Abstract
Genetic characterization of the hemagglutinin gene of the 6 selected Thai Swine influenza virus (SIV) isolates (4 H1 and 2 H3 isolates) used in the establishment of a hemagglutination inhibition (HI) assay was analyzed. Based on the phylogenetic analysis, Thai SIVs could be divided into 3 clusters of the H1 viruses (clusters I and II belonging to classical swine H1α, and cluster III belonging to classical swine H1γ), and 2 clusters of the H3 viruses both belonging to human-like 1970s. The serological results indicated that swH1N1-06 (H1 cluster I) is a suitable representative SIV for the HI test antigen to detect H1 SIV-specific antibodies in the Thai swine population, while both swH3N2-05 and swH3N2-07 should be used for Thai H3 SIV-specific antibody detection. The HI test results of swine sera collected from pigs in the 4 highest pig population provinces of Thailand indicated that the percentage of pigs seropositive to swH3N2-07 was highest compared to swH1N1-06, swH1N1-09, and swH3N2-05 (85.4%, 50.1%, 18.6%, and 15.8%, respectively). It should be noted that countries lacking SIV genetic information should be concerned with determining the most suitable HI test antigens to use when performing the tests due to the genetic variation and limited cross-reaction of SIVs. The results of the current study demonstrated that HI tests should be implemented with the suitable field strains as the representative test antigen to ascertain accurate SIV serostatus in Thailand and that test antigens should be genetically analyzed and compared with circulating strains regularly.
Introduction
Swine influenza is an acute respiratory disease in pigs caused by
Retrospective and cross-sectional studies of
Hemagglutination inhibition (HI) is a gold standard for influenza A–specific antibody detection recommended by the WHO.
40
The HI test is highly subtype-specific and requires a suitable representative virus from the locally circulating field strains for the test antigens. In general, SIV is genetically unstable.3,39 Many subclusters have emerged within the different H1N1 and H3N2 SIV subtypes around the world.
21
Historic data indicates that H1N1 SIV is comprised of 2 major lineages: the North American and Eurasian swine lineages.
2
