Abstract
Four outbreaks of bovine respiratory disease (BRD) associated with bovine Coronavirus (BCoV) infection in Italian cattle herds were reported. In 3 outbreaks, BRD was observed only in 2–3-month-old feedlot calves, whereas in the remaining outbreak, lactating cows, heifers, and calves were simultaneously affected. By using reverse transcription polymerase chain reaction (RT-PCR), BCoV RNA was detected in all outbreaks without evidence of concurrent viral pathogens (i.e., bovine respiratory syncytial virus, bovine herpesvirus type 1, bovine viral diarrhea virus, bovine parainfluenza virus). Common bacteria of cattle were recovered only from 2 outbreaks of BRD: Staphylococcus spp. and Proteus mirabilis (outbreak 1) and Mannheimia haemolytica (outbreak 4). A recently established real-time RT-PCR assay showed that viral RNA loads in nasal secretions ranged between 3.10 × 10 2 and 7.50 × 10 7 RNA copies/μl of template. Bovine Coronavirus was isolated from respiratory specimens from all outbreaks except outbreak 1, in which real-time RT-PCR found very low viral titers in nasal swabs.
Introduction
Coronaviruses (CoVs) are large, enveloped, single-stranded RNA viruses that are responsible for enteric, respiratory, or neurological signs in mammals and birds. Belonging to the family Coronaviridae within the order Nidovirales, CoVs are classified into 3 groups on the basis of antigenic and genetic properties. 6 Bovine Coronavirus (BCoV) is included in group 2a together with mouse hepatitis virus, sialodacryoadenitis virus, porcine hemagglutinating encephalomyelitis virus, human Coronavirus (HCoV) OC43, human enteric Coronavirus 4408, 6 the newly recognized HCoV-HKUl, 28 canine respiratory Coronavirus (CRCoV), 5,7 giraffe Coronavirus, 12 and alpaca Coronavirus. 16 Bovine Coronavirus was first identified as the agent of severe diarrhea in neonatal calves (neonatal calf diarrhea), 21 as well as in adult cattle (winter dysentery). 4,20 Subsequently, BCoV was also associated with the occurrence of respiratory distress in calves and adults. 19,22 The respiratory syndrome is often observed during or after transportation because the shipping of cattle represents a stressing factor that may facilitate the onset of BCoV-induced respiratory disease, mainly in feedlot calves. The present study reports the results of molecular investigations on outbreaks of respiratory disease due to BCoV infection in Italian cattle herds.
Materials and methods
Outbreaks and sample collection
Outbreak 1. In January 2006, respiratory disease was observed in an abattoir in the province of Taranto (Apulia), where three 3-month-old calves exhibited severe dyspnea and fever (40.3°C-40.5°C) at routine antemortem examination. The calves were immediately slaughtered to prevent the spread of infection to other cattle. At postmortem examination, severe signs of acute rhinotracheitis were observed, whereas lungs did not show remarkable lesions. Nasal swabs collected from the affected animals (61/06-A, B) before their slaughter were sent to the University of Bari laboratory for virological investigation.
Outbreak 2. In September 2006, respiratory and enteric disease affected a dairy cattle herd in the province of Brindisi (Apulia). Vaccination against bovine viral diarrhea virus (BVDV) had been routinely performed using a modified live virus administered at 6-month intervals. Clinical signs were observed in 2–3-month-old calves as well as in heifers and lactating cows. Despite the severity of the disease in calves and the decrease in milk production in cows, all animals recovered progressively after treatment with Oxytetracycline. Rectal (or fecal samples), ocular, and nasal swabs were collected 48 hours after the onset of the clinical signs from 10 lactating cows (339/06–1 to 10), 10 heifers (339/06–11 to 20), and 10 calves (339/06–21 to 30).
Outbreak 3. During the same month, extensive mortality (about 50%) occurred in a beef cattle herd in the province of Taranto (Apulia), where 42 of 50 feedlot calves showed respiratory distress and fever (39.8°C-0.6°C). Twenty-six calves died even though they were treated with various antimicrobials (i.e., macrolides, sulphamides, quinolones). The herd had been regularly vaccinated against infectious bovine rhinotracheitis. Nasal swabs and fecal samples were taken from 6 ill calves (341/06–1 to 6) for laboratory investigation.
Outbreak 4. Outbreak 4 occurred in December 2006 in a cattle herd in the province of Cosenza (Calabria). Sixty-two 2–3-month-old feedlot calves were severely affected by fever (40.0°C-41.2°C), diarrhea, and respiratory distress, resulting in 18 deaths. Nasal swabs and fecal samples were collected from 8 animals (438/06–1 to 8).
