Abstract
Bovine coronavirus (BCoV) is a known cause of enteric disease in cattle; however, its role in bovine respiratory disease (BRD) is poorly understood, with a dearth of evidence of the detection of the virus in respiratory tract lesions. We coupled histologic evaluation of tracheal and lower airway tissues from 104 calves with BRD in which BCoV was detected in the lungs via PCR followed by direct detection of BCoV by immunohistochemistry and an RNA in situ hybridization assay (ISH; RNAscope technology). RNAscope ISH detected BCoV in respiratory epithelium in more cases than did IHC. Using both methods of direct detection, tracheal epithelial attenuation and identification of the virus within lesions were observed commonly. Our results confirm a role of BCoV in respiratory tract infection and pathology, and show that the virus likely plays a role in the development of BRD.
Bovine coronavirus (BCoV; Coronaviridae, Betacoronavirus, Betacoronavirus 1) is closely related to porcine hemagglutinating encephalomyelitis virus (PHEV), canine respiratory coronavirus (CRCoV), equine coronavirus, and the human common cold coronavirus (Human coronavirus OC43, HCoV-OC43).17,18 BCoV is a well-known cause of enterocolitis in young calves and winter dysentery in adult cattle.2,14,19 BCoV has also been implicated as a cause of bovine respiratory disease (BRD) through detection of the virus in nasal swabs from animals with clinical respiratory disease.3,6,15,16 However, its role in BRD remains unsettled, largely because of the inability to consistently reproduce clinical respiratory disease in inoculated calves.9,11,13 There is a dearth of physical evidence utilizing direct detection methods to identify BCoV infection within microscopic lesions in the bovine respiratory tract. 4
Two BCoV challenge studies have utilized immunohistochemistry (IHC) to identify the virus within respiratory epithelium. In one study, degeneration of epithelium was shown in the nasal turbinates post-challenge, but IHC staining was displayed only in non-attenuated epithelium of the nasal turbinates. 9 In another study, BCoV IHC intracytoplasmic immunoreactivity was noted in pseudostratified columnar epithelium with mixed inflammation in the subjacent lamina propria. 5 Although both studies showed BCoV infection of tracheal epithelium, neither displayed direct detection of the virus within a microscopic lesion, which would be consistent with epitheliotropic viral injury.
At the Iowa State University Veterinary Diagnostic Laboratory (ISU-VDL; Ames, IA, USA) from November 2012 to August 2020, BCoV was detected via PCR in >700 lung samples from field submissions involving BRD. Our aim was to characterize the involvement of BCoV in clinical BRD through histologic evaluation of PCR-positive lung and trachea. Additionally, IHC and RNAscope (RNA in situ hybridization; ISH) technology were used to identify the virus in microscopic lesions within the respiratory tract.
Materials and methods
Case selection
The ISU-VDL laboratory database was searched using the following criteria for case selection: BCoV PCR-positive lung tissue and the availability of formalin-fixed, paraffin-embedded (FFPE) respiratory tissue. The availability of FFPE tissue limited the search to submissions from November 2012 to August 2020. Over this time period, 6,515 BRD cases were submitted to the ISU-VDL in which the bovine viral PCR panel was run; 1,265 of these cases were positive for BCoV via PCR. However, not all of these cases included fixed tissue because many of these submissions were nasal swabs. Of ~700 cases with histopathology, 95 cases (100 animals) with the lowest BCoV PCR Ct levels (Ct < 26) were selected; 5 of the selected cases included 2 animals in the submission. H&E-stained slides from all cases were assessed for quality of tissue. Eight animals were excluded because of significantly autolyzed lung tissue or a lack of sufficient lung parenchyma for evaluation. Overall, we included in our study lung tissue from 92 animals from 87 case submissions. In addition, we included 12 BCoV-positive animals from 9 case submissions as a result of the availability of tracheal tissue; these cases had higher BCoV PCR Ct values (26–30).
Histologic evaluation and direct detection assessment
Three diagnostic pathologists (M.C. Rahe, C.L. Siepker, D.R. Magstadt) evaluated all H&E-stained lung sections from selected cases for the presence of microscopic lesions, specifically respiratory epithelial attenuation and bronchiolitis obliterans within lung sections, as well as epithelial attenuation and inflammation within sections of trachea. Additionally, the pathologists evaluated BCoV IHC and RNAscope ISH slides and identified them as positive or negative. The pathologists were anonymized to the assessment or results of the other researchers, and a two-thirds majority was used to settle any discordance in results.