Interestingly, in Thailand, the reassortant H1N1 SIV between the North American and Eurasian lineage has been reported as swine breeds have been continuingly imported from both continents.30,34 Similarly, antigenic and genetic diversity of swine H1N1 and H1N2 viruses in Europe have been documented.
19
This indicates that a constant update of the representative filed strain virus within different regions of the world is required to reflect true SIV serologic data. It is also important that the selected virus have a wide cross-reactivity to antibodies against different SIV strains within the same subtype.1,29 Previous SIV genetic data in Thailand showed that H1 and H3 SIVs circulating between the years 2000–2005 formed at least 2 clusters within each subtype.
34
Data published in 2010 showed that the pandemic H1N1 2009 (pH1N1) virus is now circulating in Thai swine herds.
31
It should be noted that before the introduction of pH1N1 in Thailand, SIV HI test antigens and serological data had not yet been evaluated. The objectives of the current study were 1) to genetically characterize the hemagglutinin (
Materials and methods
Analysis of HA gene
All 38 HA nucleotide sequences of both H1 and H3 Thai SIVs available from the GenBank database were analyzed by multiple alignment and phylogeny construction.
35
Homology analysis of amino acid sequences were evaluated using ClustalW2.
18
The differences in the
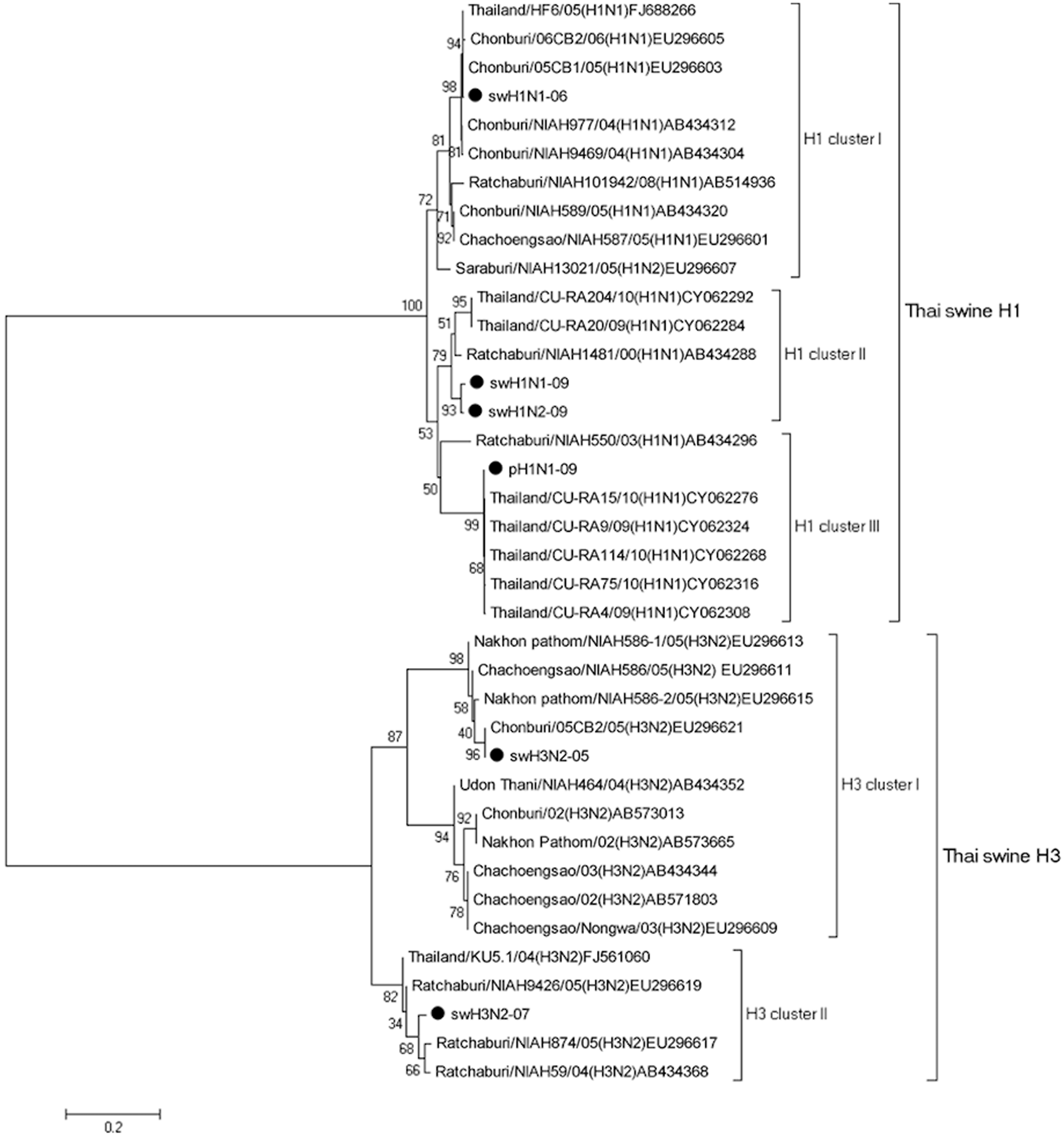
Phylogenetic analysis of the nucleotide sequence of the hemagglutinin (
Rabbit groups used in Swine influenza virus antiserum preparation.
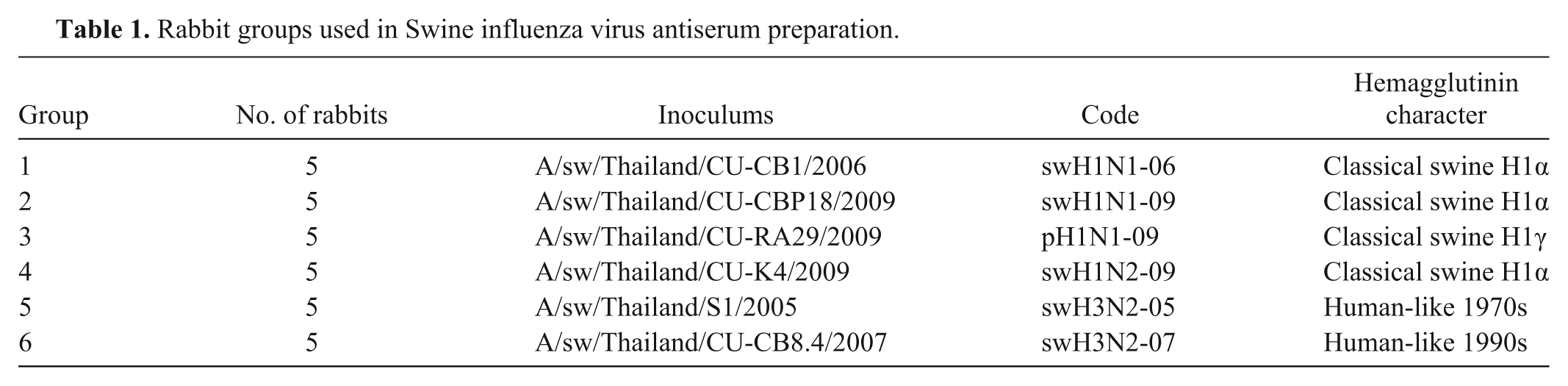
Viruses
The following 6 Thai SIV isolates were evaluated: A/sw/Thailand/CU-CB1/2006 (hereafter, swH1N1-06; H1 cluster I virus), A/sw/Thailand/CU-CB918/2009 (hereafter, swH1N1-09; H1 cluster II virus), and A/sw/Thailand/CU-K4/2009 (hereafter, swH1N2-09; H1 cluster II virus) belonging to classical swine H1α; A/sw/Thailand/CU-RA29/2009 (hereafter, pH1N1-09; H1 cluster III virus) belonging to classical swine H1γ; and A/sw/Thailand/S1/2005 (hereafter, swH3N2-05; H3 cluster I virus) and A/sw/Thailand/CU-CB8.4/2007 (hereafter, swH3N2-07; H3 cluster II virus) belonging to human-like 1990s (Fig. 1; Table 1). All 6 SIVs were obtained from Chulalongkorn University, Veterinary Diagnostic Laboratory (CU-VDL; Bangkok, Thailand). Viruses used for rabbit inoculation and for test antigens in the HI assay were propagated in Madin–Darby canine kidney cells as described previously 12 and stored at −80°C until used.
