Abstract
The H2 subtypes of avian influenza A viruses (avian IAVs) have been circulating in poultry, and they have the potential to infect humans. Therefore, establishing a method to quickly detect this subtype is pivotal. We developed a TaqMan minor groove binder real-time RT-PCR assay that involved probes and primers based on conserved sequences of the matrix and hemagglutinin genes. The detection limit of this assay was as low as one 50% egg infectious dose (EID50)/mL per reaction. This assay is specific, sensitive, and rapid for detecting avian IAV H2 subtypes.
Influenza A viruses (IAVs;
The availability of appropriate testing methods is pivotal for the prompt detection of viruses. In the past, the detection of influenza viruses was based on the clinical signs of patients, serologic tests, and isolation of viruses; however, these methods have their own limitations, such as long turnaround times for results. 17 The reverse-transcription real-time PCR (RT-rtPCR) assay is simple to perform, and exhibits high specificity, sensitivity, and reproducibility. 3 This method has been used widely to detect various viruses, such as influenza, Zika, and dengue viruses.7,14 We aimed to develop a TaqMan minor groove binder (MGB) RT-rtPCR assay for the rapid detection of avian IAV H2 subtypes.
We used 33 reference virus isolates from the virus repositories at our laboratory (State Key Laboratory for Diagnosis and Treatment of Infectious Diseases, Hangzhou, China; Table 1).18,19,21 IAVs had been stored and propagated using 10-d-old chicken embryos, as described previously.
19
Other viruses, including infectious bursal disease virus, Newcastle disease virus (