BCoV PCR
Lung was processed into a 10% homogenate using 3 g of tissue with 27 mL of minimal essential medium (MEM; Thermo Fisher) in a grinder (2010 Geno/Grinder; Spex SamplePrep). Samples were extracted using an automated magnetic particle processor system (KingFisher; Thermo Fisher), with a software program available through the manufacturer, and a commercial extraction kit (MagMAX pathogen RNA/DNA kit; Thermo Fisher) according to the manufacturer’s high-volume protocol. Specifically, the high-volume protocol contained: 100 µL of sample, 120 µL of lysis/binding solution concentrate, 120 µL of 100% isopropanol, 2 µL of carrier RNA (1 µg/µL), and 2 µL of Xeno RNA (10,000 copies/µL). Each high-volume wash step was conducted twice using 300 µL of wash buffer 1 and 450 µL of wash buffer 2. Extracted nucleic acids were eluted into 90 µL of elution buffer.
The BCoV reverse-transcription real-time PCR was conducted as a multiplex panel with additional primers and probes targeting bovine respiratory syncytial virus (BRSV; Bovine orthopneumovirus), bovine viral diarrhea virus (BVDV), and bovine herpesvirus 1 (BoHV1; Bovine alphaherpesvirus 1) in a 25-µL reaction using 5 µL of extracted nucleic acid and 20 µL of master mix. The master mix consisted of 5 µL of 5× QuantiTect virus master reverse-transcription mix (Qiagen), 0.25 µL of QuantiTect virus mix (Qiagen), 12.5 µL of nuclease-free water, 0.125 µL (500 nM) of each forward and reverse primer, 0.025 µL (100 nM) of each probe, and 1 µL of Xeno primer/probe mix (Thermo Fisher). Each plate included one each of positive extraction control, positive amplification control, negative extraction control, and negative amplification control.
The assay was conducted on the Rotor-Gene Q (RGQ) 5-plex HRM system (Qiagen) using standard RGQ series software. The RGQ used the following cycling conditions: 1 cycle at 50°C for 20 min; 1 cycle at 95°C for 5 min; and 40 cycles of 95°C for 15 s and 60°C for 1 min. Run data were analyzed with a 0.02 threshold with outliers removed between 0% and 10%. Primer and probe sequences are available upon request.
BCoV IHC
FFPE tissues were sectioned at 4 µm, placed on slides (Superfrost Plus; VWR), and dried in a 60°C oven for 20 min prior to staining. Two antibodies and immunostaining platforms were used, given the discontinuation of one of the antibodies during the course of the study: 1) Biosystems Bond RX (Leica), and 2) Ventana Medical Systems Discovery Ultra (Roche). Ready-to-use detection and ancillary reagents from Leica and Roche were used on the corresponding instrument unless otherwise indicated below.
In addition, BCoV antibodies and dilutions were used on each automated instrument, as indicated below:
1) Biosystems Bond RX
BC-N+ (Ohio Agricultural Research and Development Center [OARDC], Ohio State University, Wooster, OH, USA) cocktail with equal parts of ascites IDs BC26C8.2C, BC22H5.3C, BC22F8.3C of BC-N+ at a 1:400 dilution BC-N+ (OARDC) with ascites ID BC-N+ at a 1:200 dilution
2) Ventana Discovery Ultra
BC6-4-A A (RTI) at a dilution of 1:5,000
The BCoV protocol on the Biosystems Bond RX consisted of 1) a deparaffinization step with Bond dewax solution; 2) antigen retrieval with Bond epitope retrieval solution 1 for 20 min at 100°C; 3) BCoV antibody incubation for 15 min at room temperature (RT); 4) PowerVision Poly-HRP anti-mouse incubation for 25 min at RT; 5) a 3% H2O2 (Fisher) application for 7 min; 6) chromogenic detection using the Bond DAB refine kit with an incubation of 10 min; and 7) chromogenic counterstain with hematoxylin from the DAB refine kit for 5 min. Dehydration and coverslipping were accomplished by using an automated protocol through a series of alcohols and xylenes on the slide stainer and coverslipper (Tissue-Tek Prisma, Tissue-Tek Glas g2; Sakura).