Rabbit antiserum preparation
In the current study, rabbits were used for SIV antiserum production because rabbits are easy to handle for production of polyclonal antibodies and will produce adequate volumes of high-titer and high-affinity antiserum. The selected 6 virus isolates mentioned above were purified by sucrose gradient ultracentrifugation at 28,000 rpm for 2 hr and resuspended in phosphate buffered saline. Each virus was formalin-inactivated and later used as an antigen for immunization with (ratio 1:1) Freund adjuvant a at 400 HA unit per ml. 15 A total 30 rabbits (H1 and H3 SIV seronegative) were tagged, divided into 6 groups with 5 animals in each group, and intramuscularly injected with killed selected Thai SIV antigen (1 ml each) as shown in Table 1. Each group was housed in separate cages and was provided with commercial food and clean water as reviewed by animal use protocol number 0931049 of Chulalongkorn University Animal Use Committee. Before starting the experiment, rabbit blood was collected from each animal and subsequently used as the negative control sera. After being acclimatized for a week, the animals were immunized as assigned previously every 2 weeks, and sera were collected every week to test for specific HI titer to the inoculated homologous virus antigen. When the HI titers reached ≥1:160, animals were humanely euthanized, and sera were collected and stored at −20°C until used.
Swine serum collection
From June 2008 to May 2009, 850 pig sera were cross section sampled from pigs of various age groups (gilts, sows, finishers, growers, and weaning pigs) in the 4 provinces located in the highest pig density of Thailand (Fig. 2). Pigs from 12 pig farms reporting respiratory symptoms from Ratchaburi (
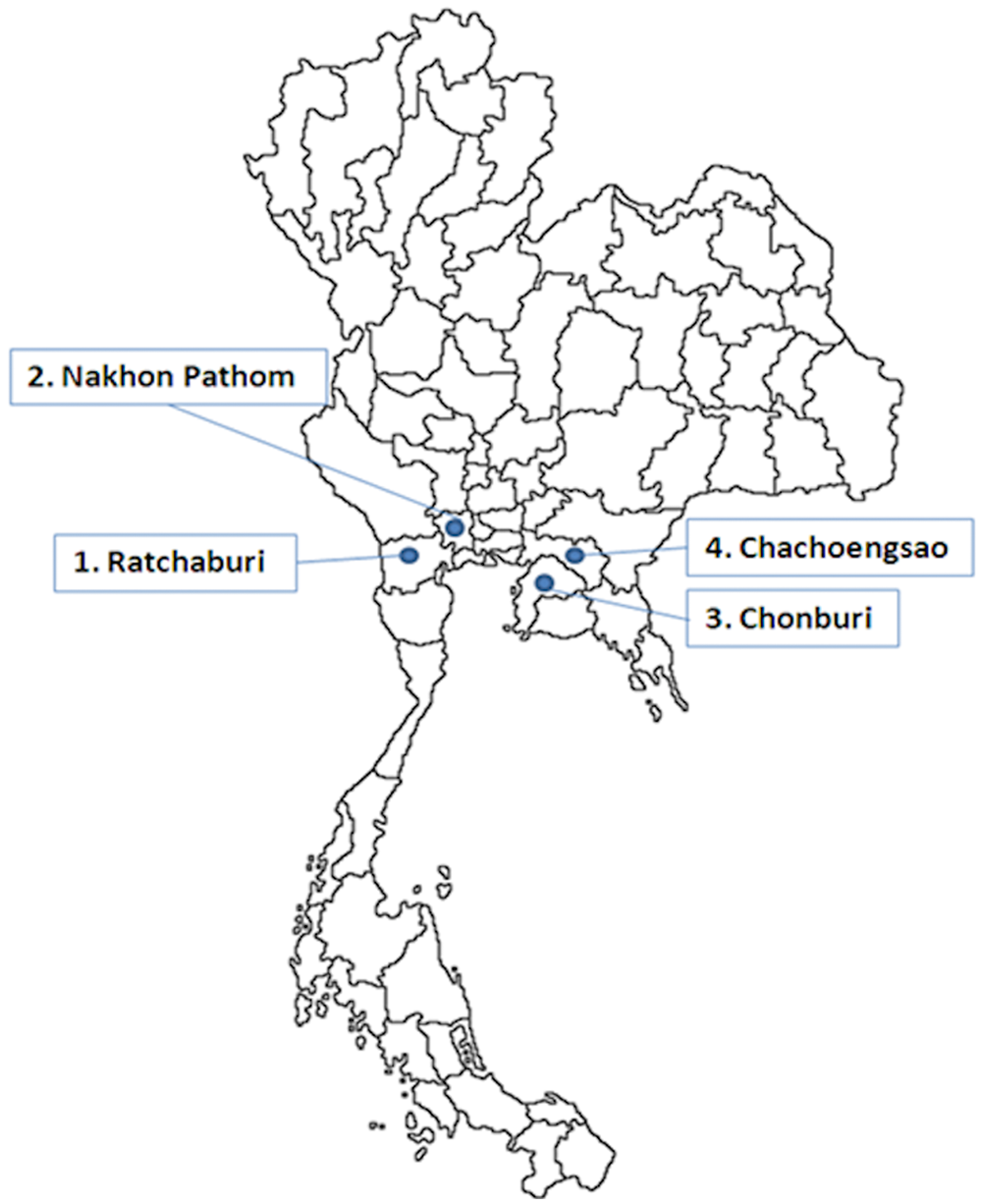
Map of Thailand showing the 4 high pig density provinces included in the current study.
