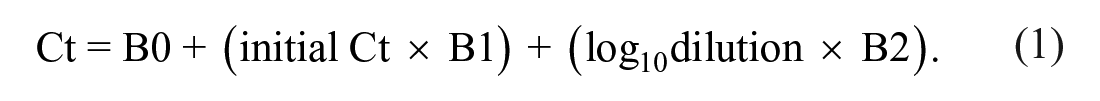Abstract
Influenza A virus (IAV) active surveillance in pigs prior to weaning is commonly conducted by collecting individual samples, mostly nasal swabs. Recently, the use of udder skin wipes collected from lactating sows was identified as an effective sampling method to indicate IAV status of suckling piglets prior to weaning. However, there is limited information on the effect of pooling multiple udder wipes on the ability to detect IAV. We evaluated the effect of pooling 3, 5, or 10 udder wipes on the sensitivity of detecting IAV and compared the results with testing the wipes individually. The likelihood of detecting positive udder wipes decreased with pooling when the initial positive cycle threshold value was ≥31.5; pooling of up to 3 samples could be performed without affecting sensitivity significantly. Our results support pooling of udder skin wipes to conduct surveillance of IAV in pigs prior to weaning.
Influenza A virus (IAV; Orthomyxoviridae, Alphainfluenzavirus) is endemic in swine and causes respiratory disease that results in productivity and economic losses. 2 Pigs prior to weaning are one of the main subpopulations that help maintain infections endemic in breeding herds. Cost-effective surveillance programs directed at detecting IAV in piglets prior to weaning are needed to design effective IAV control and elimination programs, in particular if the desired outcome is to wean an IAV-negative pig. Traditionally, nasal swabs have been used as the reference sample to monitor pigs, not only to detect but also isolate IAV. 5 A 2019 study identified udder skin wipes collected from lactating sows as a suitable sample type to monitor litters of pigs prior to weaning. 1 Udder skin wipes contain the nasal and oral secretions deposited on the skin of the lactating sows by suckling piglets and thus represent the litter status. Following the report of the use of udder wipes for IAV sampling, udder wipes have become a popular sample type to detect IAV in breeding herds in North America (Torremorell M, pers. comm., 2021). Pooling of samples prior to reverse-transcription PCR (RT-PCR) assays is common for some diseases, 6 given that it can help reduce diagnostic laboratory fees or increase the number of samples collected and tested at the same cost. Although studies have been performed using pooling of nasal swabs,1,4 information on pooling udder skin wipes and its effect on sensitivity for IAV detection is missing. Therefore, the aim of our study was to assess the impact of pooling udder wipes collected from lactating sows on the sensitivity of IAV detection by RT-PCR compared to individual wipes.
Udder skin wipes collected from sows nursing piglets close to weaning (~21 d old) were selected from a previous study. 3 Briefly, 45 lactating sows in a 3,000-sow farm located in the midwestern United States were selected randomly. Sampling was performed as described previously, 1 using 8 × 8 cm sterile gauze pads with 8 mL of Dulbecco modified Eagle medium (Gibco) supplemented with 4% of 7.5% bovine serum albumin, fraction V, and 1× antibiotics (Anti-Anti, 100× [composed of 10,000 units/mL of penicillin, 10,000 µg/mL of streptomycin, 25 µg/mL of amphotericin B]; Gibco). Samples were placed in a cooler box containing ice packs and transported to the laboratory for processing and testing. Samples were processed for RNA extraction (Ambion MagMAX 1835 viral RNA isolation kit; Applied Biosystems) and tested by RT-PCR to detect the IAV matrix gene. 7 Samples with a cycle threshold (Ct) value <35 were considered positive. Of 45 samples, 19 samples tested positive for IAV with Ct values of 27.5–33.8. These samples were used as the starting material to evaluate the sensitivity of pooling. Negative samples were combined to serve as the IAV-negative control. To evaluate the effect of pooling, 0.1 mL of the positive samples were diluted in 0.2 mL of sample solution for 1:2 ratio, in 0.4 mL for 1:4 ratio, and in 0.9 mL for 1:9 ratio, using the negative control udder wipe solution (Fig. 1) that had been confirmed IAV negative by RT-PCR. The 19 IAV-positive undiluted samples and their respective pools of 1:2, 1:4, and 1:9 were individually analyzed by RT-PCR, as described above.