Avian influenza A viruses and some other viruses used in our study.
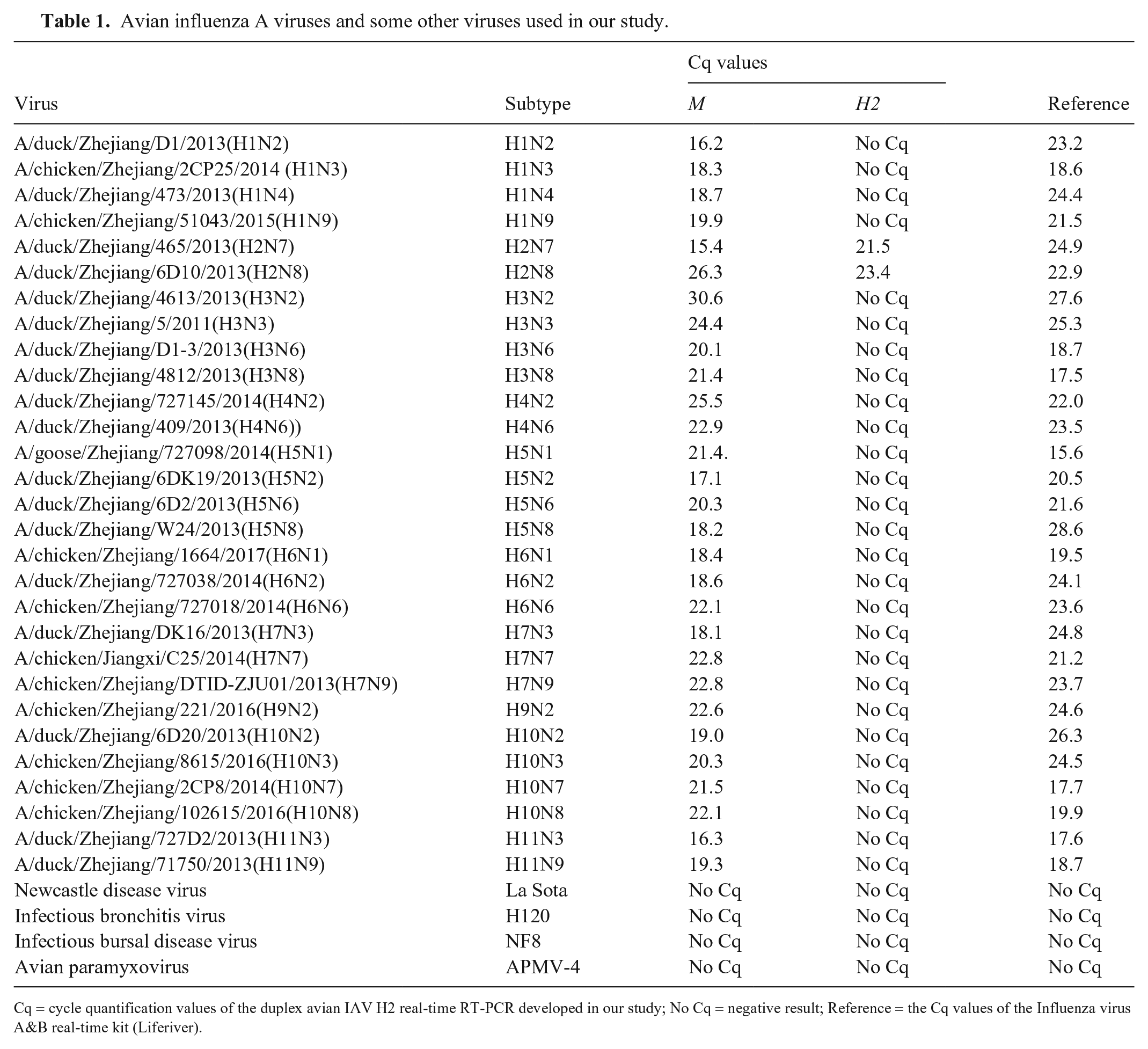
Cq = cycle quantification values of the duplex avian IAV H2 real-time RT-PCR developed in our study; No Cq = negative result; Reference = the Cq values of the Influenza virus A&B real-time kit (Liferiver).
The matrix (
Sequences of primers and probes used for RT-rtPCR assays.
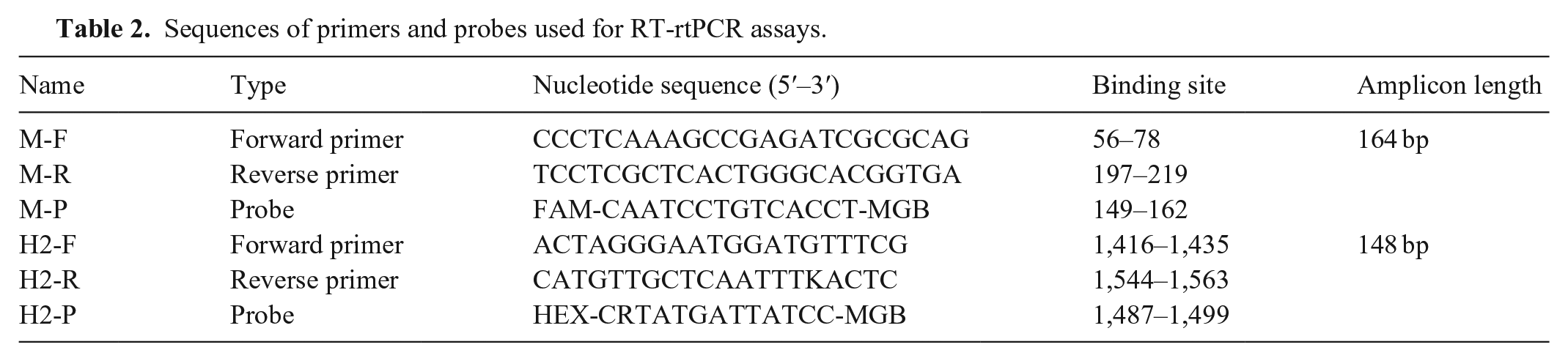
Viral RNA was extracted from respiratory specimens, cloacal swabs, allantoic fluid, and cell culture supernatants (MagPure viral nucleic acid LQ kit; Magen), according to the manufacturer’s instructions. Purified RNA was eluted in 20 µL of nuclease-free water and was either used immediately or stored at −80°C until required. The RT-rtPCR assay was optimized by varying a single parameter while other parameters were maintained constant. The variables evaluated included target primer and probe concentrations, reverse transcription time, and annealing and extension temperature and time. The RT-rtPCR assay was performed in a 20-μL reaction mixture, which was comprised of 10 μL of 2× one-step PCR mix (One-step qRT-PCR kit; Vazyme), 1 μL of enzyme mix, 1.25 μL each of
To test the sensitivity of this method, 10-fold serial dilutions of H2N8 subtype virus RNA, ranging from 1 × 106 to 1 × 100 50% egg infectious dose (EID50)/mL per reaction, were used as the templates for RT-rtPCR. Each dilution was tested 10 times for estimation of sensitivity. Standard curves were constructed by plotting the obtained cycle quantification (Cq) values against the EID50. The standard curves showed a linear correlation from 1 × 106 to 1 × 100 EID50/mL of target virus in multiplex detection. The linear equations of the standard curves of
