Abstract
Numerous canine papillomaviruses (CPVs) have been identified (CPV1–23). CPV1, 2, and 6 have been associated with inverted papillomas (IPs). We retrieved 19 IPs from 3 histopathology archives, and evaluated and scored koilocytes, inclusion bodies, giant keratohyalin granules, cytoplasmic pallor, ballooning degeneration, and parakeratosis. IHC targeting major capsid proteins of PV was performed, and CPV genotyping was achieved by PCR testing. Tissue localization of CPV DNA and RNA was studied by chromogenic and RNAscope in situ hybridization (DNA-CISH, RNA-ISH, respectively). IPs were localized to the limbs (50%), trunk (30%), and head (20%), mainly as single nodules (16 of 19). In 15 of 19 cases, immunopositivity was detected within the nuclei in corneal and subcorneal epidermal layers. PCR revealed CPV1 in 11 IPs and CPV2 DNA in 3 IPs. Overall, 14 of 17 cases were positive by both DNA-CISH and RNA-ISH, in accord with PCR results. A histologic score >5 was always obtained in cases in which the viral etiology was demonstrated by IHC, DNA-CISH, and RNA-ISH. IHC and molecular approaches were useful to ascertain the viral etiology of IPs. Although IHC is the first choice for diagnostic purposes, ISH testing allows identification of PV type and the infection phase. RNA-ISH seems a promising tool to deepen our understanding of the pathogenesis of different PV types in animal species.
Keywords
Inverted papillomas (IPs) are uncommon papillomavirus-induced skin lesions consisting of epidermal endophytic papillary projections extending into the dermis, with typical cup- to dome-shaped gross appearance and often central umbilication. 11 Approximately 20 cases of canine IPs have been described in published articles (Suppl. Table 1).2,4,10,14–17,22,27,28,30 In the first report in 1988, the viral etiology was identified by immunohistochemistry (IHC), with the use of papillomavirus (PV) group–specific antigens, and transmission electron microscopy (TEM). 4 Of note, the negative result of a DNA in situ hybridization (ISH) assay, using a probe based on the sequences of PVs identified at that time, suggested the presence of a new PV. Furthermore, the number of cases included in the study (n = 5) was too small to draw a hypothesis on the epidemiology of the disease and to characterize the lesions. After 2 decades, a few more papers presented data on canine IP: other PVs (canine papillomavirus 1 [CPV1; Lambdapapillomavirus 2, previously known as canine oral papillomavirus], CPV2, and other unknown PVs) were invoked as causative agents of IPs in 4 dogs 16 ; 3 cases of IP among 24 papillomatous cutaneous lesions were diagnosed over 10 y (2001–2011). 2 All of the other reported cases are unique descriptions of IPs in different breeds, some of them seen as multiple grouped lesions and others showing transformation to in situ carcinoma.14–17,22,23
To confirm the viral etiology of IPs, various techniques have been used, including PCR to prove the presence of viral genome, histopathology (H&E) to detect the classic inward neoplastic tissue growth and cytopathic effects, IHC to detect L1 viral antigen,4,30 and TEM to visualize viral particles within the lesions.4,23,27 A DNA-ISH technique to detect CPV in IPs has only been mentioned once in the literature; however, the hybridization mixture used by the authors was prepared using cloned canine oral papillomavirus DNA and no positive signal was detected. 4
Only a small number of CPVs belonging to different genera have been fully sequenced. 17 Various clinical presentations and biologic behaviors are described for different IPs, including multiple nodules,14,15,17,22,27 involvement of the ventral abdomen, inguinal region,2,4,14–17,22,27 footpads,10,16,23,30 or other anatomic locations,16,17,27,28 but also comprising spontaneous regression 16 and transformation into squamous cell carcinoma.10,28,30 However, the genome of the involved PVs has been sequenced only rarely, and therefore it is unknown whether the PV type could influence localization or clinical behavior. To date, only CPV1, CPV2, and CPV6 have been associated with IPs in dogs (Suppl. Table 1).10,16,17,22,28,30
Given that data in the literature are rather fragmentary, we examined the clinical and histopathologic findings of IPs in a larger series of cases; furthermore, we investigated the viral etiology of IPs through IHC and ISH techniques.
Materials and methods
Sample collection and histopathology
Cases with a histologic diagnosis of IP were retrieved retrospectively from 3 histopathology archives in 2016 and 2017. Three veterinary pathologists (M. Orlandi, M. Vascellari, F. Albanese) reviewed the slides independently and agreed on the selection of 19 IP cases with complete signalment, history, clinical data, and anatomic location of the lesion(s) (Table 1). Several breeds were represented, including 1 of each of the following: American Staffordshire, Beagle, Bolognese, Border Collie, Bull Terrier, Doberman Pinscher, French Bulldog, Golden Retriever, Italian Bracco, Labrador Retriever, Weimaraner, and 7 cross-breed dogs. The mean age was 5.3 y; for 7 dogs, the age at first presentation was <2 y. There were similar numbers of females and males. Nine nodules were localized on the limbs, 6 on the trunk, and 4 on the head (Fig. 1A). Nodule size ranged from a few mm to 3 cm; some nodules were ulcerated or umbilicated (Fig. 1B), and growth was variably fast. Multiple nodules were reported in 3 dogs: a Beagle, an American Staffordshire, and a cross-breed dog; in the first case, the nodules were on 2 limbs, whereas in the other 2 cases they were grouped on the abdomen.
Signalment and clinical data for 19 canine inverted papillomas.