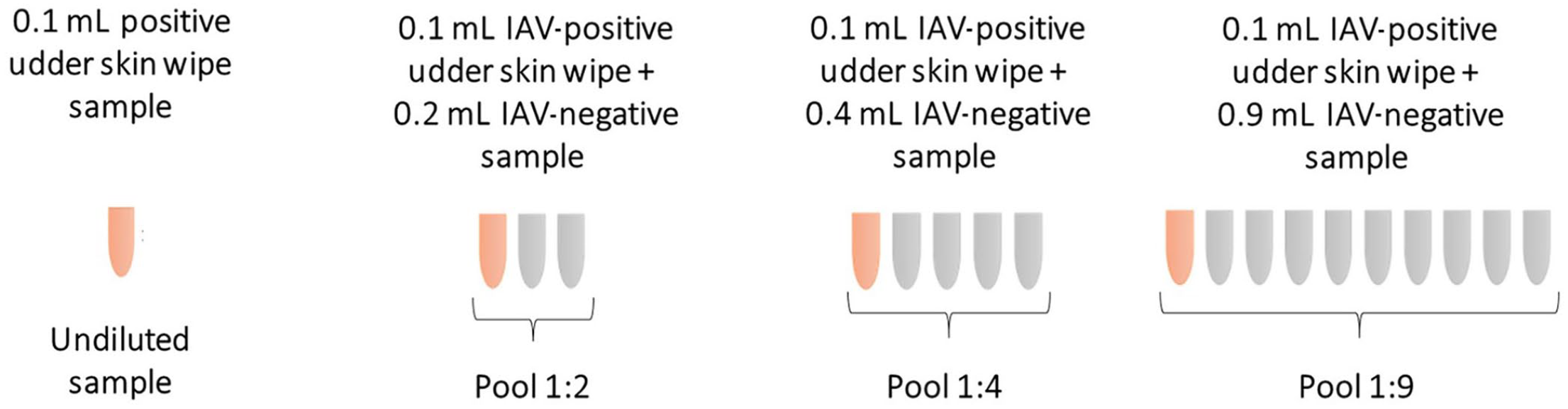
Summary of pooling procedures using positive (orange) and negative (gray) influenza A virus (IAV) udder skin wipes indicating the pooling ratio of no pooling (undiluted sample), 1:2, 1:4, and 1:9.
To calculate sensitivity, results from undiluted samples were considered reference values, and sensitivity was calculated as the proportion of positive samples after pooling divided by the total number of positive samples. The 95% confidence interval (CI) was estimated using the Jeffreys interval method. Briefly, samples were divided into 2 groups using the median of the undiluted samples, and sensitivity was calculated for the high- and low-Ct groups. A polynomial regression of a first-order model was used to estimate the increase in Ct values by each unit of dilution (equation 1). Prior to inclusion in the model, undiluted samples and dilutions 1:2, 1:4, and 1:9 were converted to the log10 scale: 0, 0.47, 0.70, 1.00, respectively. The number of dilutions that could be performed before reaching the positive cutoff (Ct < 35) was calculated, with B0 = intercept, and B1 and B2 = coefficients associated with predictors, respectively:
Pooling of the positive udder wipes in pools of 3, 5, and 10 resulted in sensitivities of 84.2%, 68.0%, and 63.0%, respectively (Table 1). As expected, there was a slight increase in the Ct values of the pooled samples compared to the individual samples. The median Ct value of the undiluted positive samples was 31.5.
Sensitivity, mean and median cycle threshold (Ct) value of undiluted samples, and samples pooled in a final ratio of 1:2, 1:4, and 1:9.
