Abstract
MicroRNAs (miRNAs) are small non-coding RNAs that regulate gene expression at a post-transcriptional level by silencing targeted messenger RNA (mRNA). Most studies concerning miRNA expression use solid tissue samples. However, circulating miRNAs from different body fluids have recently emerged as diagnostic and prognostic molecules, given that they hold informative value and have increased stability in cell-free form. Blood sampling of cats can be challenging given their small body size and because they often experience distress when handled. We quantified miR-20a, -192, -365, -15b-5p, and -16-5p from plasma and serum samples of 10 healthy domestic cats. Our RT-rtPCR procedure used 100 µL of either plasma or serum samples as sources of biomarker molecules. However, serum provided higher amounts of miRNA than plasma samples, with a p < 0.0001 for miR-20a and p < 0.0002 for miR-16-5p.
MicroRNAs (miRNAs) are small non-coding RNAs that regulate gene expression at a post-transcriptional level by suppressing blocks of translation of targeted messenger RNAs (mRNAs). 1 The concentration of miRNAs can vary significantly among the different sources in the organism. Circulating miRNAs from fluids such as serum and plasma are much more stable than those found in solid tissue cells, facilitating their manipulation and storage in laboratory settings. 12 Several miRNA-related techniques have been reported; peripheral blood can be a valuable source of miRNA.2,12
To assess the utility of serum and plasma samples, we must consider many factors that affect the concentration of circulating miRNAs. Critical factors to be carefully monitored are preanalytical variables (sampling and processing), purification efficiency, and technical issues regarding the reverse-transcription real-time PCR (RT-rtPCR) procedure and data analysis. 12 Indeed, the amount of circulating miRNA may vary between plasma and serum samples obtained from the same individual’s blood. 17 Sample preparation and miRNA measurement methods are necessary for the development of a test that could use circulating miRNA as a biomarker.4,12