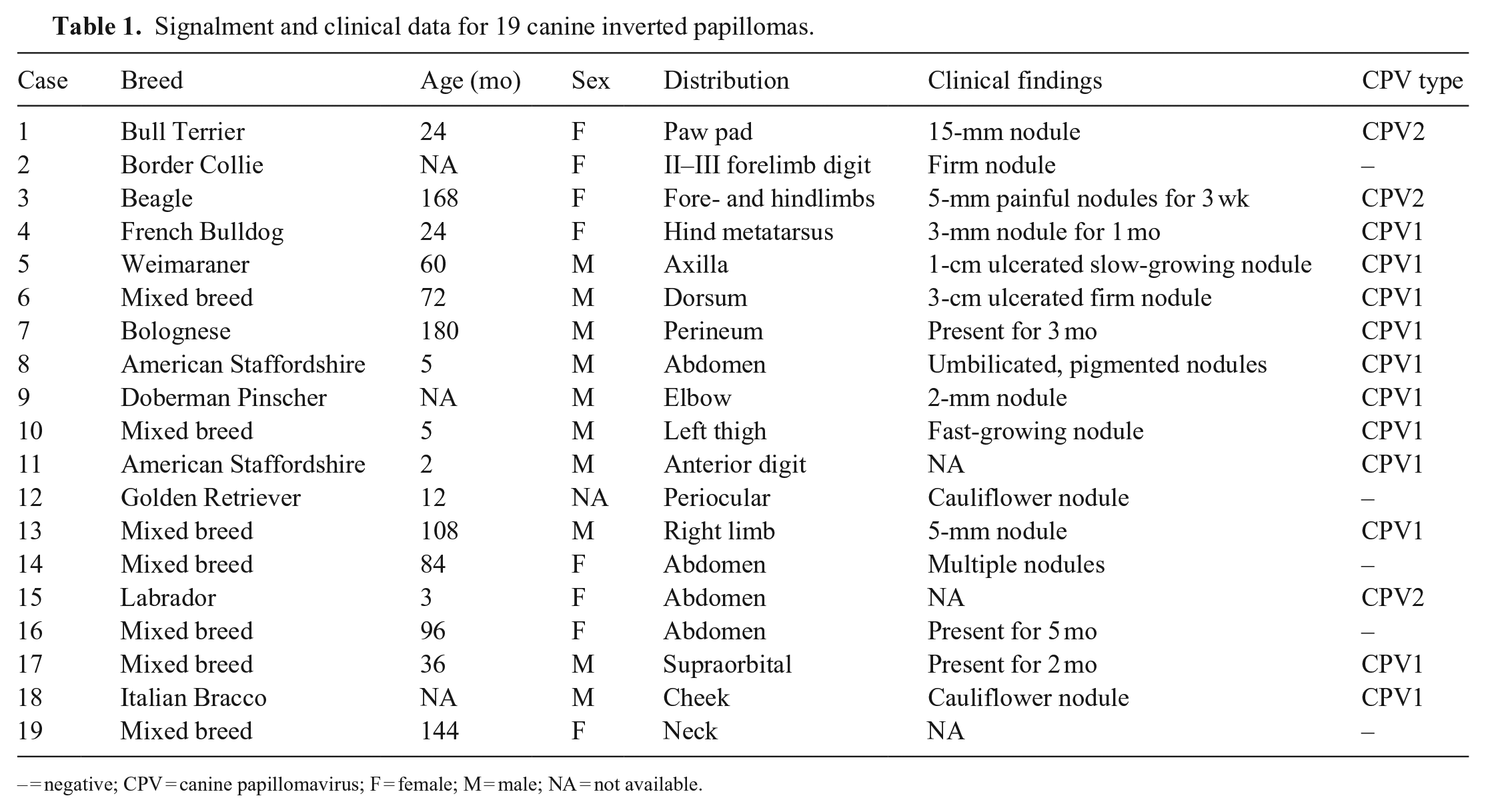
– = negative; CPV = canine papillomavirus; F = female; M = male; NA = not available.
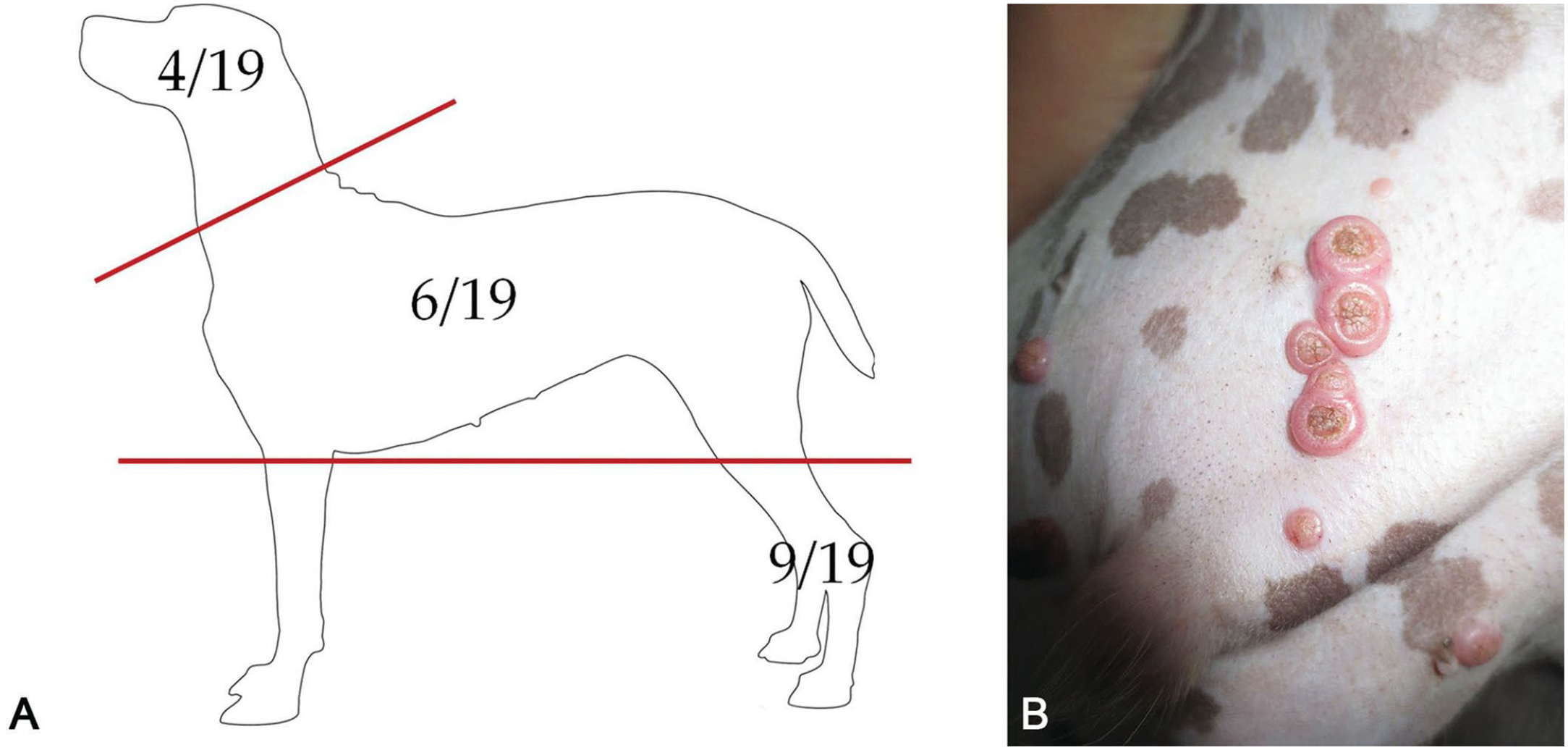
Canine inverted papilloma.