RT-PCR for detection of bovine Coronavirus RNA
Nucleic acids for reverse transcription polymerase chain reaction (RT-PCR) and PCR assays were extracted using commercial kits a,b specific for DNA and RNA purifications. Detection of BCoV RNA was carried out using a 1-step RT-PCR kit c and primers specific for the spike protein gene that are able to detect both BCoV and CRCoV RNAs. 7 The following thermal protocol was used: reverse transcription at 50°C for 30 minutes, inactivation of Superscript II RT c at 94°C for 2 minutes, 40 cycles of 94°C for 30 seconds, 55°C for 30 seconds, 68°C for 30 seconds, and a final extension at 68°C for 10 minutes. The PCR products were detected by electrophoresis in a 1.5% agarose gel and visualized under ultraviolet light after bromide ethidium staining.
Real-time RT-PCR for quantification of bovine Coronavirus RNA
A real-time RT-PCR assay based on TaqMan technology (Gabriella Elia, unpublished data) was used to quantify the viral load in samples that tested positive for BCoV by conventional RT-PCR. Reverse transcription was carried out using a commercial kit, d following the manufacturer's recommendations. The quantitative assay targeting the M gene was conducted in a 50-μl reaction mixture containing 25 μl IQ Supermix, e 600 nM of primers BCoV-F (CCTTCATATCTATACACATCAAGTTGTT) and BCoV-R (ACCAGCCATTTTAAATCCTTCA), 200 nM of probe BCoV-Pb (6FAM-CCTTCATATCTATACA-CATCAAGTTGTT-BHQ1), and 20 μl of cDNA. The thermal profile consisted of activation of iTaq DNA polymerase at 95°C for 10 minutes, followed by 45 cycles of denaturation at 95°C for 15 seconds and annealing extension at 60°C for 1 minute.
Virus isolation
Virus isolation was carried out on samples representative of each outbreak of BCoV respiratory disease. Nasal swabs were homogenized in Dulbecco's minimal essential medium f plus antibiotics (penicillin 5000 IU/ml, streptomycin 2500 μg/ml, amphotericin B 10 μg/ml). g After centrifugation at 1,500 × g for 15 minutes, the supernatants were used to inoculate confluent monolayers of human rectal tumor (HRT-18) in the presence of trypsin (5 μg/ml). h Viral growth was monitored by an immunofluorescent (IF) assay using a bovine serum positive for BCoV antibodies and a rabbit antibovine IgG conjugated with fluorescein isothiocyanate. i Each sample was passaged 3 times prior to being considered as negative for BCoV.
RT-PCR assay for screening for other bovine viral pathogens
RNA and DNA extracts from nasal and ocular swabs were tested by nested PCR for bovine respiratory syncytial virus (BRSV) 26 and bovine parainfluenza virus type 3 (BPIV-3) and by PCR for bovine herpesviruses types 1 (BoHV-1) 27 and 4 (BoHV-4). 2 RNA extracts from rectal swabs were used for RT-PCR detection of rotaviruses 10 and caliciviruses, 15 whereas RT-PCR for bovine viral diarrhea virus (BVDV) 24 was carried out on RNAs purified from nasal and fecal samples. The RT-PCR and PCR assays were performed using specific kits. c,j
Bacteriological investigations
Standardized procedures were carried out on respiratory specimens for in vitro isolation of the most common bacteria responsible for respiratory disease in cattle. Samples were plated out on 5% sheep blood agar and cultured aerobically at 37°C for 24 hours for detection of aerobic pathogens. Bacteria were identified by standard biochemical procedures and analytical profile index. k For isolation of mycoplasmas, samples were inoculated into modified Hayflick broth at 37°C, as previously described. 25
Results
Detection and quantification of bovine Coronavirus RNA in clinical samples
The results of molecular detection of BCoV RNA in clinical samples are summarized in Table 1. Conventional RT-PCR detected the BCoV RNA in both nasal swabs from outbreak 1. With regard to outbreak 2, 30 of 30 rectal swabs or fecal samples, 28 of 30 nasal swabs, and 23 of 30 ocular swabs tested BCoV positive. BCoV was also detected in 2 of 6 (341/06–1, 3) nasal swabs collected from outbreak 3 and in 6 of 8 nasal swabs and 7 of 8 fecal samples collected from outbreak 4.
Viral loads calculated by real-time RT-PCR yielded different results on the basis of outbreak and sample type. In outbreaks in which only respiratory disease was observed, BCoV RNA titers detected in nasal swabs were substantially low in outbreak 1 (4.50 × 103, 7.69 × 103 RNA copies/μl of template) and slightly high in outbreak 3 (9.15 × 105, 3.13 × 106 RNA copies/μl of template). In outbreak 2, in which both the respiratory and enteric forms occurred, viral titers ranged from 2.38 × 102 to 3.00 × 108, 5.10 × 102 to 4.61 × 107, and 3.87 × 101 to 8.04 × 106 RNA copies/μl of template in fecal specimens, nasal, and ocular swabs, respectively. A similar pattern of shedding was observed in cattle in outbreak 4, which analogously displayed respiratory distress and gastroenteritis, with virus shedding comprising between 8.98 × 102 and 7.14 × 107 in nasal swabs and between 8.34 × 101 and 8.54 × 107 RNA copies/μl of template in rectal swabs.