Hemagglutination inhibition
All rabbit and pig sera were subjected to HI testing. Sera subjected to testing with H1N1 SIV were treated with 20% kaolin and absorbed with 50% chicken red blood cells (RBCs). Sera tested with H3N2 virus were treated with receptor-destroying enzyme and absorbed with 50% chicken RBC to remove nonspecific inhibitors of agglutination and natural serum agglutinins. Positive SIV infection was considered when the HI titer measured ≥1:40. 26 Virus antigen used in the HI test was 8 HA unit per 50 µl; 0.5% chicken RBCs was used for titer evaluation based on the OIE 2010 Terrestrial Manual (http://www.oie.int/en/international-standard-setting/terrestrial-manual/access-online/). The SIV antigens used for antibody reactivity of both homologous and heterologous strains in the rabbit sera included swH1N1-06, swH1N1-09, pH1N1-09, swH3N2-05, swH3N2-07, and swH1N2-09 virus. For serologic surveillance from pig sera, 4 isolates (swH1N1-06, swH1N1-09, swH3N2-05, and swH3N2-07) were used as test antigens. Isolate swH1N2-09 was excluded because it was isolated in late 2009 after the period of serum collection.
Enzyme-linked immunosorbent assay
Pig sera with known HI titers against H1N1 (swH1N1-06) and negative for H3N2 (swH3N2-05 and swH3N2-07) were evaluated by a commercial ELISA H1N1 kit. b Similarly, sera with known HI titers against H3N2 (swH3N2-07) and negative for H1N1 (swH1N1-06) were evaluated by a commercial ELISA H3N2 kit. c Ninety-one pig sera were tested for H1N1, and 85 sera were tested for H3N2 according to the protocol recommended by the manufacturers.
Results
Analysis of HA gene
Based on phylogenic analysis, 5 clusters of Thai SIVs were divided into H1 viruses (H1 clusters I–III) and H3 viruses (H3 clusters I and II; Fig. 1). One selected virus in each cluster was selected based on its antigenic property with the exception of the H1 cluster II, which used both swH1N1-09 and swH1N2-09. For homology analysis of 4 H1 viruses, there was approximately 82.2–86.7% homology to each other except for swH1N1-09, which demonstrated 96.4% homology to swH1N2-09. Similarly, swH3N2-05
For amino acid substitution, the HA1 amino acid sequences were compared and showed important residues of the H1 and H3 strains. The amino acid residues 133, 96, and 183 are characterized as host-restrictive residues or PIRs of human H1 influenza virus. All 4 swH1 viruses (swH1N1-06, swH1N1-09, swH1N2-09, and pH1N1-09) were compared to A/sw/NC/00573/2005, a human-like swine isolate, and A/sw/Minnesota/37866/1999, a classical swine lineage showing serum cross-reactivity to the swine viruses from 1930–2004 in the United States (Table 4). Among the 4 swine H1 viruses in the current study, the PIR was similar except for pH1N1-09 at residue 96. The amino acid residue 96 of swH1N1-06, swH1N1-09, and swH1N2-09 is threonine (T), which is similar to A/sw/NC/00573/2005, but pH1N1-09 contains isoleucine (I) similar to A/sw/Minnesota/37866/1999. The amino acid residues 133 and 183 are similar to A/sw/Minnesota/37866/1999 (valine [V] and proline [P], respectively). In addition, amino acid change on the presumed antigenic site (Ca, Cb, Sa, and Sb) of the HA1 region was evaluated in Table 5. A total 19 amino acid residues were different among the 4 H1 viruses. There were 4 residues similar in 3 H1 viruses (swH1N1-09, swpH1N1-09, and swH1N2-09), but different from swH1N1-06 (Ca141, Cb73, Sa162, and Sa155). For the HA H3 subtype, the 8 major potential immunodominant positions (121, 135, 144, 145, 155, 156, 157, and 188) were compared (Table 6). Seven amino acid positions were different, and only an amino acid on residue 156 was similar in swH3N2-05 and swH3N2-07.