From all cases, 5–10-μm thick tissue sections were cut serially: 1 section was routinely stained with H&E; the other 10- and 5-μm thick sections were used for PCR, IHC, DNA-CISH, and RNA-ISH (RNAscope; ACD) assays. To prevent DNA cross-contamination, the microtome blade holder was covered with a new piece of adhesive tape and a new blade was used for each sample. Sections for DNA-CISH and RNA-ISH were mounted on adhesive glass slides. H&E-stained sections were evaluated for the type of growth and the presence of inclusion bodies, koilocytes, giant keratohyalin granules, slate-gray cytoplasmic material, ballooning degeneration, and parakeratosis. 11 These changes were scored semi-quantitatively as follows: 0 = absence, 1 = mild (presence of the change in <10% of the section), 2 = moderate (presence of the change in 10–60% of the section), 3 = marked (presence of the change in >60% of the section).
IHC
Cases were evaluated for the detection of PV using a mixture of mouse monoclonal antibodies generated against major capsid proteins of bovine PV1 and human PV16 (ab-2417; Abcam), which recognizes multiple PV types. 18 Briefly, antigen unmasking was carried out at 120°C for 3 min in a pressure cooker with Tris-EDTA buffer (1.2 g/L Tris, 0.36 g/L EDTA, pH 9.0). Sections were pretreated for 10 min with 1% H2O2 in 0.1 M PBS (pH 7.4) to quench endogenous peroxidase activity and then blocked for 30 min at room temperature (RT) in PBS with 2% normal horse serum (PK-7200; Vector Laboratories) and 0.05% Triton X-100. Sections were incubated overnight at 4°C with the primary antibody (dilution 1:100). After washing in PBS, incubation with a secondary universal biotinylated anti-mouse/rabbit antibody (PK-7200; Vector Laboratories) was performed for 1 h at RT. Staining was visualized with a diaminobenzidine (SK-4105; Vector Laboratories) solution under a light microscope (Eclipse 80i; Nikon) and slides counterstained with hematoxylin. Positive controls, consisting of tissue from canine oral papilloma that showed strong and diffuse nuclear immunolabeling for PV, were included in each run.
Molecular biology analyses
DNA extraction
DNA was extracted from formalin-fixed, paraffin-embedded (FFPE) sections (QIAamp DNA FFPE tissue kit and deparaffinization solution; Qiagen) following the manufacturer’s instructions. DNA was eluted in 100 µL and stored at −20°C until analyses were performed.
For primer specificity, the entire Canis familiaris (taxid: 9615) genome was studied to avoid nonspecific host genome amplification. Moreover, to validate the efficiency of DNA extraction and its integrity, primers were designed to amplify a 212-bp amplicon on the reference gene corresponding to ribosomal protein RPL15 of Canis familiaris (NG_030479.2). 25
PCR
Primers sets were designed (Primer 3; NCBI) using the major capsid protein L1 gene of CPV1, CPV2, and CPV6 reference genomic sequences (Suppl. Table 2). PCR was undertaken in a thermocycler (iCycler; Bio-Rad) using a thermostable DNA polymerase (Wonder Taq Hot Start; EuroClone) using 2.5 µL of extracted DNA. Three long oligos spanning CPV1, CPV2, and CPV6 amplicons were synthesized (Merck Life Science; Suppl. Table 3) based on GenBank CPV reference genomes and then used as positive controls. Each PCR assay also included negative controls, to ascertain the specificity of the reaction. All PCR products were analyzed by electrophoresis in 1% agarose gel containing ethidium bromide with a quantitative 100-bp DNA marker (SharpMass 100; EuroClone).
Sequence analysis
Amplicons of expected length were submitted for sequence analysis (BMR Genomics, Padova, Italy) by the Sanger method. 24 Nucleotide sequence data were subjected to BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi) analysis to confirm the CPV type.
CISH
Probes for DNA-CISH analysis were generated by PCR on CPV-positive DNA identified previously, using a DNA labeling kit with digoxigenin-dUTP. Briefly, dig-labeled DNA probes were synthesized (HotStarTaq DNA polymerase; Qiagen) with the addition of 200 µM DIG DNA labeling mix (Roche). The PCR protocol consisted of 30 amplification cycles using CPV1 and 2 specific primers, followed by 10 amplification cycles after the addition of 2 units of Taq polymerase (@Taq; EuroClone). The purity and concentration of the probes were quantified by running an aliquot on 2% agarose gel with a quantitative 100-bp DNA marker (SharpMass 100; EuroClone).