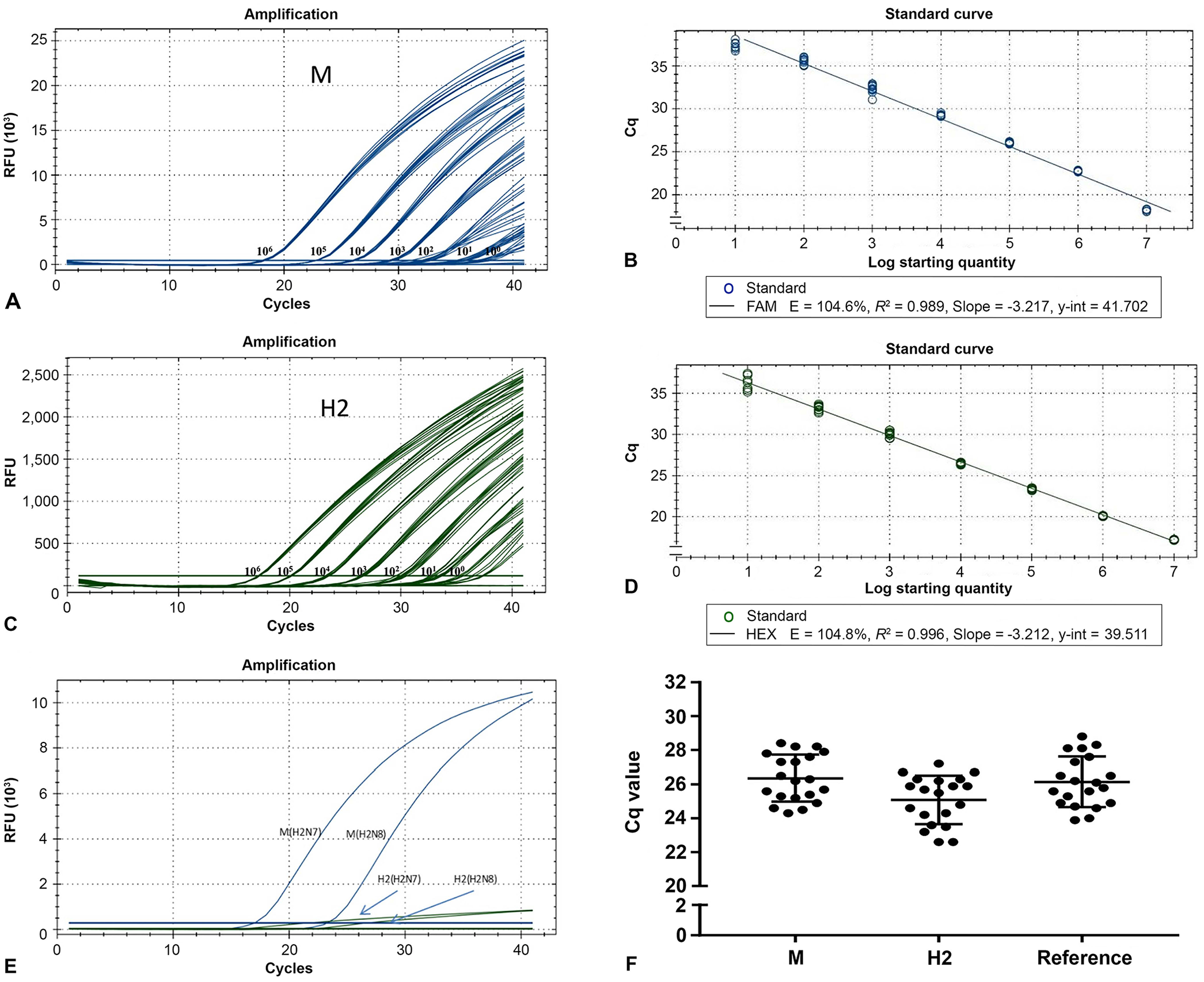
Amplification plots and standard curves of
To compare differences between the intra- and inter-run, experiments were performed in 10 repetitions on a single plate, and the same experiments were repeated at 10 different times. We found that the RT-rtPCR assay had good reproducibility given that the coefficients of variation were <3% (Suppl. Tables 1, 2).
To further evaluate the clinical application of the RT-rtPCR assay, twenty 6-wk-old specific pathogen–free (SPF) chickens were inoculated with avian IAV H2N8 strain A/duck/Zhejiang/6D10/2013 (106 EID50/mL) intranasally (0.1 mL/chicken); 10 healthy 6-wk-old SPF chickens served as controls. All animal experiments were approved by the Institutional Animal Care and Use Committee (IACUC) of Zhejiang University (2017-015). Oropharyngeal swabs were collected at 48 h post-infection. A RT-PCR kit (Influenza virus A&B real time RT-PCR kit; Liferiver) was used as the reference standard. All samples had positive amplification of
IAV H2 subtypes reportedly infect a wide range of species, including poultry, pigs, and humans. The continuous circulation of zoonotic IAVs in poultry and humans can lead to the emergence of new pandemic H2 subtype strains.
2
Virus isolation is the gold standard for the detection of IAVs. Subtypes of viruses can be determined by hemagglutination inhibition assays.
16
Sequencing has also become a way to identify influenza subtypes; it plays an important role in monitoring the reassortment of IAVs, antiviral resistance, and mutations related to antigenic changes.
