Abstract
Mycoplasma hyopneumoniae is the primary etiologic agent of swine enzootic pneumonia (EP), in which the immune response is reduced, making pigs susceptible to secondary infections. We surveyed commercial pig herds in Brazil for viral and bacterial respiratory coinfections that could complicate EP. Over a 2-y period (2015–2016), we found that 854 of 2,206 pigs (38.7%) were positive for M. hyopneumoniae in herds from various production systems in 3 Brazilian regions (Central-West, Southeast, South). We collected samples of 321 lungs positive for M. hyopneumoniae from 169 farms to also screen for Pasteurella multocida, Actinobacillus pleuropneumoniae, Glaesserella parasuis, influenza A virus (IAV), and porcine circovirus 2 (PCV2) by real-time PCR. The prevalence of pathogens found in addition to M. hyopneumoniae varied: P. multocida (141; 43.9%), G. parasuis (71; 22.1%), PCV2 (50; 15.6%), IAV (23; 7.2%), and A. pleuropneumoniae (18; 5.6%). G. parasuis was more frequent in farrowing or nursery herds (48.7%) than in breeding and fattening herds (10% and 18.6%, respectively; p < 0.01); A. pleuropneumoniae was found only in herds on farrow-to-finish and fattening farms.
Mycoplasma hyopneumoniae causes enzootic pneumonia (EP), the main chronic respiratory disease of swine in commercial herds. M. hyopneumoniae is highly prevalent in important pig-producing regions, and results in economic losses worldwide. After detection in a pig-producing farm, attempts to control the spread of EP include management practices, housing conditions, and antimicrobial medication. 23
Animals with EP typically have a dry cough. M. hyopneumoniae colonizes ciliated respiratory tract cells and predisposes to coinfection with other bacteria, which results in more severe pulmonary lesions. 23 EP can also be potentiated with viral coinfections, such as by influenza A virus (IAV; Orthomyxoviridae, Alphainfluenzavirus) and porcine circovirus 2 (PCV2; Circoviridae, Circovirus).6,23 PCV2 coinfection may further progress to post-weaning multi-systemic wasting syndrome. 11
M. hyopneumoniae and related respiratory infections have been reported on a limited geographic scope in Brazil.2,9,21 Direct molecular detection is described less commonly and is restricted to specific regions.21,22 We aimed to detect M. hyopneumoniae and other bacterial and viral pathogens associated with EP by PCR assays in commercial pig herds in Brazil.
Veterinarians and technicians collected swine lung samples from field autopsies from January 2015 to December 2016. Both lungs were removed from each of 2,206 animals, and tissue samples were selected randomly from cranioventral and caudal lobes, with or without pneumonic lesions, according to procedures described previously. 3 Lung samples were maintained at −20°C until analysis. Cotton-tipped swabs were used to scrape different parts of the lung samples, and nucleic acids were extracted with commercial reagents (NewGene PreAmp/Prep; Simbios Biotecnologia) according to the supplier’s protocol. Samples positive for M. hyopneumoniae (from 321 animals) were used to investigate coinfections with other pathogens. This convenience sampling included animals from 169 herds located in 14 municipalities in 3 important pig-producing Brazilian regions: Central-West, Southeast, and South (Fig. 1). The herds were also classified according to the production systems: 1) 20 breeding or multiplier units; 2) 39 farrowing and nursery units; 3) 253 fattening units; and 4) 9 one-site farrow-to-finish systems.