The domestic cat is a companion animal that is often considered a “family member.” In scientific research, cats represent relevant models for ~250 genetic diseases. 13 Although renal disease is common in domestic cats,13,19 we only found 2 studies that had identified miRNAs involved in renal changes. Analysis of renal tissues from healthy cats identified miRNAs in both the renal cortex and medulla. 9 Urinary miRNAs were also detected in the urine of cats. 10 Although the miRNAome of many species has been well characterized, feline miRNA is not recorded in miRNA databases, which impedes research on spontaneous diseases in cats with both feline and human applicability.13,15 Previous studies have evaluated only serum and tissue miRNAs in samples from diabetic cats, and also in cats with liver and heart diseases.3,6,18 We found no reports using feline plasma samples for miRNA detection.
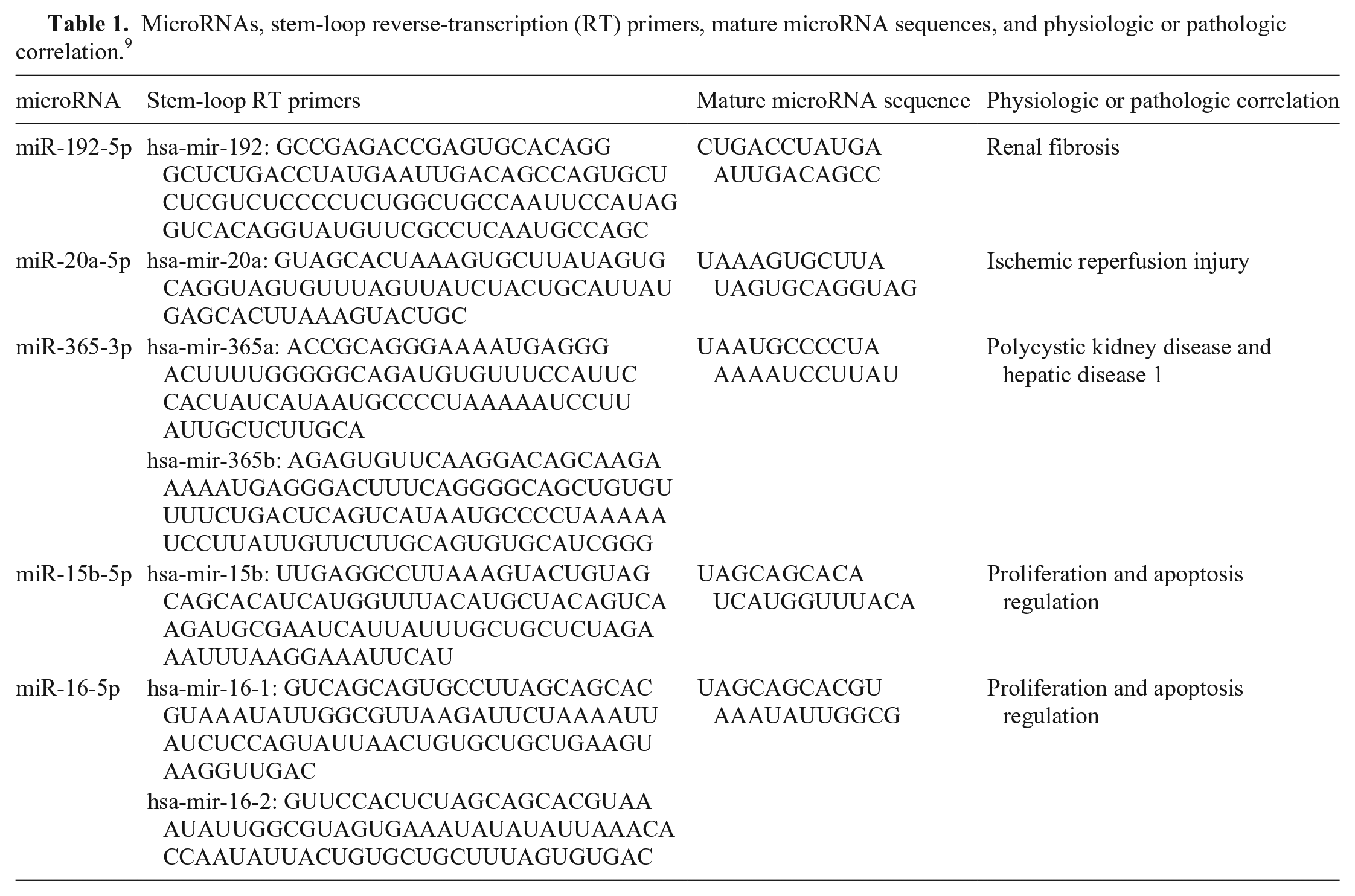
We developed a method for quantifying circulating miRNAs from feline blood samples. We compared and verified plasma and serum samples as a reliable source of miRNAs, examining 5 different targets (miR-20a, -192, -365, -15b-5p, -16-5p) in blood samples from healthy domestic cats. We chose these targets from a set of miRNAs showing differential patterns of expression during physiologic and pathologic conditions of kidney tissue in dogs and cats (Table 1). 9 In mice, the progression of renal fibrosis has been correlated with an increased expression of miR-192. 11 miR-20a was overexpressed after reperfusion in a mouse model of ischemic injury. 7 In humans, miR-365 was related to gene 1 transcription in polycystic kidney disease. 5 miR-15b-5p and miR-16-5p are regulators of cell proliferation and apoptosis in rats. 20
MicroRNAs, stem-loop reverse-transcription (RT) primers, mature microRNA sequences, and physiologic or pathologic correlation. 9
The Animal Use Ethics Committee, CEUA-UnB (protocol 82/2018) approved our experimental methods. The 10 selected animals were client-owned, clinically healthy domestic cats, 6 males and 4 females, 1 Persian and 9 mixed-breed, with ages of 7 mo to 7 y. All cats were patients from the Veterinary Hospital at the University of Brasília (Hvet-UnB). Samples were collected from cats by venipuncture of cephalic or jugular veins into K3EDTA-containing tubes for plasma, and non-additive tubes for serum. Samples were processed at the Veterinary Clinical Pathology and Technology for Gene Therapy Laboratories, at the College of Agronomy and Veterinary Medicine, University of Brasilia, Brazil.