DNA-CISH for CPV1 and CPV2 was performed on the BenchMark Ultra platform (Ventana, Roche). FFPE tissue sections were deparaffinized, rehydrated, and permeabilized as described below. Target site unmasking was performed by incubating the slides 3 times in cell-conditioning buffer (Ultra CC2; Roche) at 86°C, followed by ISH protease 3 treatment, at 36°C for 16 min. Denaturation was obtained at 80°C for 20 min. Each section was incubated with each specific digoxigenin-dUTP–labeled probe at 1.5 µg/mL concentration, and underwent hybridization at 42°C for 9 h. After being washed 3 times with buffer for 8 min, the detection substrate (UltraView red ISH DIG detection kit; Roche) was applied, and eventually the sections were counterstained with hematoxylin. A positive signal was detected as a magenta deposit within the nucleus of epithelial cells. Negative control sections, consisting of non-PV lesions and normal skin, were included in each run.
RNA-ISH
Detection of CPV1 and CPV2 mRNA was performed with the RNA-ISH 2.5 detection kit using 2 specific probes (RNAscope; ACD); the RNAscope probe V-COPV-E6E7 is a 13ZZ probe targeting the region corresponding to E6 and E7 genes found between nt 102 and 826 of the CPV1 complete genome sequence (GenBank D55633.1); the RNAscope probe V-CPV2-E6E7 is a 15ZZ probe targeting the region corresponding to E6 and E7 genes found between nt 105 and 801 of the CPV2 complete genome sequence (GenBank AY722648.1). RNA-ISH was performed manually on 4-µm thick FFPE serial sections, according to the manufacturer’s protocol. Briefly, sections were deparaffinized and pretreated with heat and protease before hybridization. Successful hybridization requires binding of adjacent probe pairs on the targeted nucleic acid, initiating a cascade that leads to deposition of the fast red chromogen. The final deposit is detected as a red, punctate precipitate. From each sample, 4 adjacent sections were stained using probes for CPV1, CPV2, Fc-PPIB (peptidylprolyl isomerase B [cyclophilin B]), and negative control probe dapB (Bacillus subtilis dihydrodipicolinate reductase [dapB] gene). PPIB was used as an endogenous control to assess RNA integrity; the bacterial gene dapB served as a negative control to assess background staining. To validate the hybridization with mRNA sequences, in a limited number of samples, DNase I (500 μg; MilliporeSigma), diluted 1:50 in 1× DNase buffer (10 mM Tris-Cl, 2.5 mM MgCl2, 0.5 mM CaCl2, pH 7.5) for 30 min at 37°C, was added before the protease treatment.
Results
Histopathology
Histologically, all cases were classified as inverted papilloma based on the endophytic growth of folded epidermis (Fig. 2A); further microscopic features observed frequently consisted of parakeratosis (19 of 19), giant keratohyalin granules (16 of 19), intracytoplasmic slate-gray fibrillar material (14 of 19), and koilocytes (12 of 19). Ballooning degeneration (7 of 19) and intranuclear inclusion bodies (4 of 19) were observed occasionally (Fig. 2B–D). Case 15 was considered a regressing inverted papilloma (Fig. 4A), based on the endophytic, but not completely developed, growth of folded epidermis around crateriform keratin (reminiscent of the inverted papilloma), the dense lymphoplasmacytic infiltrate obscuring the dermal-epidermal interface, and scattered apoptotic cells, indicative of an ongoing immune-mediated process. The overall highest histologic scores were 11 (case 1) and 9 (cases 2, 3, 17); cases 13 and 15 had the lowest scores, with 2 and 3 points, respectively (Table 2).
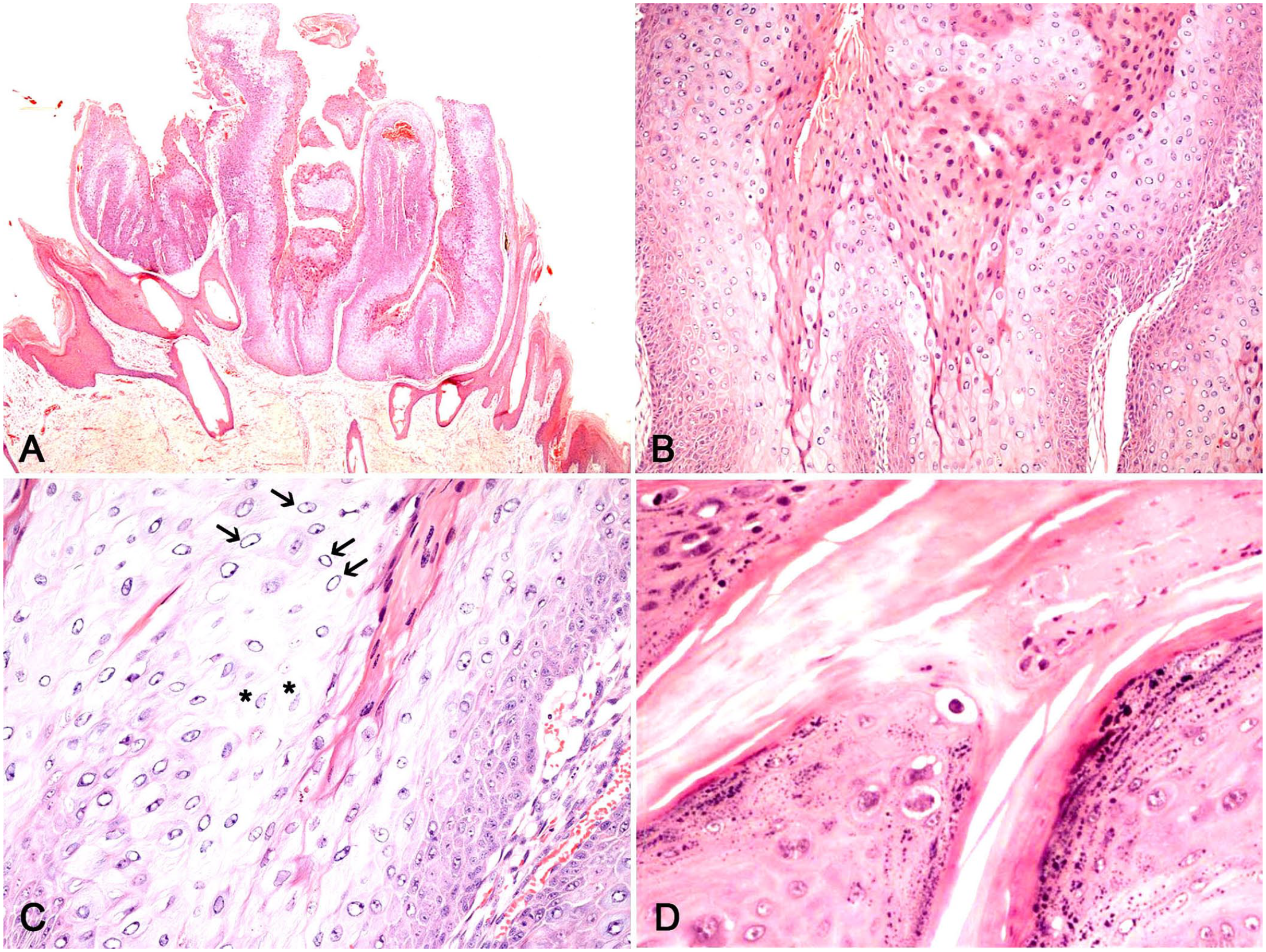
Histopathology of canine inverted papilloma.