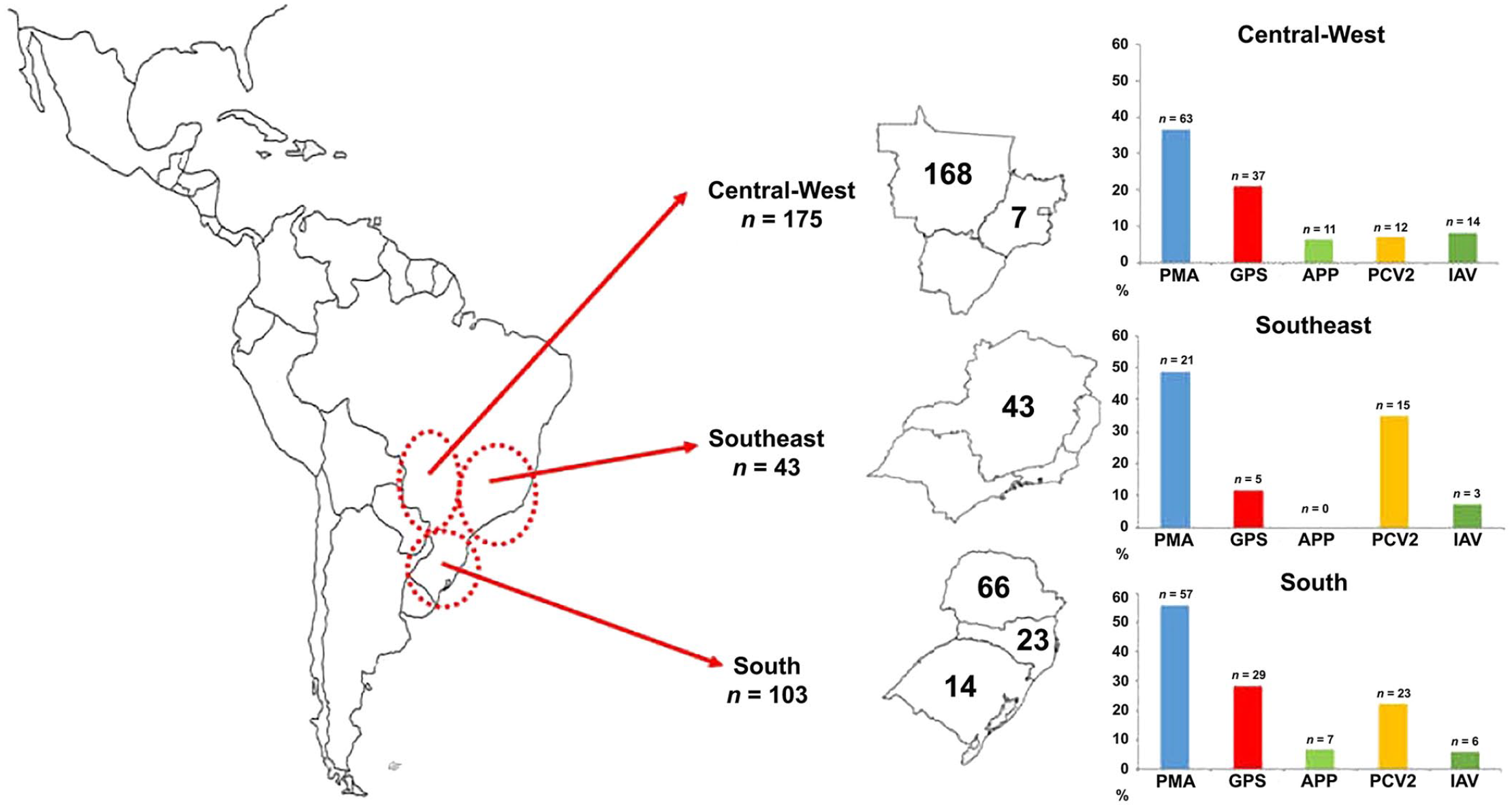
Geographic distribution of Mycoplasma hyopneumoniae coinfections in 3 Brazilian regions (n = the number of pigs sampled in each region and state); these 3 regions hold >80% of the swine herds in Brazil. Graphs show the pathogens present as percentages of coinfections in samples positive for M. hyopneumoniae in each region. APP = Actinobacillus pleuropneumoniae; GPS = Glaesserella parasuis; IAV = influenza A virus; PCV2 = porcine circovirus 2; PMA = Pasteurella multocida.