After centrifugation of blood at 1,814 × g for 5 min, the serum was carefully aspirated to avoid contamination with cellular debris. Plasma in K3EDTA-containing tubes were separated from blood within 1 h of collection, at 1,300 × g for 10 min in a 4°C refrigerated centrifuge for removal of formed elements. A second centrifugation was performed at 3,000 × g for 10 min at 4°C to remove any residual platelets. 16
Samples were frozen at −80°C until the extraction procedure. The samples were separated into 100-µL aliquots to minimize the effects of repeated cycles of thawing and freezing, given that these temperature changes reduce the number of miRNA molecules available. 12
Extraction was performed (miRNeasy mini kit; Qiagen), using the manufacturer’s recommendations, with addition of a supplementary protocol. 18 Initially, we set up an extraction method testing volumes of 400 µL, 100 µL, and 50 µL of plasma and serum obtained from 2 clinically healthy domestic cats. The miRNA measurement was undetectable on 400 µL and 50 µL of plasma, and 50 µL of serum. These results were probably caused by a low performance of the extraction kit when using these volumes and type of samples. Using 100 µL of plasma or serum, total miRNA concentrations were 0.25–0.96 ng/µL. From then on, extraction was carried out only with 100 µL of plasma or serum. As an extraction and normalizing control, 3.5 µL of the synthetic cel-miR-39-3p (miRNeasy serum/plasma spiked-in control; Qiagen) was spiked into each sample at a concentration of 1.6 × 108 copies/µL. 14 Final elution volumes in RNase-free water were 30–50 µL.
Fluorometric quantification of miRNA was done using 2 µL of extracted sample (Qubit 2.0, microRNA assay kits; Life Technologies) following the manufacturer’s recommendations. The miRNA concentrations were 0.25–1.0 ng/µL. All samples were diluted to 0.25 ng/µL to standardize the miRNA concentration for the following RT-rtPCR protocol.
The TaqMan RT kit (Thermo Fisher) was used for complementary DNA synthesis, followed by real-time PCR (rtPCR). Stem-loop RT primers were used for the quantification of the miRNAs (Table 1). Preparation of the reagent mix followed the manufacturer’s instructions (TaqMan; Life Technologies). The rtPCR assays were performed using a QuantStudio 12K (Life Technologies) thermocycler. All samples were tested in triplicate, and results were expressed as cycle threshold (Ct; Table 2). All assays included RNase-free water as a negative control. 8
Cycle threshold values (mean ± SEM) for microRNA derived from feline plasma and serum samples.
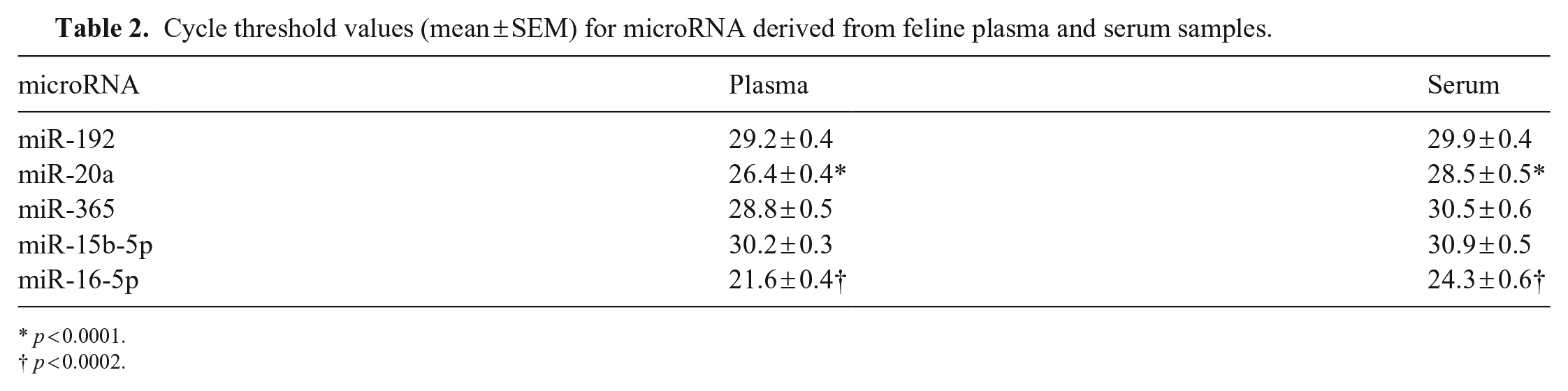
p < 0.0001.