Hemagglutination inhibition
For antigenic properties using rabbit antisera, HI titers of each group of the studied rabbits reached ≥1:160 against the homologous virus within 5 weeks post inoculation. Antisera from those rabbits injected with the H1 antigens had no antibody titers to the other 2 H3 subtypes and vice versa. The HI geometric mean titers in each group against H1 and H3 antigens are shown in Table 2 and 3, respectively. Rabbit antisera against swH1N1-09, pH1N1-09, and swH1N2-09 isolates had high cross-reactive HI titers (>1:160) against swH1N1-06. Interestingly, swH1N1-06 antiserum had no cross-reactivity against current H1 isolates. Similarly, swH3N2-05 and swH3N2-07 antisera had no cross-reactivity to the heterologous viral antigen. These findings indicated that swH1N1-06 is a suitable representative isolate for use as the HI test antigens for H1 SIV-specific antibody detection in the Thai swine population. Furthermore, both swH3N2-05 and swH3N2-07 are suitable for use as the HI test antigens for H3 SIV-specific antibody detection in Thailand.
Geometric means of rabbit hemagglutination inhibition (HI) titers evaluated against the 3 selected H1N1 and 1 H1N2
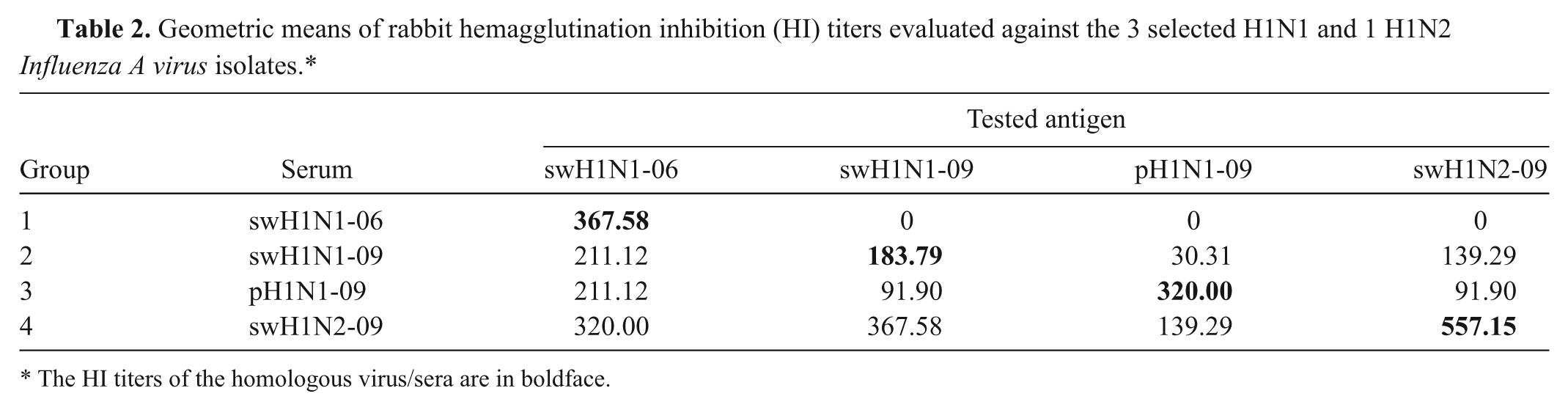
The HI titers of the homologous virus/sera are in boldface.
Geometric means of rabbit hemagglutination inhibition (HI) titers against the 2 selected H3N2
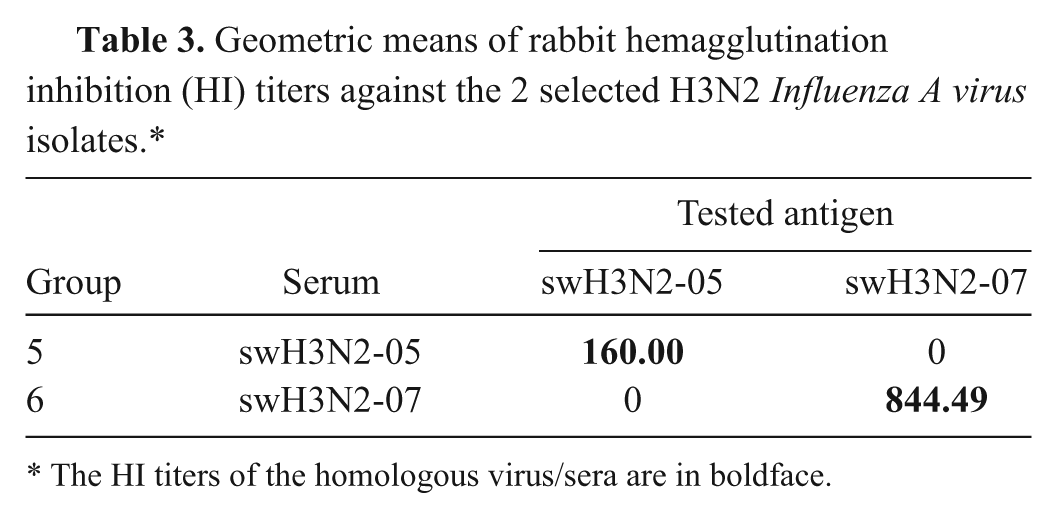

The HI titers of the homologous virus/sera are in boldface.