M. hyopneumoniae was detected by real-time PCR (rtPCR; NewGene MHYAmp reagents; Simbios Biotecnologia). Four other rtPCR assays were carried out to detect Pasteurella multocida, Actinobacillus pleuropneumoniae (APP), Glaesserella parasuis, and PCV2 (NewGene PMAmp, APPAmp, HPSAmp, and PCVAmp, respectively; Simbios Biotecnologia). IAV was detected using one-step reverse-transcription rtPCR (AgPath-ID; Thermo Fisher).
M. hyopneumoniae was detected in 854 of 2,206 (38.7%) lung samples. Among the 3 main Brazilian geographic regions, 106 of 206 (36%) animals from the Central-West region were M. hyopneumoniae infected, as were 460 of 774 (59%) from the Southeast region, and 228 of 1,136 (25%) from the South region.
In the 321 convenience lung samples positive for M. hyopneumoniae, the sole pathogen was M. hyopneumoniae in 106 of 321 (33.0%); 140 of 321 (43.6%) samples contained 1 other agent (virus and/or bacterium), 62 of 321 (19.3%) had 2 other agents, and 13 of 321 (4.0%) had 3 other agents. Most coinfections occurred with 1 additional bacterial species (n = 113; 35.2%), but there were also infections with 2 (n = 29; 9%) and 3 (n = 3; 0.9%) different bacteria. PCV2 and IAV were also detected; only 1 virus in most animals (n = 27, 8.4%), but both viruses were present in 3 samples (0.9%). Additionally, we observed M. hyopneumoniae with 1 bacterium and 1 virus (n = 30; 9.3%), as well as with 2 bacteria and 1 virus (n = 10; 3.1%). The most frequent co-pathogens were: P. multocida (n = 141; 43.9%), G. parasuis (n = 71; 22.1%), PCV2 (n = 50; 15.6%), IAV (n = 23; 7.2%), and APP (n = 18; 5.6%; Fig. 2).
