Abstract
Bovine parvovirus 1 (BPV1) is a causative agent of respiratory, gastrointestinal, and reproductive cattle diseases. We collected 149 yak diarrhea fecal samples from 9 farms in the Qinghai-Tibet Plateau. The samples were screened for BPV1 by PCR, and 2 samples were positive for BPV1. The complete genomes of these BPV1 isolates were sequenced successfully. The sequences of these 2 variants were both 5,515 bp in length and shared 96.5–96.8% identity with 2 previously reported BPV1 genomes (GenBank DQ335247, NC_001540). Twenty-six identical amino acid mutations were found in the 2 yak variants, including 7 amino acid substitutions in receptor-binding regions of the VP2 protein, and 5 amino acid substitutions in the NS1 protein C-terminal region that functions to activate transcription. The new genome sequences contribute to better understanding of the evolution and molecular characteristics of BPV1.
Bovine parvovirus 1 (BPV1; Parvoviridae, Parvovirinae, Bocaparvovirus, Ungulate bocaparvovirus 1) 2 is a single-stranded DNA virus that is a causative agent of respiratory, gastrointestinal, and reproductive diseases in cattle. 19 BPV1 has been detected in many countries, indicating a worldwide distribution. 19 The 2 previously reported BPV1 genomes (GenBank DQ335247, NC_001540) are 5,515 and 5,517 nucleotides (nt) in length, respectively, and contain 3 open reading frames (ORFs): ORF1, ORF2, and ORF3, encoding the nonstructural protein NS1, nuclear phosphoprotein NP1, and viral capsid proteins VP1/VP2, respectively. 17 Notably, NS1 protein is vital for viral DNA replication, and the C-terminal region of NS1 protein contains an activating domain for transcription.14,16 The VP2 protein is involved in receptor attachment, antigenicity, and pathogenicity. 7 The crystal structure of the VP2 protein of BPV1 reveals the important determinants of receptor binding. 8
The yak (Bos grunniens) is a unique long-haired bovine species. 9 There are over 14 million yaks worldwide, and these animals are distributed mostly in the high-altitude regions (>2,500–6,000 m) of India, China, and Nepal. The yak is an indispensable bovine species for native people living in the Qinghai-Tibet Plateau. 18 Diarrhea is a common disease that causes major losses to the yak industry; many viruses have been identified as the pathogens causing diarrhea in yak, including bovine rotavirus, bovine viral diarrhea virus, and bovine nebovirus.3,6,20 However, little is known about BPV1 and whether yak are susceptible to infection by this bovine pathogen. We investigated whether yaks were infected by BPV1 and determined the molecular characteristics of BPV1 genomes in yaks.
In June 2020, we collected 149 fecal samples on 9 farms from yak (≤2 mo old) with diarrhea in Hongyuan, Sichuan Province, China. All samples were screened for BPV1 by a validated PCR detection method. 5 Two samples from 1 farm were BPV1 positive. Although this is a low infection rate by BPV1, this detection confirms the presence of this virus in yaks, and this finding may contribute to the diagnosis and prevention of yak diarrhea.
To determine the molecular characteristics of BPV1 in yak, primers were designed to amplify the genome sequences from the 2 positive samples. The homologies of nucleotide and deduced amino acid sequences were determined using MegAlign 7.0 (DNASTAR). MEGA X (https://www.megasoftware.net/) was used to perform multiple sequence alignment and to subsequently build neighbor-joining phylogenetic trees with 1,000 bootstrap replicates. A 3-dimensional model of the Yak/HY-HC11/2020/CH capsid VP2 protein was generated on the Swiss-Model server (https://swissmodel.expasy.org/) from the crystal structure of the BPV1 capsid VP2 protein (PDB ID 4qc8.2.). Chimera (https://www.cgl.ucsf.edu/chimera) was used to visualize the structure.