Amino acid changes in host residues or phylogenetically important regions (PIRs) of the
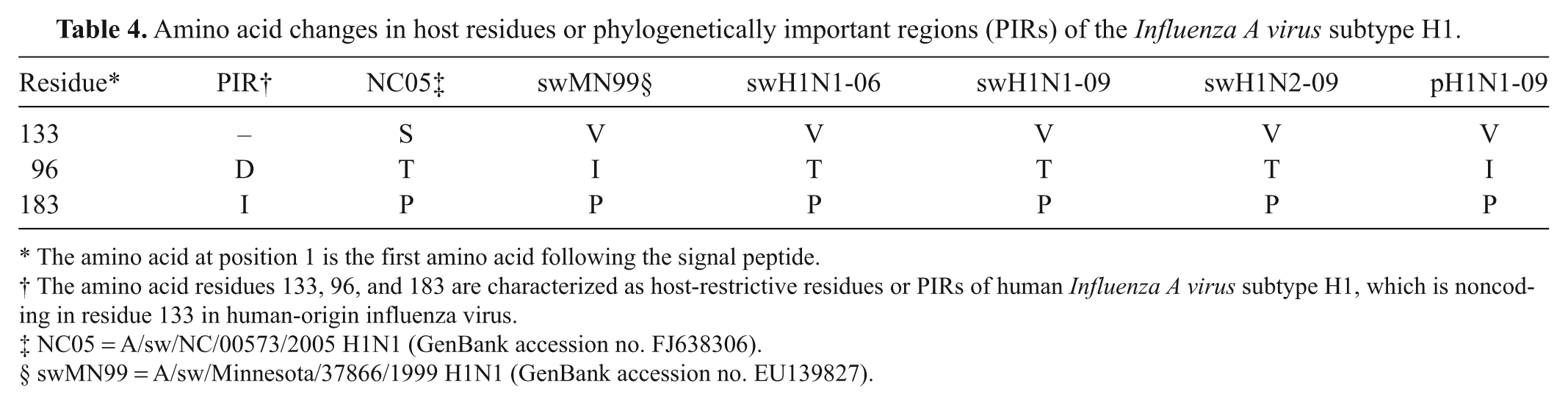

The amino acid at position 1 is the first amino acid following the signal peptide.
The amino acid residues 133, 96, and 183 are characterized as host-restrictive residues or PIRs of human
NC05 = A/sw/NC/00573/2005 H1N1 (GenBank accession no. FJ638306).
swMN99 = A/sw/Minnesota/37866/1999 H1N1 (GenBank accession no. EU139827).
Amino acid changes on antigenic sites of the HA1 region of swine
Eight major potential immunodominant positions of both swH3N2-05 and swH3N2-07.
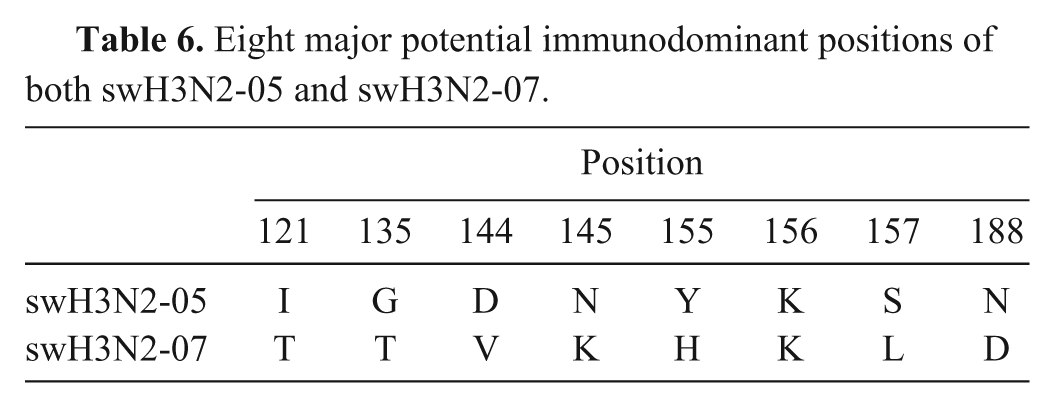

A total of 850 field pig sera from 12 farms were evaluated for HI testing using swH1N1-06, swH3N2-05, and swH3N2-07 as the representative Thai SIV antigens. In addition, the current isolate (swH1N1-09) was also tested for seroprevalence of the previous pig sera. The findings demonstrated that SIV serostatus in pigs during 2008–2009 were seropositive to both H1 and H3 viruses (Fig. 3). The number of pigs seropositive to swH3N2-07 were highest compared to swH1N1-06, swH1N1-09, and swH3N2-05 (85.4%, 50.1%, 18.6%, and 15.8%, respectively; Fig. 3A). In addition, the sow seropositive to Thai swH1N1 was highest compared to growing and weaning pigs (73.2%, 43.9%, and 30.4%, respectively). However, there were no differences in seropositivity in each age group tested against Thai swH3N2 (Fig. 3B).