p < 0.0002.
The rtPCR results were evaluated by the mean Ct value from each sample. 8 Data for each miRNA was normalized by the Ct value of the target reference spike-in miRNA control cel-miR-39-3p to obtain the ΔCt. D’Agostino–Pearson was used to analyze normal distribution. An unpaired student t test was used for comparisons between groups, and data were expressed as the mean and standard error of the mean (Fig. 1; Table 2). The level of significance was p ≤ 0.05. All analyses were performed using Prism 6 (GraphPad).
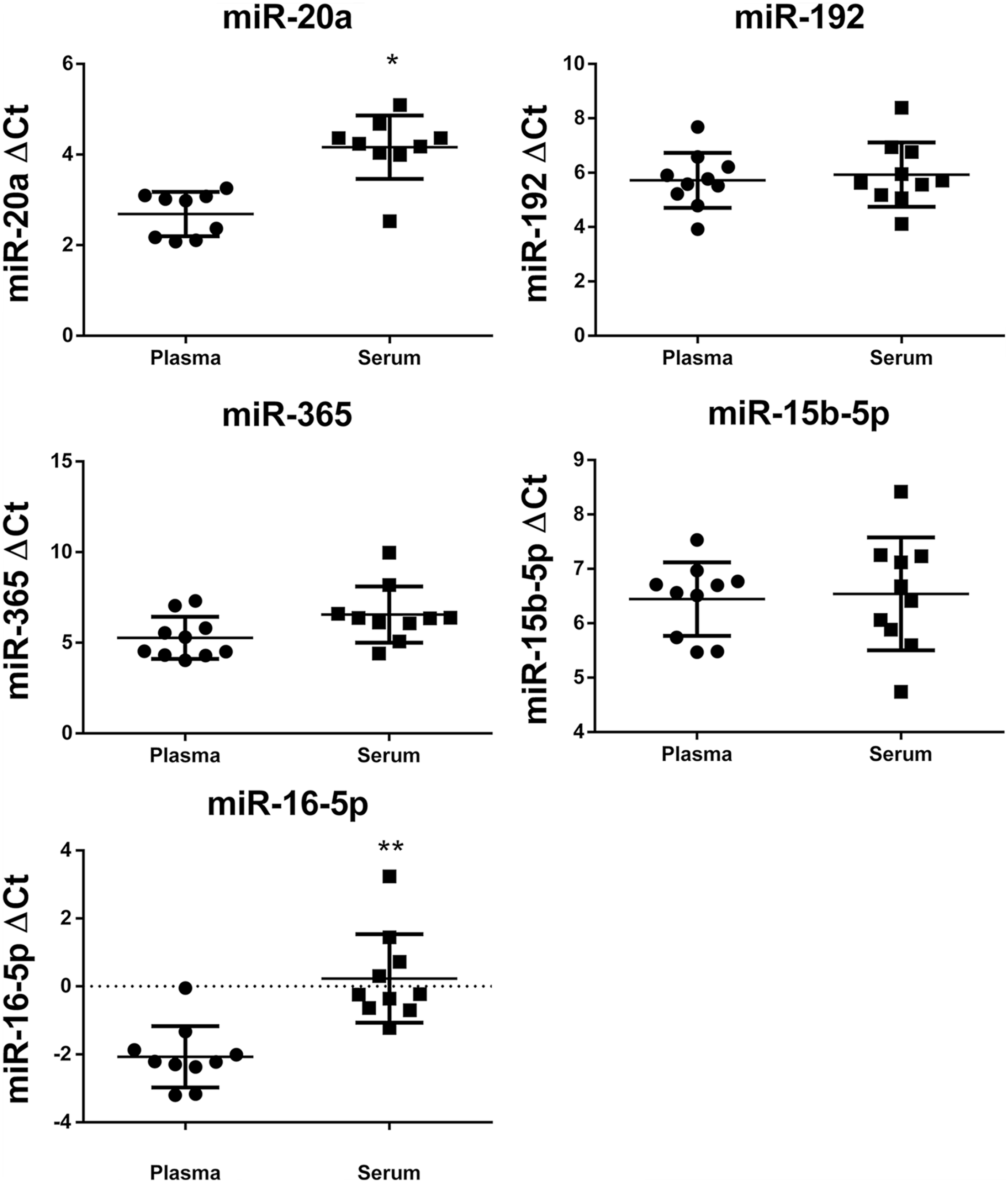
ΔCt of microRNAs (miR-20a, miR-192, miR-365, miR-16-5p, miR-15b-5p) from domestic cat plasma and serum reverse-transcription real-time PCR assays, normalized with cel-miR-39-3p, and p value by unpaired student t test. * p < 0.0001; ** p < 0.0002.
All miRNA targets were successfully detected in serum and plasma samples (Table 2). It was possible to identify different patterns of expression among the various miRNAs. Based on our results, blood samples were viable sources for miRNA measurement in domestic cats (Fig. 1).
More miRNA was found in serum than in plasma samples for both miR-20a (p < 0.0001) and miR-16-5p (p < 0.0002; Fig. 1). In general, plasma samples are preferred for miRNA analysis, mainly to avoid variation caused by the coagulation process and release of small RNAs from blood cells. 17 Our finding of more miR-20a and miR-16-5p in serum than in plasma can be explained by the release of small RNAs from blood cells ruptured during coagulation.
In the studies of diabetic cats and cats with hypertrophic cardiomyopathy, miRNA in serum was compared with healthy cats,6,18 but miRNA was not quantified in plasma samples. Simultaneous identification of miRNA in plasma and serum from the same cats has not been reported previously, to our knowledge. Considering miR-192, miR-365, and miR-15b-5p, plasma and serum concentrations values were not statistically different (Fig. 1).