Two complete genomes of yak variants were sequenced successfully and named Yak/HY-HC11/2020/CH and Yak/HY-HC12/2020/CH (GenBank MW032436, MW032437). Our 2 BPV1 genomes obtained from yak bring the number of BPV1 genomes in GenBank to 4. The similarities of genome sequences for these 4 BPV1 variants were 96.5–98.6%. A neighbor-joining phylogenetic tree was constructed based on all complete genomes of BPV1, and yak variants were located on an independent branch (Fig. 1). Further analysis of these sequences revealed identical lengths of NP, VP1, and VP2 genes. Notably, the NS1 gene of strain NC_001540 differed from that of the other stains by lacking nt 336–740 as a result of an additional A residue, as well as a G residue at nt 507 and nt 778, in the NS1 coding region. Like other parvoviruses, greater conservation was seen for the NS1 (96.4–98.9%) and NP (97.4–98.6%) genes than for the VP1 (93.8–93.9%) and VP2 (94.1–94.2%) genes. 4 The higher sequence variation of BPV1 VP1 and VP2 genes may reflect the selective pressure of the environment and host immunity. 10 Interestingly, 26 unique and identical amino acids were found in our 2 variants from yaks, with 7 amino acid substitutions in NS1, 5 in NP1, 4 in the VP1 unique region of VP1, and 10 in the overlapping region of VP1 and VP2 (Fig. 2) proteins. Given these amino acid substitutions, the genomes in our study that generated these variant proteins were of 2 new BPV1 variants, and it is necessary to do further investigation to analyze the function of the variant proteins. The unique amino acid substitutions of BPV1 in yaks may be related to the special geographic environment of the Qinghai-Tibet Plateau, including high altitude (average altitude >4,000 m), low oxygen, low temperature, and low atmospheric pressure, as well as the specific characteristics of yaks.

Neighbor-joining phylogenetic analysis based on alignments of bovine parvovirus 1 complete genomes available in GenBank. The number at nodes represent the percentage branching following 1,000 bootstrap replicates. Bars indicate the evolutionary distance in substitutions per amino acid site. The sequences from our study are marked with black triangles.
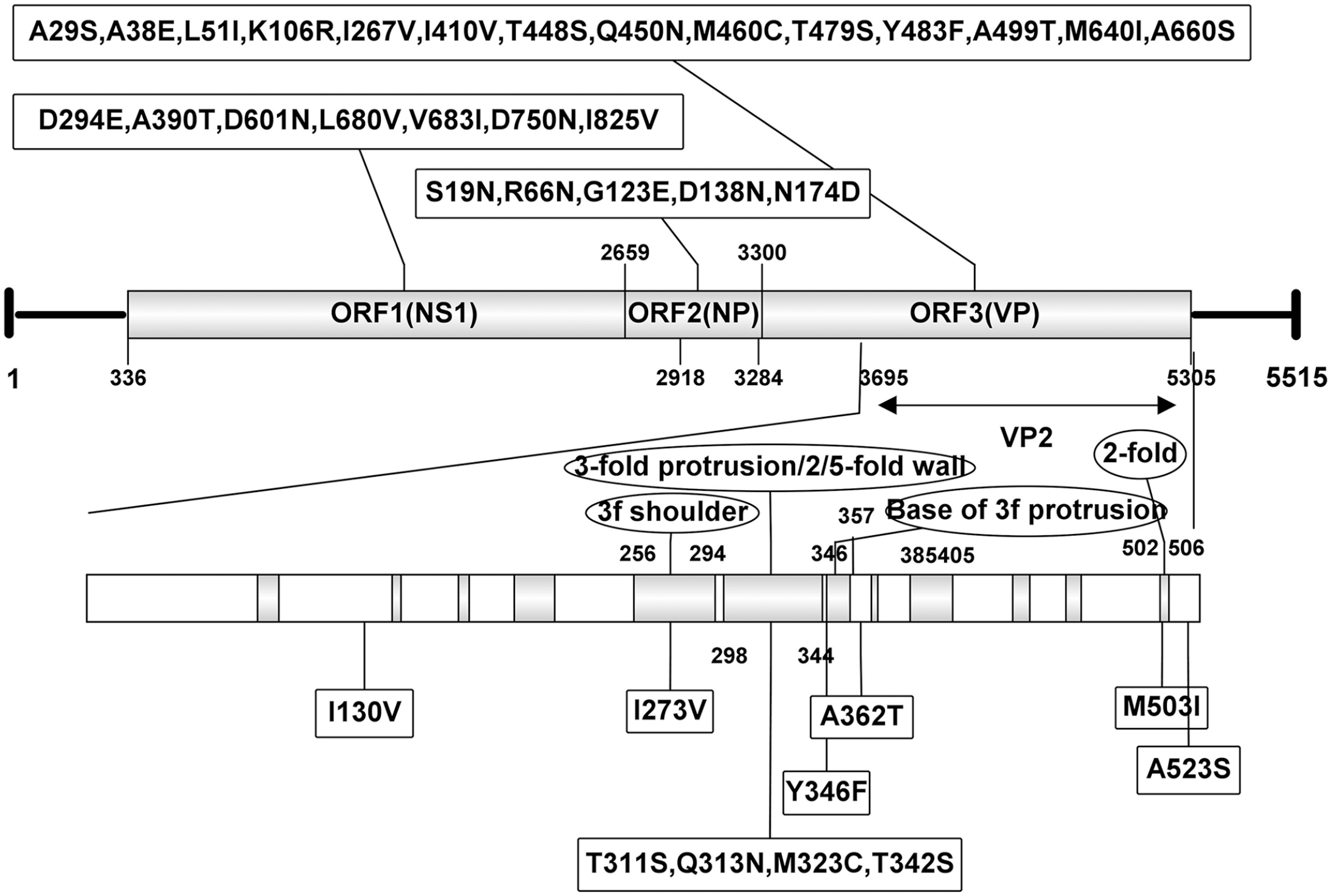
Genome organization of Yak/HY-HC11/2020/CH and unique identical amino acid variations of the 2 complete bovine parvovirus 1 sequences from yaks in our study. The numbers in the boxes indicate identical amino acid mutations in the 2 variants compared with the other 2 genomes in GenBank. The shaded boxes in the VP2 region indicate the defined variable regions.