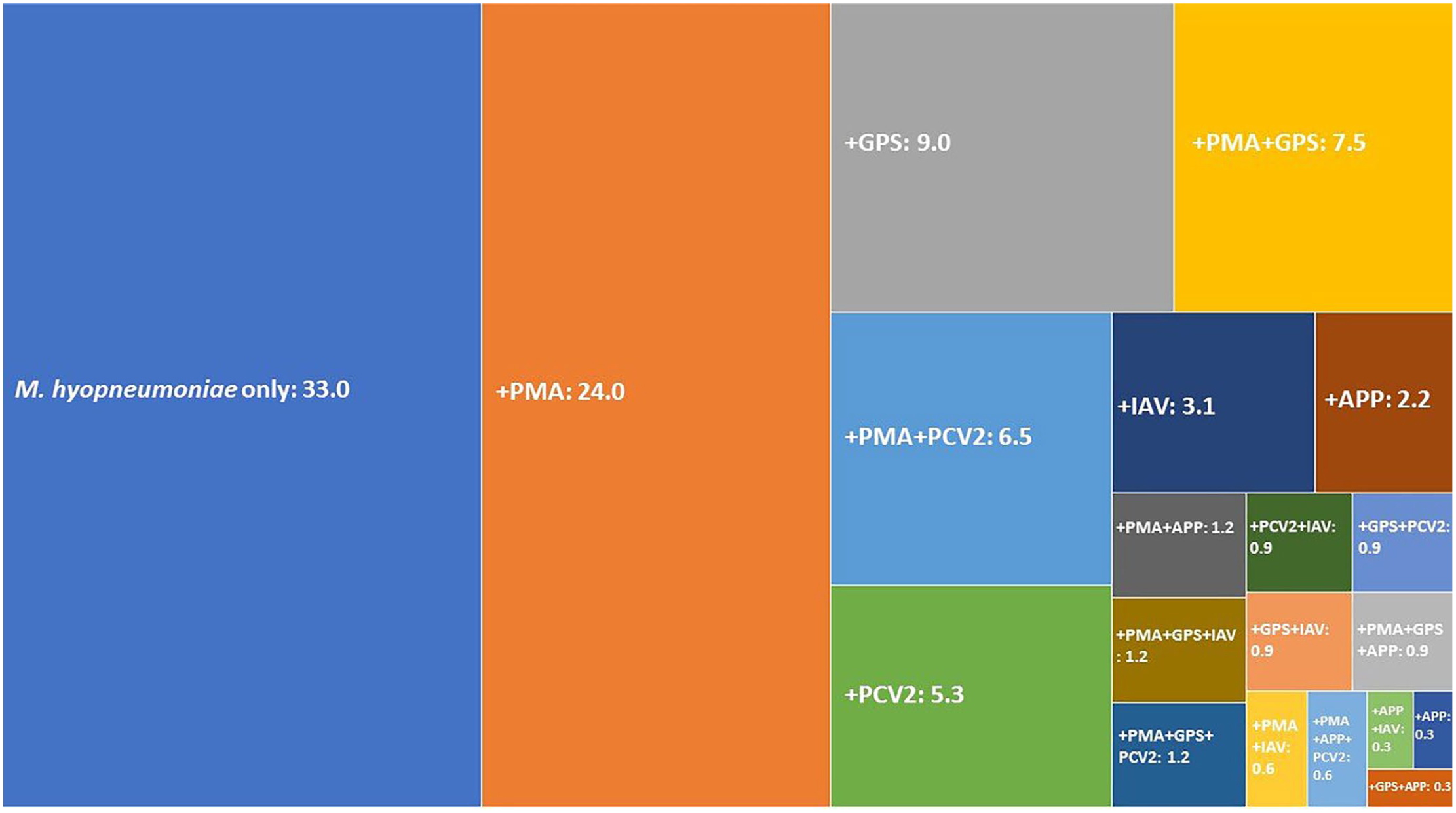
Tree map depicting the percent positive of coinfecting pathogens detected in 321 Mycoplasma hyopneumoniae–positive samples. APP = Actinobacillus pleuropneumoniae; GPS = Glaesserella parasuis; IAV = influenza A virus; PCV2 = porcine circovirus 2; PMA = Pasteurella multocida. + = M. hyopneumoniae plus the other named pathogen(s).
We also evaluated M. hyopneumoniae coinfections according to geographic region. PCV2 was more prevalent in farms from the Southeast region (34.9%) than from farms in the South (22.3%) and Central-West (6.9%) regions (p < 0.01; Pearson chi-square test). G. parasuis was significantly more common in farms from the South (28.2%) than from the Central-West (21.1%) and Southeast (11.6%) regions (p = 0.04; Pearson chi-square test). APP was detected in farms from the Central-West (6.3%) and South (6.8%), but not in the Southeast farms (p = 0.04; Pearson chi-square test). Finally, coinfections were evaluated according to production system: breeding or multipliers, farrowing or nursery (including samples from sows and piglets), fattening, and farrow-to-finish. G. parasuis was more frequent in farrowing or nursery (48.7%) than in the breeding or multipliers and fattening farms (10% and 18.6%, respectively; p < 0.01; Pearson chi-square test). Another interesting finding was the detection of APP only in herds from fattening and farrow-to-finish farms.