The BCoV protocol on the Ventana Discovery Ultra consisted of 1) a deparaffinization step with EZ Prep solution; 2) antigen retrieval with cell conditioning 1 for 16 min at 100°C; 3) suppression of endogenous peroxidase with Inhibitor ChromoMap for 8 min; 4) BCoV antibody incubation for 28 min at 37°C; 5) Discovery OmniMap anti-MS HRP application and incubation for 16 min at RT; 6) detection with the Discovery ChromoMap DAB kit (RUO) for 8 min; and 7) chromogenic counterstain and bluing with hematoxylin for 8 min and bluing for 4 min. Slides were removed from the instrument, agitated in soapy water, and rinsed in tap water before following the same dehydration and cover slipping protocol described previously.
BCoV RNAscope ISH
The RNA ISH assay was performed utilizing RNAscope technology, probes, kits, and procedures (Advanced Cell Diagnostics, ACD). RNAscope 2.5 documents 322452 and 322360-USM were followed, unless otherwise noted.
FFPE tissues were sectioned at 4 µm and placed on slides (Superfrost Plus; VWR). After drying overnight, slides were heated at 60°C for 1 h before deparaffinizing with 2 changes each of xylene and 100% alcohol. RNAscope H2O2 was applied to each dry tissue section for 10 min at RT. Slides were then immersed in the pre-heated RNAscope target retrieval solution for 15 min at 100°C, with subsequent washes in distilled water and 100% alcohol. RNAscope protease plus was then dropped onto the tissue sections after a hydrophobic barrier was created. Slides were placed in the oven (HybEZ; ACD) for 30 min at 40°C, followed by a distilled water rinse.
Upon completion of deparaffinization, pretreatment, and retrieval, the following ACD probes were applied and hybridized for 2 h at 40°C: 1) target probe V-BCoV-Mebus-S; 2) negative control probe DapB; and 3) positive control probe Bt-PPIB. Six amplification probes from the RNAscope 2.5 HD reagent kit (red assay) were applied in series and incubated as instructed, with wash steps between applications. Amplification was followed with chromogen detection by applying the red mixture to the slides for 10 min at RT. Slides were counterstained with 50% hematoxylin solution for 2 min, dipped in a 0.02% ammonia solution for 10 s, and washed in distilled water. Slides were then dried at 60°C for 15 min, cooled and submerged in a 100% xylene solution, and coverslipped immediately (EcoMount; Biocare Medical). Slides were examined microscopically (BX41 bright-field microscope; Olympus).
Results
Signalment
Of 104 BCoV PCR-positive animals examined, 74 were submitted from Iowa, 7 from Minnesota, 4 from South Dakota, 3 each from Illinois, Wisconsin, and Missouri, 2 each from Florida, Indiana, Texas, and Ohio, and 1 each from Arkansas and Nebraska. The reported age for 82 calves ranged from 5-d-old to 1-y-old, with an average of 18-wk-old and a median of 16-wk-old. The breed was indicated for 81 animals and included 33 crossbreds, 23 Holsteins, 14 Angus, 3 Jersey, and 1 each of Limousin, Pinzgauer, Gelbvieh, Charolais, Shorthorn, Devon, Hereford, and Simmental. The sex was identified for 58 calves: 25 steers, 25 bulls, and 8 heifers.
Bovine respiratory pathogen PCRs
Initially, 89 BCoV PCR-positive animals, from 87 case submissions, with Ct values <26, were identified. Two cases included 2 animals with requested individual molecular testing, with Ct values <26. The BCoV PCR assay was within a PCR panel that also included BoHV1, BRSV, and BVDV. Of the 89 BCoV-positive animals, 81 calves had PCR panels run for bacterial respiratory pathogens that included Mycoplasma bovis, Mannheimia haemolytica, Histophilus somni, and Pasteurella multocida. At least one other viral or bacterial pathogen was detected in 79 of the 89 BCoV-positive calves (Table 1). Only 2 animals were BCoV PCR-positive and also negative for the screened bacteria via PCR. Eight other calves were positive for BCoV and negative for other viruses; however, these animals were not screened for bacterial pathogens via PCR.
Pathogens detected in cattle that were bovine coronavirus (BCoV)-positive in lung by PCR, by BCoV Ct group.