The parvoviral NS1 protein is essential for viral DNA replication. 16 The NS1 gene contains Walker loop motifs (ASTGKT) that are conserved and required for viral infectivity of bocaparvoviruses 13 ; the C-terminal region of NS1 contains an activating domain that could be fused with a heterologous DNA-binding domain to activate transcription. 14 The 4 complete NS1 genes in GenBank contain Walker loop motifs in residues 446–451, and these sequences are 100% identical in the 4 BPV1 genomes, indicating the strong conservation of these functional elements. However, 7 unique identical amino acid substitutions in the NS1 protein of yak variants have been found, and 5 (D601N, L680V, V683I, D750N, I825V) were distributed in the C-terminal region. Given that the C-terminal region is important for virus replication, 14 further investigation is needed to understand if these mutations influence the function, especially transcription activation, of the NS1 protein.
The VP2 protein is involved in cell entry, receptor recognition, tissue tropism, antigenicity, and pathogenicity. 7 According to the crystal structure of BPV1, 11 there are 12 variable regions (VR-I to VR-IX, plus 3 additional VRs: VR-IIA, VR-IIIA, VR-VIIIB) of the BPV1 VP2 gene (HADEN strain; GenBank ABC69731.1) responsible for receptor binding, tissue tropism, and neutralization antibody recognition across the Parvovirinae subfamily. 11 Including our 2 newly acquired sequences, there are now 5 complete VP2 genes in GenBank. Compared with 3 amino acid sequences of VP2 reported previously, the 7 unique amino acid substitutions of yak variants were distributed in the VRs, with 4 substitutions (T311S, Q313N, M323C, T342S) in the VR-V motif (298–344 residues of VP2), 1 (Y346F) in the VR-VI motif (346–357 residues of VP2), and another 2 substitutions (I273V and M503I) in the VR-IV (256–294 residues of VP2) and VR-IX (502–506 residues of VP2) motifs, respectively. Although the precise functional changes of BPV1 VP2 caused by these substitutions need further studies to verify, previous work on 3 other parvoviruses (adeno-associated virus type 2 [Adeno-associated dependoparvovirus A], minute virus of mice [Rodent protoparvovirus 1], and Aleutian mink disease virus [Carnivore amdoparvovirus 1]) found that substitutions in similar regions changed the binding capacity of the neutralizing antibody and resulted in immune escape.1,12,15 The 3-dimensional model of Yak/HY-HC11/2020/CH was constructed, and results showed that, compared with the previously determined VP2 structure of strain DQ335247, there were 2 significant changes predicted by the Swiss-Model server: 1) Yak/HY-HC11/2020/CH variant VP2 protein contained a loop at residues 311–314, but in strain DQ335247 this was α-helix; and 2) the protein of the variant consisted of loops and an α-helix at residues 507–525, whereas in the strain DQ335247 the protein was in β-sheets and loops. Future work should investigate the biological significance of these unique amino acid substitutions in VP2 proteins of BPV1 from yak for a better understanding of BPV1 molecular evolution.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was funded by the Innovation Team for Emerging Animal Diseases, Southwest Minzu University (grant 2020NTD02), and Program Sichuan Veterinary Medicine and Drug Innovation Group of China Agricultural Research System (SCCXTD-2020-18).