Our results clearly demonstrate that M. hyopneumoniae is widely disseminated in Brazilian pig-producing farms, reinforcing previous reports.2,9,17 The prevalence of M. hyopneumoniae is ~39% in the regions surveyed, which is similar to that in other important pig-producing countries; prevalence ranged from a low of 9.3% in Germany 14 to a high of 69.3% in France. 7 We also observed prevalence differences in Brazilian geographic regions (59% in the Southeast, 36% in the Central-West, and 25% in the South), which could be related to vaccination practices and farm types, as reported previously. 2 In addition, the M. hyopneumoniae strains that we detected in Brazilian herds were previously found to be highly clonal and belonged to a few multi-locus sequence types disseminated in the pig-producing chain. 1 Other studies have also reported the high genetic identity of M. hyopneumoniae in specific pig-producing chains and/or geographic regions.8,17,24
Our study also highlights M. hyopneumoniae coinfection with 5 other important respiratory pathogens: P. multocida, G. parasuis, APP, PCV2, and IAV. Coinfection with P. multocida was more frequent (43.9%) than with other pathogens, consistent with its common occurrence as a secondary pathogen in pigs with EP. 16 G. parasuis was also detected frequently (22.1%), as reported elsewhere. 14 These 2 bacteria have been identified commonly in commercial pig herds in Brazil and other countries.18,19,21,22 Coinfection with APP was a less common finding. Other studies have demonstrated dual infection based on serologic analyses.2,9 Further, the prevalence of APP that we observed (5.6%) was mainly in fattening pigs, as reported in herds from Germany. 10 This pathogen probably spreads in sow herds and is an infection source to piglets. 2 In chronically infected herds, APP is more frequent in fatteners given the additional infection risk factors, such as crowding, mixing, and adverse climatic conditions. 12
We detected coinfections with PCV2 and IAV in all production systems and in the 3 regions of the country evaluated in our study. There was a predominance of PCV2 with a prevalence of 15.6%, (6.9–34.9%, according to the geographic region). A study has demonstrated a prevalence of PCV2 of ~25% in swine herds from the Rio de Janeiro State in Brazil. 5 In addition, we detected IAV in 7.2% of our samples. This finding reinforces the occurrence of IAV in commercial Brazilian swine herds, as demonstrated previously with serologic testing.4,21 Various swine viruses have been demonstrated to circulate intermittently and continuously in pig populations from the same production system. 20
Overall, we found that coinfections with other pathogens were common (67%), as has been reported in other studies.15,22 Coinfection by M. hyopneumoniae and other microorganisms probably occurs as a result of the similar processes of adhesion to, and infection of, the animals’ bronchial cells. Further, the occurrence of M. hyopneumoniae in the lungs of animals increases the L-fucose composition significantly, which enhances the adherence of P. multocida to epithelial ciliated cells of the respiratory tract. 18
To compare the coinfection results with the severity of the disease, some clinicopathologic data were also obtained. Dyspnea and cough were the main clinical signs; pulmonary consolidation was detected in >50% of the evaluated animals. No statistical differences between mono-infection (only M. hyopneumoniae) and coinfection (M. hyopneumoniae and one or more microorganisms) were observed (data not shown). Histologic examination was not carried out on the samples, preventing a more complete analysis of the lung lesions. Hence, our survey has limitations in associating lesions with infection by a specific pathogen. 3 Clinical signs and macroscopic lesions are suggestive of, but not exclusive to, M. hyopneumoniae infection, as reviewed previously. 13
Although we had a large sample size and surveyed the main Brazilian pig-producing regions and systems, we did not employ a formal random sampling procedure, and animals were sampled by convenience in a limited number of selected swine herds in Brazil. Therefore, the frequency of M. hyopneumoniae in the pig herds studied, as well as of the coinfection with other pathogens, should be considered an estimate of their true frequencies in Brazil.
Footnotes
Acknowledgements
We thank the staff of Simbios Biotecnologia and the Molecular Diagnostics Laboratory of ULBRA for technical support.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Eder Balestrin and Jonas M. Wolf were supported by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior–Brasil (CAPES)–Finance Code 001, and Lucas M. Wolf by the Fundação de Amparo à Pesquisa do Rio Grande do Sul (FAPERGS). Nilo Ikuta and Vagner R. Lunge were financially supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq; process 331 313564/2014-0; 311010/2017-2).