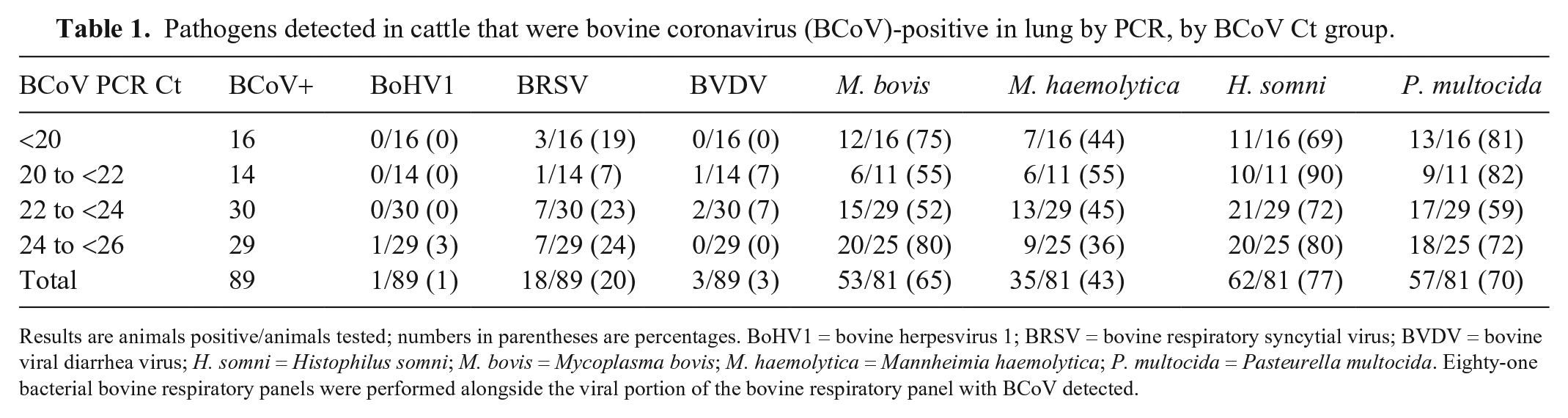
Results are animals positive/animals tested; numbers in parentheses are percentages. BoHV1 = bovine herpesvirus 1; BRSV = bovine respiratory syncytial virus; BVDV = bovine viral diarrhea virus; H. somni = Histophilus somni; M. bovis = Mycoplasma bovis; M. haemolytica = Mannheimia haemolytica; P. multocida = Pasteurella multocida. Eighty-one bacterial bovine respiratory panels were performed alongside the viral portion of the bovine respiratory panel with BCoV detected.
Histologic evaluation of lungs
Epithelial attenuation was observed histologically in the lungs of 52 of 92 animals examined; 25 calves had at least 1 airway with bronchiolitis obliterans, and 16 animals had both attenuation of respiratory epithelium and bronchiolitis obliterans. Within most sections of lung, there were microscopic lesions indicative of common bovine respiratory pathogens such as BRSV, M. haemolytica, H. somni, and M. bovis. All but 5 of the 52 BCoV-positive cases with epithelial attenuation had at least one other viral or bacterial agent detected in the lung via PCR. In 4 of the 5 cases without PCR detection of another agent, a bacterial pathogen was identified via culture.
Direct detection of BCoV in lung
Only 2 calves (BCoV Ct values of 18.1 and 18.2 in the lung) of 92 examined had positive intracytoplasmic BCoV IHC staining of intact respiratory epithelium within airways. The IHC-positive epithelium was pseudostratified or columnar with no evidence of attenuation (Fig. 1). However, 24 cases had intracytoplasmic staining of free cells within cellular debris in the lumens of airways (Fig. 2).