Results of histology, IHC, PCR, DNA-CISH, and RNA-ISH investigations for 19 canine inverted papillomas.
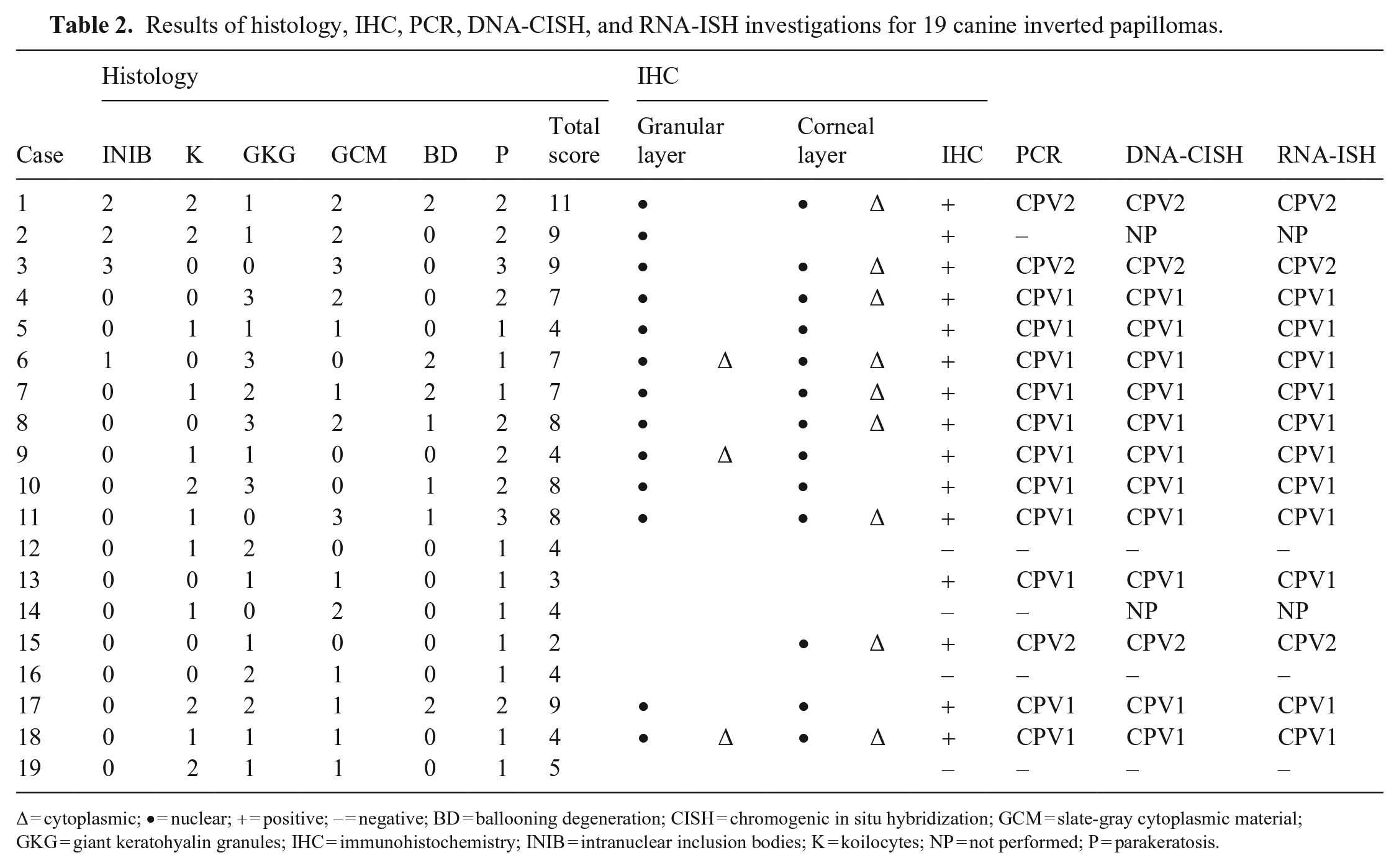
∆ = cytoplasmic; • = nuclear; + = positive; – = negative; BD = ballooning degeneration; CISH = chromogenic in situ hybridization; GCM = slate-gray cytoplasmic material; GKG = giant keratohyalin granules; IHC = immunohistochemistry; INIB = intranuclear inclusion bodies; K = koilocytes; NP = not performed; P = parakeratosis.