Bovine Coronavirus (BCoV) RNA titers detected in samples collected from Italian cattle herds with bovine respiratory disease (BRD).*
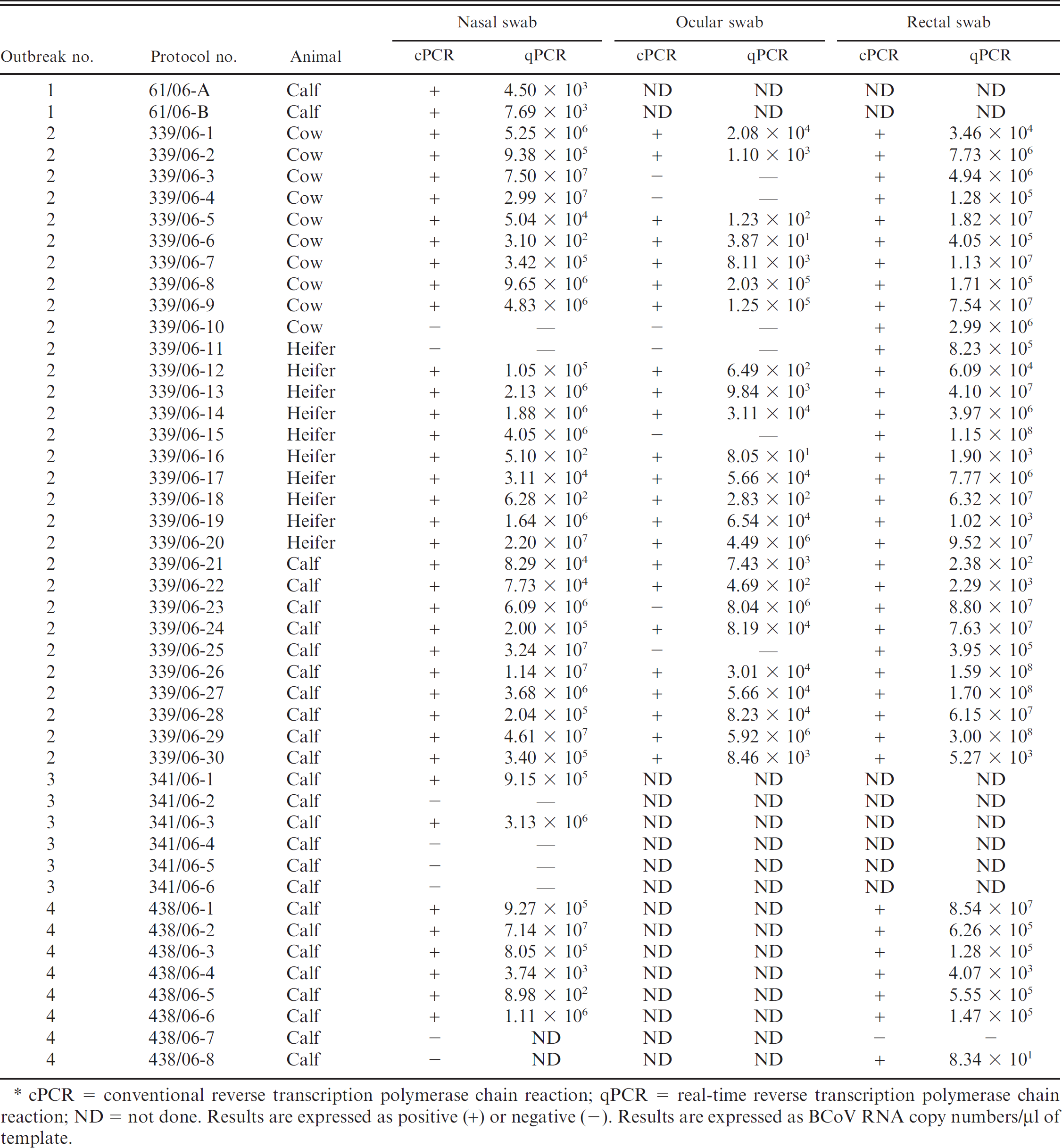
cPCR = conventional reverse transcription polymerase chain reaction; qPCR = real-time reverse transcription polymerase chain reaction; ND = not done. Results are expressed as positive (+) or negative (−). Results are expressed as BCoV RNA copy numbers/μl of template.