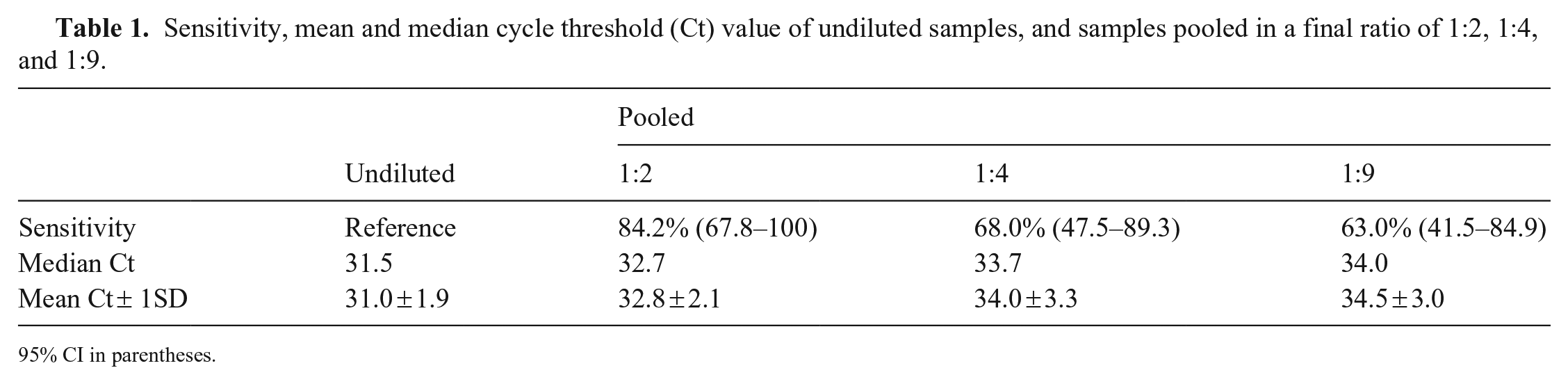
95% CI in parentheses.
There was no difference in sensitivity in the group of samples with low (≤31.5) Ct values (Fig. 2). In the group of samples with high (>31.5) Ct values, sensitivity ranged between 30% (95% CI: 42–98%) and 100% (95% CI: 2–58%), depending on the pooling ratio (Fig. 2).
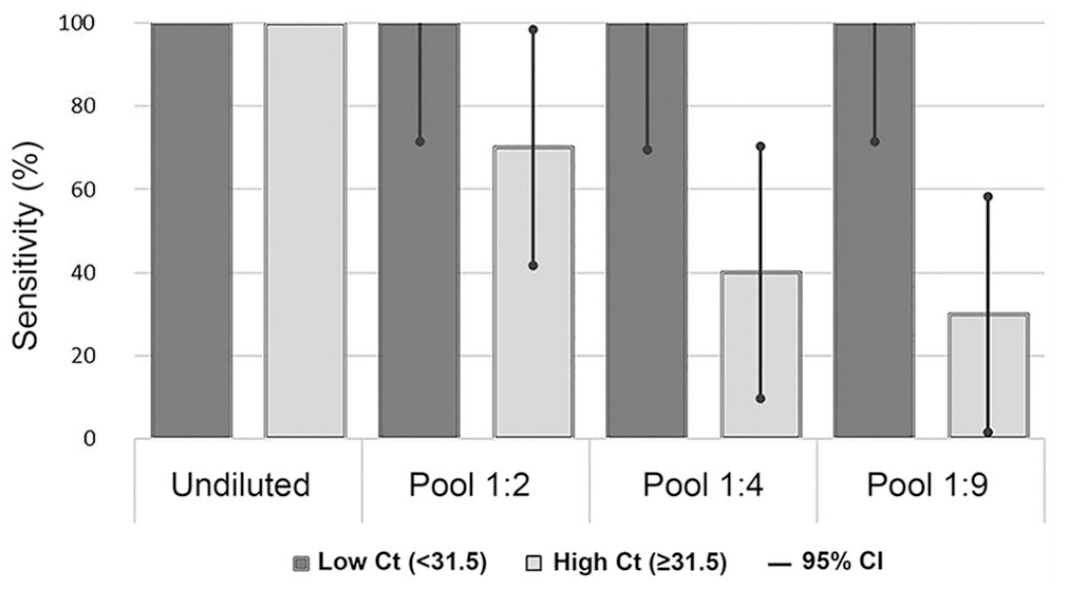
Sensitivity (%) and 95% CI in groups with low (<31.5) and high (≥31.5) initial cycle threshold (Ct) values, based on Ct values from undiluted samples (reference values).