9
However, such methods are costly, time-consuming, and require strictly controlled conditions. RT-PCR has a wide range of applications in virus detection; RT-rtPCR, in particular, has become the most popular molecular approach used in clinical laboratories.7,14 For IAV detection, many RT-rtPCR methods have been established for IAVs of different subtypes (H1–H16). These methods proved to be highly specific and revealed suitable sensitivity, allowing direct HA subtyping of clinical material.
1
In 2017, a PCR method was reported with a detection limit of 103.2 EID50/mL for serial dilutions of the IAV H2N2 subtype.
6
We designed
Supplemental Material
sj-pdf-1-vdi-10.1177_1040638721994810 – Supplemental material for Development and application of a real-time RT-PCR assay to rapidly detect H2 subtype avian influenza A viruses
Supplemental material, sj-pdf-1-vdi-10.1177_1040638721994810 for Development and application of a real-time RT-PCR assay to rapidly detect H2 subtype avian influenza A viruses by Yixin Xiao, Fan Yang, Fumin Liu, Hangping Yao, Nanping Wu and Haibo Wu in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was funded by Grants from the National Science and Technology Major Project for the Control and Prevention of Major Infectious Diseases in China (2018ZX10711001, 2018ZX10102001, 2020ZX10101016-004-002), Zhejiang Provincial Natural Science Foundation of China (LY19H260006), and the Independent Task of State Key Laboratory for Diagnosis and Treatment of Infectious Diseases (2020ZZ17).
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