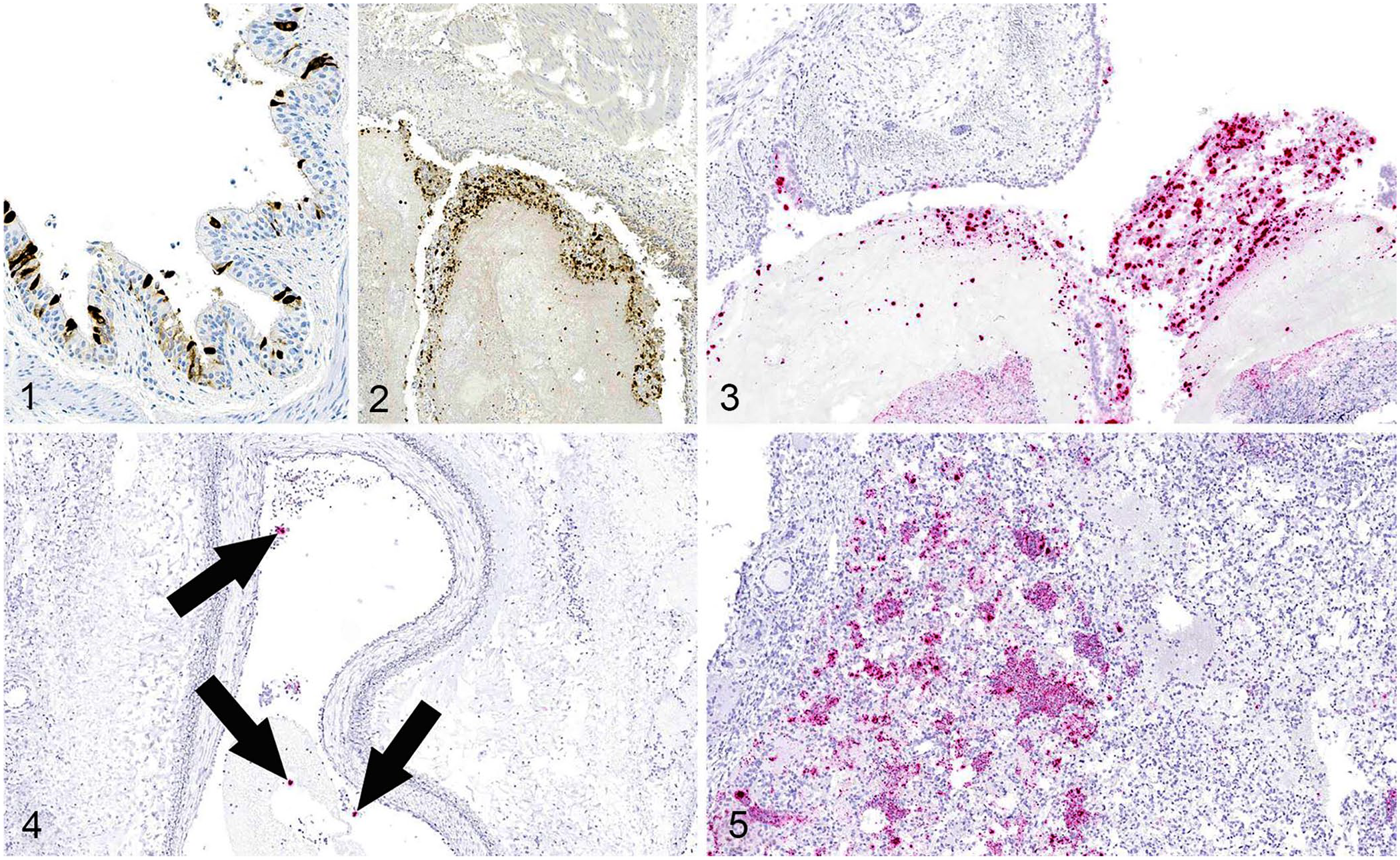
Bovine coronavirus (BCoV) in the lung of cattle with respiratory disease.
For RNAscope ISH, we selected 39 animals with BCoV PCR Ct values <22 and with intraluminal debris IHC staining. BCoV nucleic acid was detected frequently within the cytoplasm of cells free within the lumens of airways: 33 of 39 lungs had positive cytoplasmic staining of debris in bronchioles, and 18 of 19 lungs had positive staining of cellular debris within bronchi (Fig. 3). Lung tissue from 20 animals lacked cross-sections of bronchi for evaluation. Remarkably, the BCoV ISH identified cytoplasmic staining of bronchial epithelium in 17 of 19 lungs, and positive cytoplasmic staining of bronchiolar epithelium in 28 of 39 lungs. Of the 39 lungs probed with BCoV ISH, 28 had evidence of epithelial attenuation, and 20 of these lungs had ISH intracytoplasmic staining within areas of epithelial attenuation.
In several cases, ISH staining was identified in non-respiratory epithelium. For example, 5 cases had punctate staining within alveolar septa; although, interstitial pneumonia was not observed in these animals. Six lungs had intravascular staining, which was often within the cytoplasm of circulating cells (Fig. 4), and 12 lungs had intracytoplasmic staining of cells within alveolar spaces (Fig. 5).
Histologic evaluation of trachea
Direct detection of BCoV infection of cellular debris within airways resulted in the investigation of trachea as a likely site of BCoV infection. Trachea is not a common specimen submitted to the ISU-VDL in BRD cases; however, 17 cases with a total of 21 animals were identified in which BCoV was detected in lung via PCR and fixed trachea was included in the submission.
