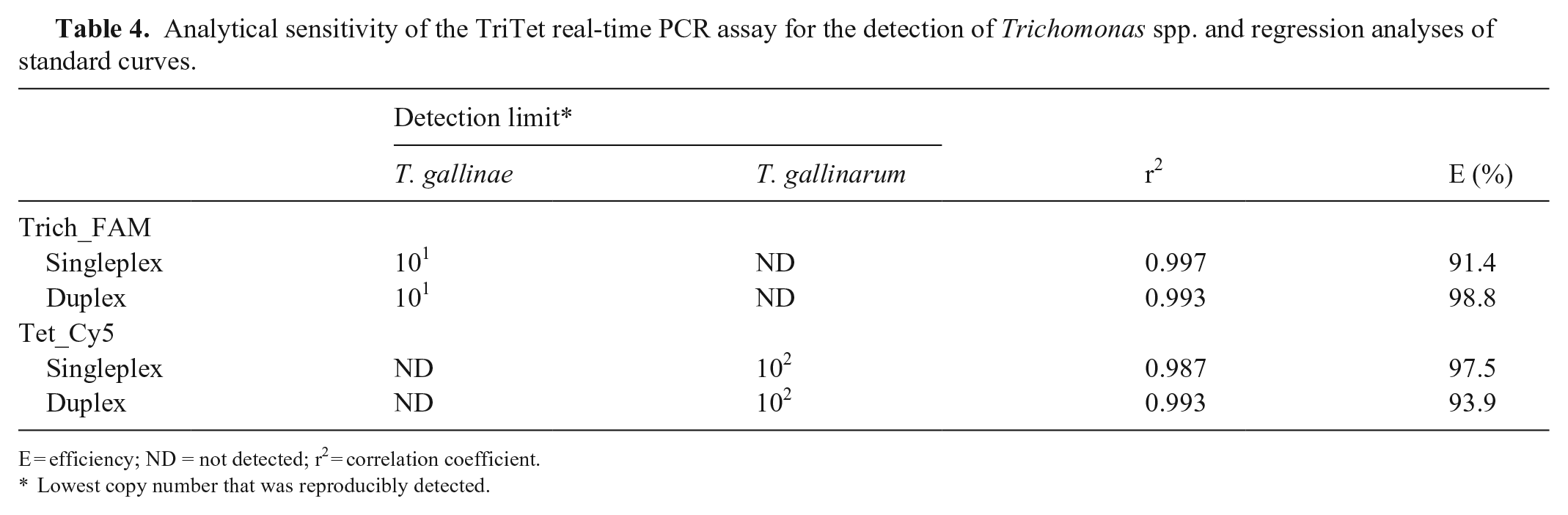
E = efficiency; ND = not detected; r2 = correlation coefficient.
Lowest copy number that was reproducibly detected.
The analytical sensitivity of the TriTet duplex assay for the desired target was unaffected compared to the respective singleplex assay. When detecting T. gallinae and T. gallinarum, E was 98.8% and 93.9%, and r2 was 0.993 and 0.993, respectively. Using probe Trich_FAM, no T. gallinarum was detected, and probe Tet_Cy5 did not recognize the T. gallinae reference plasmid. T. vaginalis was not detected with either probe (data not shown).
Detection of T. gallinarum and T. gallinae DNA in clinical samples
In applying the TriTet rtPCR assay to 46 clinical samples, 5 samples were found positive for T. gallinarum, and 39 samples were positive for T. gallinae. Ct values for the detection of T. gallinarum were 19.7–35.2. Of the 3 cases in which cecum and liver were available for analysis, T. gallinarum was found in the liver in only 1 case (sample 4), in addition to the cecum. It should be noted that the Ct values in the corresponding cecum sample were very low, suggesting a large amount of T. gallinarum DNA. In contrast, the Ct values in the liver sample were very high, which in turn indicates small amounts of T. gallinarum DNA. Ct values for the detection of T. gallinae were 13.0–36.0. No coinfection with both Trichomonas species was detected in any of the tested samples (Table 1).
Discussion
To develop a rtPCR assay that simultaneously detects and differentiates between T. gallinarum and T. gallinae, the first step was to design primers recognizing both species. Results demonstrate that using these primers in a conPCR enables detection of DNA from T. gallinarum and T. gallinae. Furthermore, the primers also allowed the detection of another tested species, T. vaginalis, a close relative of T. gallinae. Further species were not tested given a lack of reference DNA, but in silico sequence analysis suggests that the primers would also allow the detection of additional Trichomonas spp., such as Tetratrichomonas buttreyi, T. brixi, T. tenax, and T. gypaetinii. The latter is also relevant in birds, although T. gypaetinii has only been detected in birds of prey to date.23,44
When comparing our new TriTet assay with published conPCR assays,11,14 the TriTet assay was as sensitive as the ITS assay in detecting T. gallinarum and T. gallinae, 11 both detecting as few as 102 copies. Concerning the detection of T. vaginalis, the TriTet assay was more sensitive than the ITS assay (102 vs. 103 copies). The Tgf/r assay was developed to recognize T. gallinarum exclusively, 9 which could be confirmed here given that neither T. gallinae nor T. vaginalis reference DNA was detected. Nevertheless, the sensitivity was considerably lower than the TriTet assay (103 vs. 102 copies). However, it should be noted that neither the ITS nor the Tgf/r assay has been optimized for the requirements in our laboratory. Under optimal conditions, the Tgf/r assay is supposed to detect as few as 1 organism. 14
In silico sequence analysis of both probes revealed one potential off-target effect for each probe. Tet_Cy5 showed 100% nucleotide identity with the SSU rRNA gene of K. flavicollis, and the nucleotide sequence of Trich_FAM was identical to a sequence stretch of an uncharacterized protein of M. album. The primer sequences were blasted with the potential off-target sequences and revealed a maximum nucleotide identity of 62%. Thus, detection of the SSU rRNA gene of K. flavicollis or the uncharacterized protein of M. album is unlikely.
Using the new primers in combination with the TaqMan probes, the analytical sensitivity with respect to T. gallinarum remained unchanged at 102 copies, whereas the analytical sensitivity for T. gallinae was further increased, and as few as 101 copies could be detected. The sensitivity did not differ between singleplex assays using each probe individually and the duplex assay simultaneously including both probes. Efficiency (singleplex: 91.4%, 98.8%; duplex: 97.5%, 93.9%) and linearity (singleplex: 0.997, 0.993; duplex: 0.987, 0.993) were both in the desired range for rtPCR assays (E: 90–100%; r2: >0.98).36,43
The fact that T. gallinarum was detected in the liver in addition to the cecum in only one case during the analysis of the clinical samples is probably related to the amounts of T. gallinarum in the corresponding organs. This is in accordance with published findings in chicken and turkey, both infected experimentally with T. gallinarum. 2
Our TriTet rtPCR assay presented here seems to be more sensitive than previous conPCR assays, given that all 39 samples considered suspicious for T. gallinae (based on microscopic analysis) were positive. In contrast, only 16 of these 39 samples were positive using the T. gallinae–specific conPCR assay. Notably, these samples resulted in low Ct values when using the TriTet rtPCR assay (Ct < 21), suggesting lower sensitivity of the conPCR assay.
Footnotes
Acknowledgements
We thank Professor Michael Hess (University of Veterinary Medicine Vienna), Prisca Mattmann and Sebastian Dirren (Swiss Ornithological Institute, Sempach, Switzerland), and Stéphanie Borel (Institute for Fish and Wildlife Health, Vetsuisse Faculty, University Bern, Switzerland) for kindly providing Trichomonas spp. DNA and/or clinical samples.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