IHC
Overall, 15 of 19 cases had positive immunostaining, with different levels of intensity and distribution of positivity, which were detected mainly within both the corneal and subcorneal layers (14 of 15 cases); in case 15, positive immunostaining was restricted to the corneal layer (Figs. 3A, 3B, 4B). Nuclear immunostaining was reported in both corneal and subcorneal layers; cytoplasmic immunostaining was observed more frequently in the corneal layer (12 of 19) and only occasionally in the subcorneal layer (3 cases; Table 2).
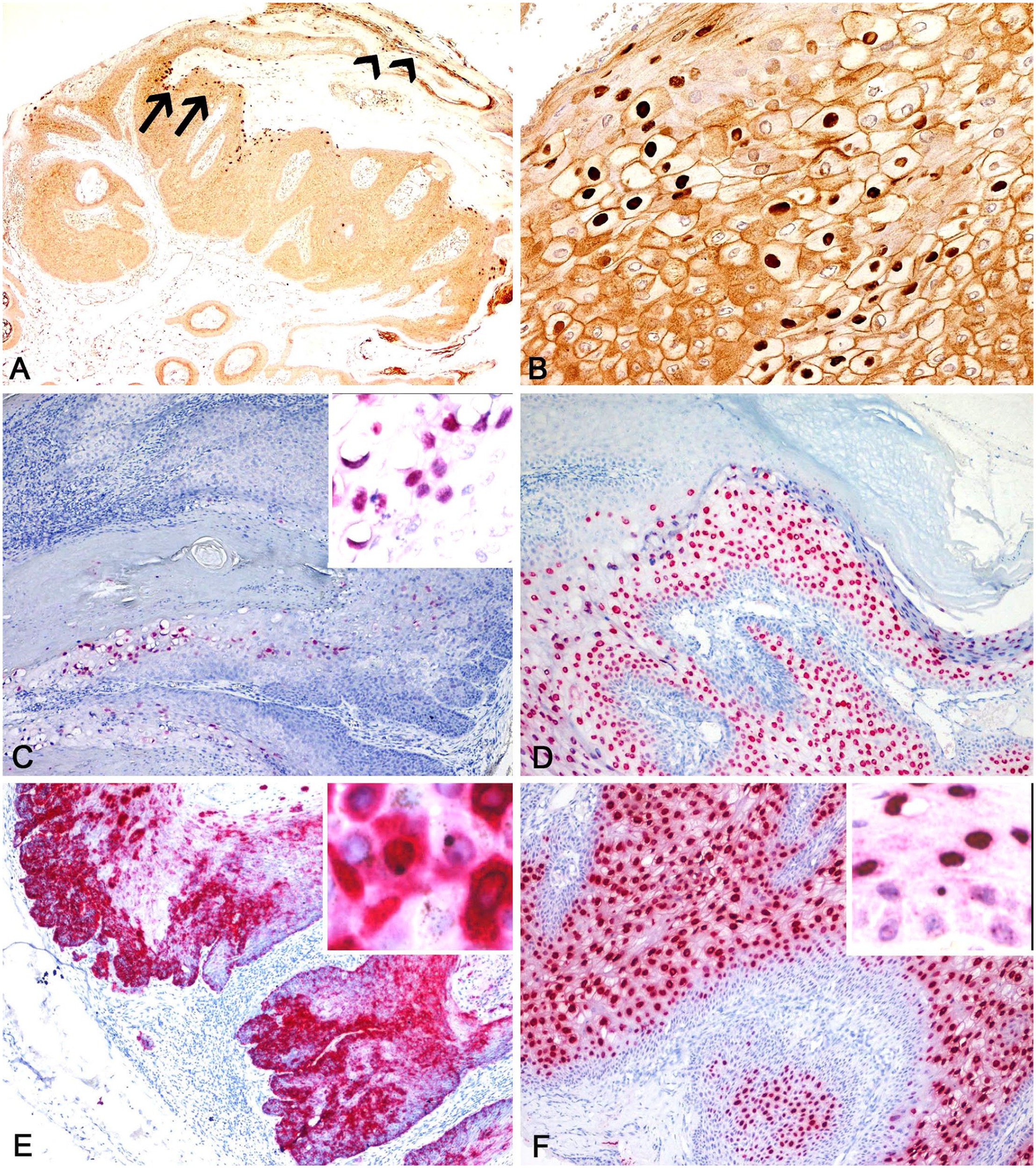
Representative photomicrographs of canine papillomavirus 1 (CPV1)- and CPV2-induced inverted papillomas. Results of immunohistochemistry (IHC), DNA chromogenic in situ hybridization (DNA-CISH), and RNA-ISH.
Molecular biology analyses
PCR
Fourteen samples had specific amplification signals for CPV1 (11 cases) and CPV2 (3 cases); CPV6 was not detected in any of the samples examined. Sequence analysis confirmed the CPV specificity identified by PCR. Five cases were negative for all CPVs tested.
All DNA samples tested yielded positive results by PCR for the Canis familiaris ribosomal protein reference gene, indicating their 212-bp DNA integrity. All 3 CPV primer sets amplified correctly the specific long oligos, either using diluted in water or in genomic canine DNA; no PCR amplicon was detected in negative controls (Suppl. Fig. 1).
DNA-CISH
Overall, 14 of 17 cases were positive by DNA-CISH, 11 for CPV1 and 3 for CPV2. Two samples (cases 2, 14) were not available for DNA-CISH investigation. Positivity was detected as a strong magenta nuclear signal, observed mainly in corneal and subcorneal epithelial cells, with a multifocal-to-diffuse pattern, and to a lesser extent in the suprabasal and basal layers, with focally extensive distribution. Positive nuclei were sometimes pushed laterally by a clear space or were completely surrounded by a clear halo, both features reminiscent of koilocytes (Fig. 3C). In case 15, the signal was dispersed within nuclei and cytoplasm of a group of exfoliating dead corneocytes (Fig. 4C). DNA-CISH was in complete agreement with IHC and PCR. Three IHC-negative samples were confirmed negative by DNA-CISH (Table 2).