Evaluation of tracheal mucosa identified epithelial attenuation, ulceration, or regeneration in 20 of 21 calves (Figs. 6, 7). Ulceration and attenuation were observed frequently within the same section of trachea, and regeneration of epithelium was characterized by hyperplastic and frequently stacked epithelium (squamous metaplasia). Within the underlying lamina propria, vessels were often congested with multifocal areas of hemorrhage, and there was frequent infiltration by lymphocytes and plasma cells with scattered neutrophils.
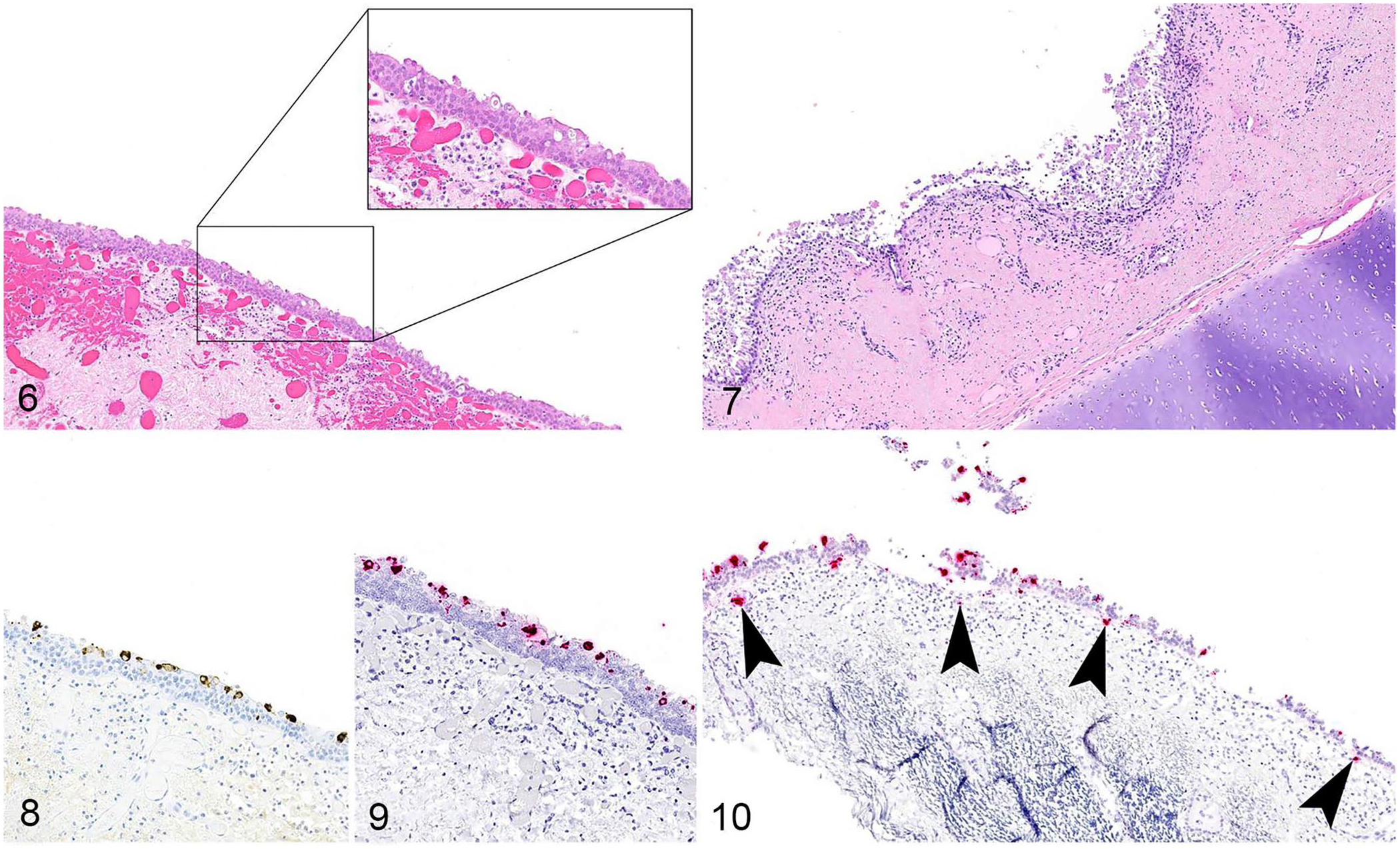
Bovine coronavirus (BCoV) in the trachea of cattle with respiratory disease.