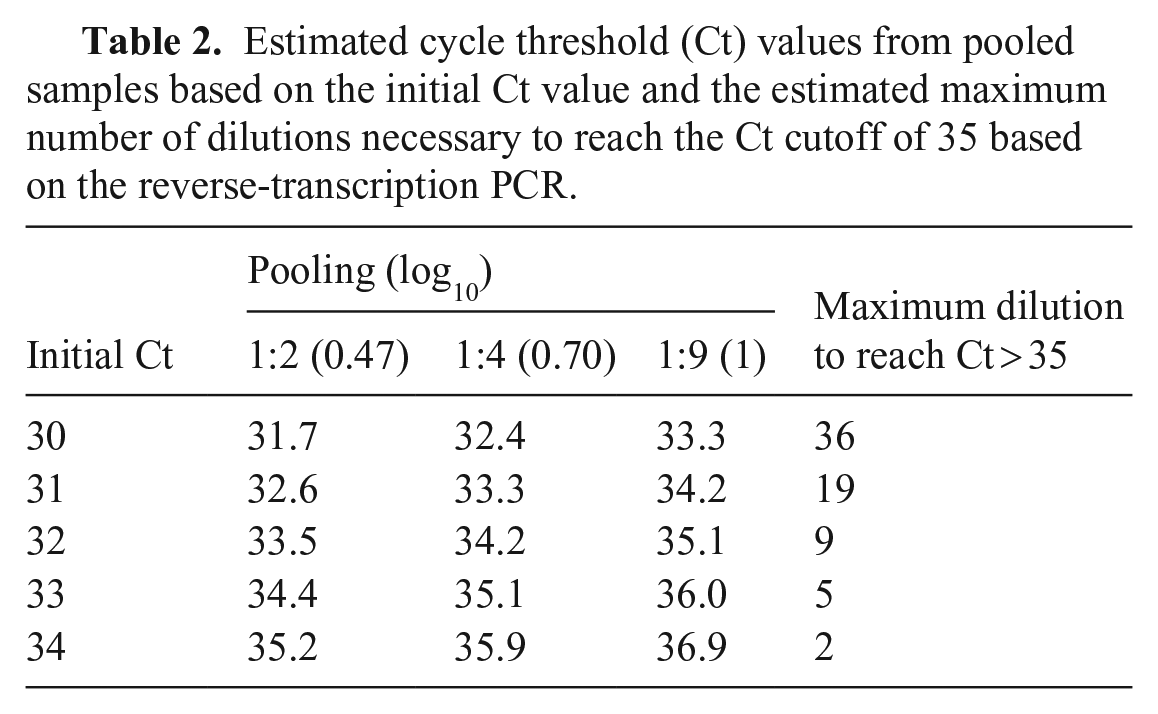
Estimated Ct values after pooling samples varied based on the initial samples’ Ct (Table 2). For samples with Ct = 34, only 2 dilutions were needed to reach the negative cutoff value of 35. In contrast, 36 dilutions were needed when the starting Ct = 30. The intercept from polynomial regression was 3.53. The coefficient associated with initial Ct was 0.89, and the dilution ratio was 3.04.
Estimated cycle threshold (Ct) values from pooled samples based on the initial Ct value and the estimated maximum number of dilutions necessary to reach the Ct cutoff of 35 based on the reverse-transcription PCR.
We conclude that pooling of udder skin wipes can be used to test for IAV by RT-PCR. However, pooling resulted in a slight decrease in sensitivity when pooling ratios were 1:2, and a moderate decrease when pooling ratios were 1:4 or higher. As expected, our study demonstrated that the likelihood of detecting a positive udder skin wipe is correlated with the initial Ct value, which reflects viral load. In the samples used in our study, sensitivity was not compromised when samples with Ct < 31.5 were pooled for up to 10 samples. Instead, sensitivity decreased with pooling of samples with initial Ct values ≥31.5. We understand that, as a cost-mitigation strategy, pooling of samples at the laboratory prior to testing them by RT-PCR is a very attractive proposition. However, pooling of more than 3 samples would significantly reduce the probability of detection, and could lead to false-negative results and the inability to detect litters infected with IAV. The decision on the number of samples in the pool should be made in conjunction with the sampling strategy at the farm level. A decrease in sensitivity caused by increased pooling may be compensated by increasing the number of litters sampled at the farm at the same cost. When increasing sampling, there is a greater likelihood of detecting at least one positive litter in the population, in particular in scenarios of low IAV prevalence when the aim is to detect IAV at the herd level. It is important to note that our study was limited to investigating the effect of pooling on the sensitivity to detect IAV by RT-PCR in udder wipes. It is likely that pooling may affect the sensitivity of virus isolation or other test methods and, as such, this should be further investigated.
To our knowledge, the effect of pooling udder skin wipes to detect IAV by RT-PCR has not been reported previously. The approach described here can help guide sampling strategies and surveillance approaches to detect IAV in litters of pigs prior to weaning.