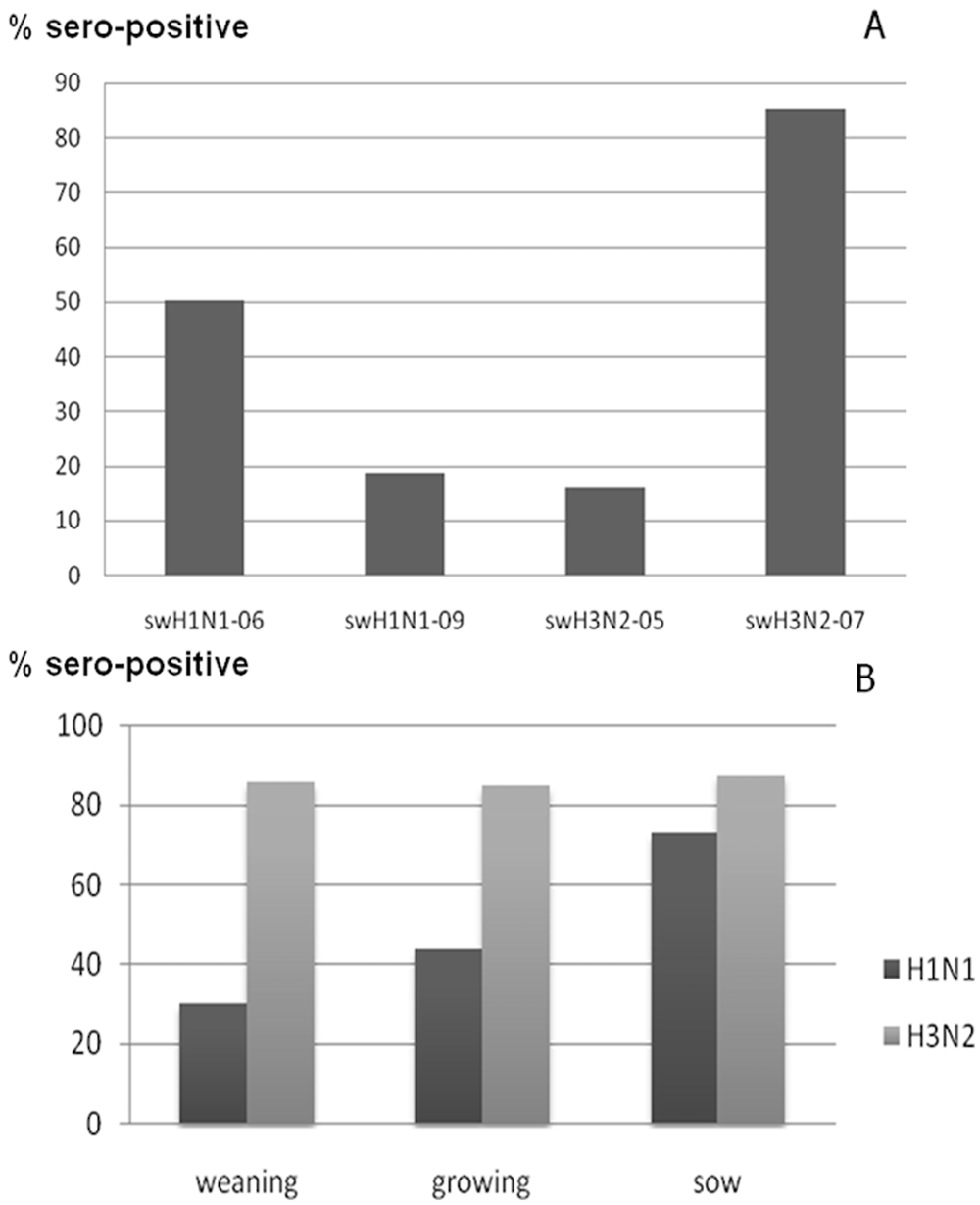
The percentages of seropositive hemagglutination inhibition (HI) test results against 4 selected Thai Swine influenza virus (SIV) isolates (
Commercial ELISA evaluation
A total of 91 pig sera were tested with a H1N1 ELISA kit, b and the results were compared to the results of the HI test using swH1N1-06 virus antigen. A 2-by-2 table of the tests was constructed (data not shown) and evaluated for sensitivity (89.36%), specificity (77.27%), and false-negative (8.5%) and false-positive (22.72%) results, respectively. Similarly, a 2-by-2 table of the test results between the H3N2 ELISA c and swH3N2-05 HI test (data not shown) showed sensitivity (70.83%), specificity (91.89%), false negative (29.16%) and false positive (8.11%), respectively.