Isolation of bovine Coronavirus strains from respiratory outbreaks
BCoV strains were isolated from all outbreaks of respiratory disease, with the exception of outbreak 1, in which the BCoV titers detected in nasal swabs were very low. Successful virus isolation was confirmed by the occurrence of cytopathic effects in the inoculated cells, as well as by the observation of cytoplasmic fluorescence after IF assay using an anti-BCoV serum.
Simultaneous detection of other viral and bacterial pathogens of cattle
BRSV, BPIV-2, BVDV, BoHV-1, and BoHV-4 were not detected in nasal or ocular swabs of cattle with respiratory distress belonging to the different outbreaks, whereas a bovine rotavirus G6P[11] strain was identified in the feces of diarrheic feedlot calves of outbreak 4. Mycoplasmas were not detected in any respiratory specimens, and only common bacteria were detected in nasal swabs, including Staphylococcus spp. and Proteus mirabilis (outbreak 1) and Mannheimia haemolytica (outbreak 4). No bacteria were isolated from respiratory specimens from outbreaks 2 and 3.
Discussion
BCoV is usually associated with the occurrence of diarrhea in calves and adult cattle, but it is also responsible for respiratory distress, being implicated as an etiological agent of bovine respiratory disease (BRD). Bovine respiratory disease is a complex multifactorial disease caused by multiple viruses such as BRSV, BoHV-1, BPIV-3, BVDV, and BVDV. 3,11,18 Concurrent bacterial and mycoplasmal infections can exacerbate the course of viral diseases, increasing morbidity and mortality rates. 9
This article describes 4 outbreaks of
The BCoV respiratory disease has been described in different areas worldwide, including the United States, 14,19,23 Canada, 8 Turkey, 13 Japan, 17 Finland, 1 and Sweden. 11 In the Finland study, 1 various microorganisms were detected in association with BCoV, including mycoplasmas (mainly Mycoplasma bovis and Mycoplasma dispar), Pasteurella multocida, Mannheimia haemolytica, and Arcanobacterium pyogenes, as well as other viral pathogens, specifically, BPIV3, BRSV, and bovine adenoviruses. In the present study, concurrent viral infections were not detected in the respiratory tract, whereas a rotavirus strain was associated with BCoV-induced gastroenteritis in 1 outbreak. Mycoplasmas were not identified in any respiratory samples, and common bacteria were detected in only outbreaks 1 and 4, probably as a result of the antibiotic treatments administered to cattle in outbreaks 2 and 3.
In 2 of the 4 outbreaks described, BCoV was simultaneously responsible for respiratory and enteric signs in most animals, whereas in the remaining 2 outbreaks, only BRD was observed. Extensive epidemiological surveys will assess whether BRD in Italian cattle is associated mainly with BCoV infection, thus requiring specific vaccination programs in feedlot calves in addition to those used in pregnant cows. Moreover, experimental infection of calves followed by immunohistochemistry or in situ hybridization on respiratory tract sections is required to identify virus localization and its correlation with histological lesions.
In 3 of the 4 outbreaks, BRD occurred only in 2–3-month-old feedlot calves as a result of a single BCoV infection without evidence of classical respiratory pathogens, such as BRSV, BoHV-1, and BVDV. The age of the calves with BCoV-induced BRD is likely related to the waning of maternally derived immunity. The occurrence of respiratory disease in 2–3-month-old calves, even if not associated with high mortality rates, may have a negative impact on farm production as a result of reduced weight gain and the cost of veterinary care. However, only a precise analysis of the economic impact of BCoV infection will assess whether there is a need for extensive vaccination of cattle herds against BCoV respiratory disease.
Acknowledgements
This work was supported by grants from the University of Bari, Italy: project ex 60% 2007, “Messa a punto di un sistema real-time RT-PCR per la identificazione e la quantificazione dell'RNA del Coronavirus bovino.”
Footnotes
a.
DNeasy Tissue Kit, QIAGEN S.p.A., Milan, Italy.
b.
QIAamp Viral RNA Mini Kit, QIAGEN S.pA., Milan, Italy.
c.
SuperScript One-Step RT-PCR for Long Templates, Life Technologies, Invitrogen, Milan, Italy.
d.
GeneAmp RNA PCR, Applied Biosystems, Applera Italia. Monza, Italy.
e.
IQ Supermix, Bio-Rad Laboratories srl, Milan, Italy.
f.
Dulbecco's minimal essential medium, BioWhittaker, Verviers, Belgium.
g.
PenStrep Amphotericin B (100×), BioWhittaker, Venders. Belgium.
h.
Trypsin, Biochrom KG, Berlin, Germany.
i.
Antibovine IgG FITC conjugate, Sigma Aldrich srl, Milan, Italy.
j.
LA PCR Kit version 2.1, TaKaRa Bio Inc., Shiga, Japan.
k.
API, BioMérieux Italia S.p.A., Rome, Italy.