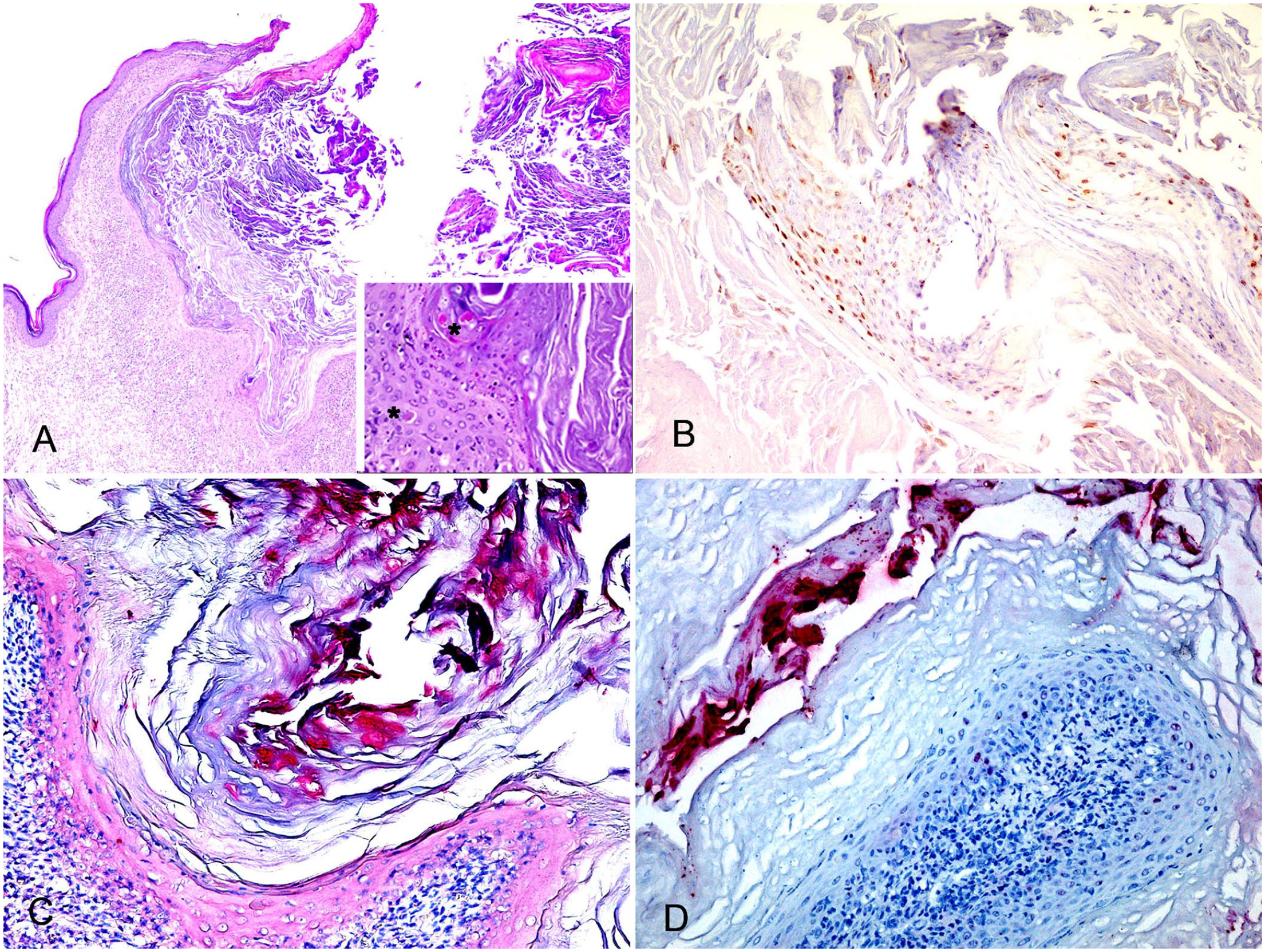
Representative photomicrographs of a canine papillomavirus 1 (CPV1)-induced regressing inverted papilloma.
RNA-ISH
Overall, 14 of 17 cases were positive by RNA-ISH, 3 were negative; cases 2 and 14 were not available for RNA-ISH investigation (Table 2). CPV1 mRNA was detected in 11 cases; CPV2 mRNA was detected in 3 cases. CPV1 positivity was detected as strong diffuse magenta cytoplasmic staining in the basal and suprabasal layers, with a medium-to-strong nuclear, multifocal, dot-like signal in the most superficial layers (Fig. 3E). Positivity for CPV2 was detected as a strong magenta nuclear signal, mainly observed in corneal and suprabasal epithelial cells, with a multifocal-to-diffuse pattern, and to a lesser extent as a punctate signal in the nuclei and cytoplasm of the basal layer (Fig. 3F). In case 15, the signal was dispersed within the nuclei and in the cytoplasm of a group of exfoliating dead keratinocytes (Fig. 4D).
No positive reaction was observed in 3 of 17 samples that also scored negative by IHC. RNA-ISH results were in complete agreement with IHC, PCR, and DNA-CISH for both of the probes tested. All of the samples incubated with the PPIB probe had good mRNA integrity and were negative with the dapB control probe. DNase I treatment gave only mild attenuation of the diffuse nuclear signal (Suppl. Fig. 2).
Discussion
The clinical histories and signalments of our 19 cases are consistent with previously reported cases.2,4,10,14–17,22,23,27,28,30 IPs are frequently reported in young adult dogs, unlike nonviral-induced epithelial tumors that arise in older subjects with no breed or sex predilection. In our survey, 2 American Staffordshire dogs were included, and 2 cases have been described previously in the Kerry Blue Terrier,4,27 although the reported sample is too small to speculate on a possible breed predisposition. The nodules more often occurred on the limbs, head, and trunk. In accordance with other reports, the abdomen and/or inguinal location was common, having been observed in 4 cases and with multiple nodules in 2 dogs. The clinical presentation was often typical, namely as umbilicated nodules present occasionally as grouped lesions.