Direct detection of BCoV in trachea
All 21 tracheas were probed with BCoV IHC; intracytoplasmic staining was observed in tracheal epithelium in 15 of 21 animals (Fig. 8; Table 2). RNAscope ISH was performed on 19 of the evaluated tracheal sections (2 animals did not have sufficient tracheal tissue remaining in the FFPE block for examination); BCoV nucleic acid was identified in the tracheal epithelium of 14 of 19 animals (Fig. 9; Table 2). Animals that were ISH- and IHC-negative either had markedly flattened epithelium or had hyperplastic and stacked epithelium, indicative of prior insult to the epithelium. Additionally, both IHC and ISH identified rare cytoplasmic BCoV-positive leukocytes within the subepithelial stroma of the trachea in 10 of 19 calves (Fig. 10).
Direct detection of bovine coronavirus in tracheal tissue with immunohistochemistry (IHC) and RNAscope in situ hybridization (ISH).
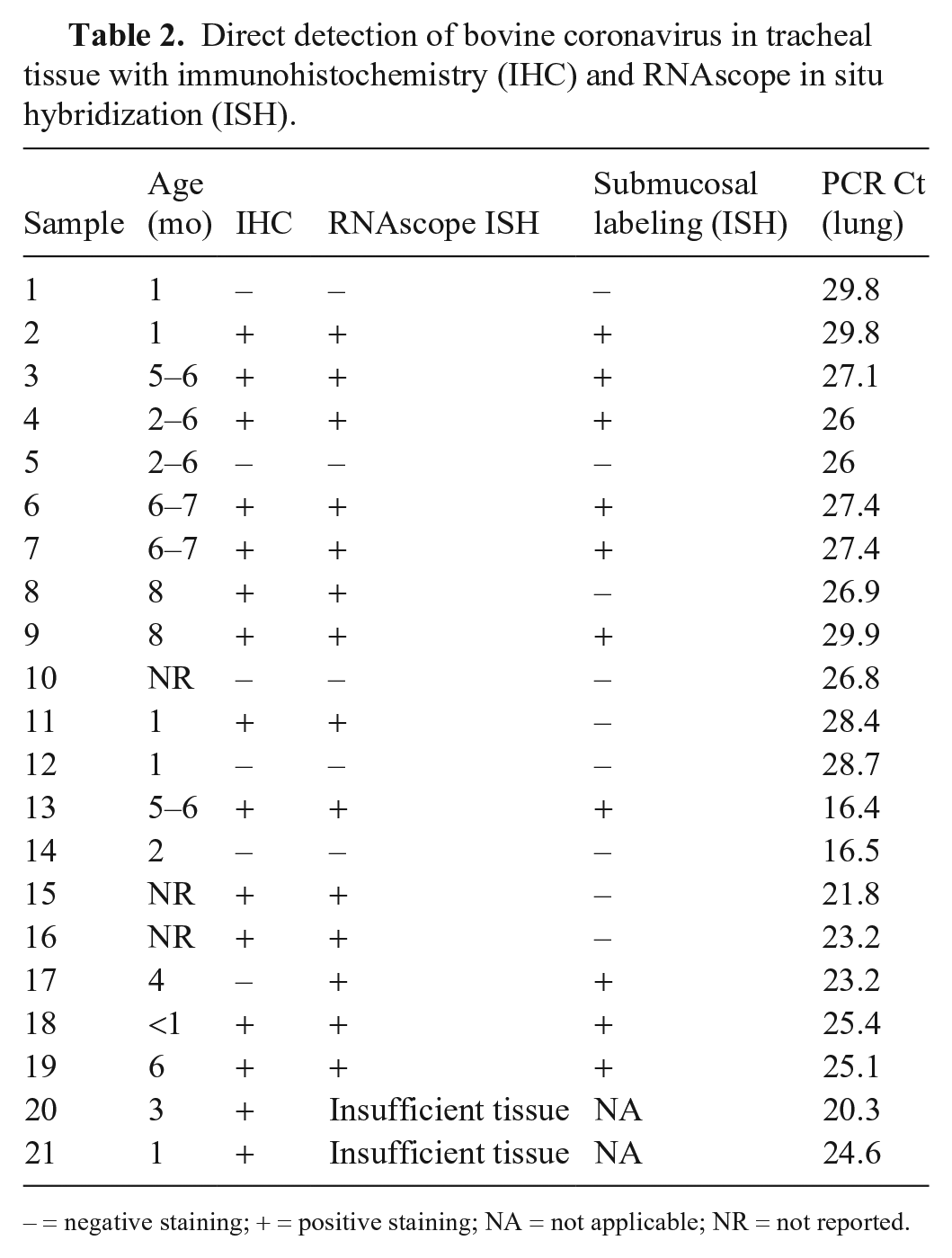
– = negative staining; + = positive staining; NA = not applicable; NR = not reported.