Several preanalytical factors must be considered in sample handling and preparation because they may interfere with the availability of miRNAs. A critical step for miRNA measurement from plasma samples is the initial separation procedure, which requires careful handling. This step must be done within one hour of sample collection and requires a refrigerated centrifuge. Therefore, it is more convenient to use serum samples, which do not require a refrigerated centrifuge. However, when choosing the blood collection tube for a serum sample, one must consider that there are at least 2 types of tubes commonly used: a dry tube in which the coagulated blood is simply centrifuged and, alternatively, a gel separator tube for rapid and complete separation of serum from the clot. It is important that the type of collection tube used in the laboratory be standardized to avoid mistakes in operational protocols. 12
A previous study demonstrated that miRNAs can be very stable in serum, even under denaturation and degradation conditions. 4 For diagnostic purposes, more information is needed, such as reference intervals for healthy animals, and standardization of spiked-in or endogenous miRNA for normalization of results.
One of the objectives of our research was to establish a minimum volume of each type of sample for work with domestic cats. Our final protocol used 100 µL of plasma or serum, enabling successful miRNA identification without the need to collect a large volume of blood. Serum samples contained more miRNA than plasma. Thus, we consider serum the ideal sample, especially for cats, which have low concentrations of miRNAs in their blood compared to human samples. 17 Studies typically consider serum as a readily accessible source for circulating biomarkers.4,12,17
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
We thank Coordenação de Aperfeiçoamento de Pessoal de Nível Superior–Brasil (CAPES-PROAP) and Decanato de Pesquisa e Inovação da Universidade de Brasilia for their financial support for our work.