The viral origin of IP can be suggested by histologic examination, although the classical viral cytopathic lesions were observed in a limited number of our cases. Inclusion bodies, the hallmark of this pathologic entity, were observed in only 4 of 19 cases, and in 1 case inclusion bodies scored 1 and therefore could be easily overlooked by a pathologist; koilocytes were scored 2 in 5 cases, and were observed in an additional 3 cases that yielded negative results by IHC and all molecular assays. Even though we cannot exclude that other CPVs were implicated in these cases, it might be assumed that koilocytes are not pathognomonic of PV infection in this disease, also considering that keratinocytes with koilocyte-like features can be observed occasionally in other lesions of non-viral origin. Giant keratohyalin granules and cytoplasmic slate-gray material were observed in 14 and 12 of 19 cases, respectively, both findings had high scores (2–3) in 50% of the cases. The aforementioned alterations were also observed in some of the cases that were negative by PCR, DNA-CISH, and RNA-ISH. Parakeratosis and ballooning degeneration were nonspecific features of PV infection. Considering the histologic score assigned to each case, a score ≤5 was less likely to be associated with PV presence given that both IHC and the molecular techniques were negative in 4 of 9 (44%) cases. On the contrary, a histologic score >5 was always obtained in cases in which the viral etiology was demonstrated by IHC, PCR, DNA-CISH, and RNA-ISH, and therefore our scoring system may be useful to suggest a viral etiology for the lesion, particularly if other more specific tests are not available.
In human medicine, several studies have provided evidence that human papillomavirus (HPV) may be a causative agent of squamous cell carcinoma in different anatomic locations, with important clinical and prognostic implications. The most frequent HPV tests used are PCR, IHC, DNA-CISH, and RNA-ISH.1,9,26,29 PCR can detect PV DNA in tissues and is useful to determine PV genotype but does not allow identification of virions and/or viral transcripts, which would prove active viral infection. 3 IHC is a cost-effective marker of viral infection that can be applied routinely on FFPE tissues. Unfortunately, IHC does not provide data on viral genotype. Thus, IHC is a suitable method for diagnostic purposes, but it is less informative for pathogenetic and research investigations. DNA-CISH has the advantage of identifying PV genotype and signal localization within tissues, but it does not confirm the active phase of infection. 29 Finally, RNA-ISH is a novel approach to detect viral RNA in paraffin-embedded tissues, thus assessing an active infection with viral DNA transcription. RNA-ISH is expensive but applicable to FFPE samples, and it seems a promising technique in both human and veterinary studies.5,8,12,13,20,21
In our evaluation of the ability of IHC, DNA-CISH, and RNA-ISH to detect CPV in paraffin-embedded IPs, 15 of 19 cases were positive for CPV, with the majority of positive cases being detected by IHC, DNA-CISH, and RNA-ISH. CPV6, which has been reported in the literature as a possible etiologic agent in IPs, was not detected by PV-specific PCR in our study, and therefore specific probes were not prepared and tested.
Both DNA-CISH and RNA-ISH are sensitive and specific methods to detect viral infection in FFPE tissues. The 2 DNA probes, targeting the L1 gene of CPV1 and CPV2, showed similar patterns of positivity between the 2 viral genotypes, revealing viral DNA in the nuclei of suprabasal cells. The RNAscope probes, that were specifically designed to detect E6 and/or E7 mRNA for CPV1 and CPV2, showed a different pattern of positivity between the 2 viral genotypes, giving different biological information compared to PCR or DNA-CISH. In fact, in human medicine, it is known that E6 and E7 transcript expression directly indicates that HPV is transcriptionally active.6,7 Particularly, the latent phase of infection is characterized by undetectable E6 and/or E7 expression; in the productive phase, low levels of E6 and/or E7 transcription are detectable in the lower third to half of the epithelium; the transformative phase has high levels of E6 and/or E7 expression throughout the full thickness of the epithelium. 8 In our survey, in CPV1-infected samples, E6 and/or E7 was detected in all of the epithelial layers, being diffusely expressed in the basal and suprabasal layers and multifocally in the most superficial layers; in CPV2-infected samples, E6 and/or E7 expression was mainly concentrated in the suprabasal layers, suggesting different phases of infection. Even though PV in dogs is mainly associated with hyperplastic lesions, PV-induced malignant transformations are reported rarely for different CPV genotypes.10,19 In human patients, infections with specific PV types are associated with progression to cancer; further studies are needed in the veterinary field to understand the viral and host factors associated with the risk of malignant transformation.
RNAscope has proved to be a sensitive and specific assay for mRNA demonstration by ISH, given a probe design strategy in which 2 independent probes (double Z probes) hybridize to the target sequence in tandem for signal amplification to occur. As described previously,8,12 the experimental conditions are such that the denaturation of the DNA to allow the probes to bind is minimal, as demonstrated by DNase I treatment prior to hybridization, which caused only mild attenuation of the diffuse nuclear signal.
Both dogs from the UK tested positive for CPV2; most of the Italian dogs were positive for CPV1, except a Beagle and a young Labrador Retriever that were infected by CPV2. CPV2 infection has not been reported previously in Italy, to our knowledge. Malignant transformation to squamous cell carcinoma has been reported for CPV2 in a group of dogs with X chromosome–linked severe combined immunodeficiency, 10 and recently also for CPV1. 28 Although progression toward malignant neoplasia is reported only rarely, it would be of interest to further investigate the etiology of PV-induced lesions to better understand their biologic behavior.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387211035799 – Supplemental material for Localization and genotyping of canine papillomavirus in canine inverted papillomas
Supplemental material, sj-pdf-1-vdi-10.1177_10406387211035799 for Localization and genotyping of canine papillomavirus in canine inverted papillomas by Margherita Orlandi, Maurizio Mazzei, Marta Vascellari, Erica Melchiotti, Claudia Zanardello, Ranieri Verin, Francesco Albanese, Fabiano Necci, Luca Pazzini, Giulia Lazzarini and Francesca Abramo in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank all of the veterinary clinicians who referred their cases, providing signalment and clinical information.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was supported financially by “Fondi di Ateneo” University of Pisa.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