Discussion
In general, SIV genetic mutation normally occurs by point mutation, reassortment, and introduction of new viruses into the local area. Unlike North American or European SIVs, Thai SIVs contain mixed genomes from both North American and European lineages. In the United States, swine H1 is divided into 3 clusters, classical H1N1, reassortant H1N1, and H1N2-like.21,38 Interestingly, genetic characterization of the
Antigenic properties by HI testing using specific rabbit antisera were evaluated in the current study. To elucidate H1 cross-reactivity, the representative isolates of subtype H1N1 in each cluster and the H1N2 isolates were selected for evaluation of the HI titers using known rabbit antisera. The results showed that sera from the previous swH1N1-06 (H1 cluster I) inoculation had no cross antibody reactivity with any of the 3 current swH1 in 2009 (H1 clusters II and III) viruses. However, all 3 swH1 rabbit sera in the present study demonstrated cross-reactivity to the previous swH1N1-06 virus. It should be noted that for routine serosurveillance using HI testing in each country, the specific endemic subtypes isolated in the local area having a broad cross-reactivity must be used for the test to obtain the most accurate results.1,29 In the current study, swH1N1-06 could be used as the representative virus for Thai H1 subtype HI test because it showed one-way cross-reactivity to other H1 viruses. For H3N2 subtype, the rabbit sera showed no cross antibody reactivity in either selected H3N2 isolates. Interestingly, both swH3N2-05 and swH3N2-07 viruses were isolated from the same farm in 2005 and 2007, respectively. However, the swine serum collected during this studied period showed different seroprofiles, with a high seroprevalence for swH3N2-07. The results suggest that the virus could have genetic mutations and become endemic in the original farm. The virus antigens used for HI testing and genetic monitoring locally, therefore, should be evaluated regularly.
From June 2008 to May 2009, serologic surveillance in the present study showed that all 4 selected provinces in Thailand had higher H3N2 seropositive percentages than those of the H1N1 subtype (swH3N2-07: >70%; swH1N1-06: >40%). It should be noted that pH1N1 emerged in Thai pigs in late 2009, 31 and the current seroprofile of the Thai SIVs could possibly be different from the current study. Continuous monitoring of SIV status in both virology and serology is suggested.
Based on the farm history, all 12 selected pig farms in the present study showed respiratory problems concurrently with the presence of
It should be noted that SIV has gained significant attention because of its public health concern, particularly, after the emergence of the pH1N1 in 2009. Moreover, this pandemic flu showed continuing reassortant with endemic SIV in pig populations worldwide.7,9,17,22,28,32,37,42 Similarly, cocirculation of both pH1N1 and endemic Thai swine H1N1 viruses was found in the same farm with no reassortant virus. 31 However, a novel reassortant virus could possibly emerge without notice. Based on intensive surveillance, the pandemic reassortant virus was demonstrated in Thailand. 13 It should be noted that SIV-infected pigs may show only acute mild respiratory distress that could be overlooked by farmers or veterinarians. In addition, human to pig transmission or vice versa may occur without notice. The current study has sought to demonstrate the benefit of HI testing, which could be useful for interspecies transmission surveillance.
The current study illustrates the importance of global swine influenza surveillance because it demonstrates that the use of SIV test antigens from nonendemic sources should be carefully scrutinized, as the genetic and antigenic variation of SIV might not be suitable for the surveillance of local SIVs. In the present study, the genetic and antigenic variation of antigens in different assays was apparent when compared using commercial ELISA test kits with the established HI tests. Therefore, evaluation of commercial test kits from other countries is necessary to ascertain current epidemiology data due to the genetic and antigenic differences of the circulating viruses.
In general, nucleotide sequences encode the amino acid sequences most related to protein function. The PIRs suggesting possible host adaptation
39
were located at the amino acid residue 96 of the H1
In conclusion, results of the current study demonstrated that at least 5 clusters of SIVs circulated in Thailand had a high prevalence of antibodies against H3N2 virus during the study period. In addition, general SIV data including genetic characterization, virology, and serology are of importance for disease diagnosis, prevention, and control in order to reduce spreading to human beings and other species. Due to the limited genetic, virologic, and serologic data on Thai SIVs in the past 10 years, the results from the present study provide necessary data for better disease diagnosis of Thai SIVs. Such data can be modified for SIV diagnosis in other countries as well. Continuous surveillance and monitoring of
Footnotes
Acknowledgements
The authors thank Drs. Padet Tammaruk, Suparlark Nuntawan Na Ayudhya, Sittikorn Triyarat, and Suwarak Wannaratana for their assistance in swine sample collection; Drs. Wijit Bunlunara, Theerayuth Kaewamatawong, Patchareeporn Nilwilai, and Kanana Rungprasert for their assistance in rabbit blood collection; and Duangdearn Pragayroongnamtip and Ranida Tuanudom for technical assistance.
a.
Sigma-Aldrich, St. Louis, MO.
b.
HerdChek H1N1 ELISA, IDEXX Laboratories, Westbrook, ME.
c.
HerdChek H3N2 ELISA, IDEXX Laboratories, Westbrook, ME.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported partly by grants from National Research Council of Thailand (NRCT, Thailand), Emerging Health Risk Cluster, the Higher Education Research Promotion and National Research University Project of Thailand, Office of the Higher Education Commission (HR1160A-55), the Rachadapiseksompoch Endowment Fund, Emerging and re-emerging diseases in animals, Research Unit, Chulalongkorn University. This work has also been funded in part with the federal funds from the National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Services, under contract no. HHSN266200700007C. Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the NIH.