Discussion
We evaluated respiratory tissue from 92 animals in which BCoV was detected by PCR in lung submitted to the ISU-VDL from cattle with BRD. Nearly two-thirds of the evaluated calves, 61 of 92 animals, had lesions consistent with either active or prior epithelial insult. Microscopically, these changes were not diagnostic for BCoV because other bacterial and viral agents were often present. Additionally, no correlating severity of microscopic changes was observed based on PCR Ct. Initially, direct detection of BCoV using IHC provided scant supportive evidence for BCoV as a pathogen of the respiratory system given that only 2 of 92 animals had positive staining of lung respiratory epithelium. However, damage to respiratory epithelium is nonspecific and known to be caused by several pathogens of the bovine respiratory system, many of which were detected in the evaluated lungs.1,7 ISH identified BCoV RNA within the respiratory epithelium of lungs in 28 of 39 calves, and 20 of these calves had evidence of epithelial attenuation. Collectively, these data show that BCoV infects airway epithelium, yet it does not provide sufficient evidence that this infection leads to epithelial damage.
It is important to note that our study was performed on clinical samples from animals that had succumbed to respiratory disease; these calves may have been through a primary viral infection followed by secondary bacterial infection. Viral infection may have occurred and resolved prior to death with only nucleic acid remaining at the time of sample collection. Future challenge studies should use the ability to control the sampling time post-inoculation to evaluate the airways and gastrointestinal tract, with histopathology and direct detection of BCoV to assess the progression of lesions throughout the course of infection.
The identification of IHC-positive intraluminal cellular debris was initially difficult to understand. The specificity of IHC staining was marginal at best; BCoV ISH staining was used to confirm the suspect staining in exfoliated cells. The source of this cellular debris was likely higher in the respiratory tract, given that IHC staining was only rarely observed in cells in alveolar spaces in the lower respiratory tract. Evaluation of available fixed tracheal sections showed that nearly all calves had some degree of epithelial attenuation, along with identification of BCoV in epithelium within lesions. Together, this evidence strongly supports the conclusion of others that BCoV can cause tracheitis in cattle. 5 In the initial analysis of 92 calves with BCoV PCR Ct values <26 in the lung, only 9 calves had trachea submitted with lung. The additional 12 cases in which trachea was evaluated were found by active searching, and although all had Ct values of ≥26 in the lung, 8 of 12 had direct detection of the virus in respiratory epithelium via IHC and ISH. The findings from our study highlight the need for the submission of both tracheal tissue and lung for the diagnostic investigation of cases of respiratory disease.
The ISH identified other areas of the lung in which the virus was present and not detected by IHC (e.g., the identification of BCoV RNA within leukocytes in alveolar spaces and the detection of viral RNA within leukocytes in vessels). This last example would support systemic macrophage trafficking, which has been proposed for BCoV and would be one way in which the virus could access the respiratory or enteric system and cause disease in 2 separate systems in the same animal. 9 Additionally, in a small number of animals (n = 5), BCoV was detected within alveolar septa, which can be observed with hematogenous spread of viruses; however, the evaluated cases lacked interstitial pneumonia as has been described with challenge studies with BCoV.9,12
Although our objective was to characterize the involvement of BCoV in the development of respiratory disease, we limited our examination to trachea and lower airways. Future challenge studies should focus on identifying infection and associated lesions throughout all levels of the respiratory tract, with particular attention paid to the turbinates and trachea. Closely related betacoronaviruses, such as HCoV-OC43, PHEV, and CRCoV, cause respiratory tract infections; our results support previous work suggesting that BCoV has similar tropism for the epithelium of the respiratory tract in cattle.8,10
Footnotes
Acknowledgements
We thank the ISU-VDL histology section for their assistance with the organization of slides from retrospective cases.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This research was funded by a USDA NIFA Animal Health Capacity Program grant GR-022989-00002.
