Abstract
Porcine reproductive and respiratory syndrome virus (PRRSV) is a highly contagious respiratory virus causing severe morbidity in pigs worldwide. Control strategies for PRRSV often rely on detecting PRRSV, culling or isolating sick pigs, disinfecting pig barns, vaccination, and monitoring for virus spread. Given the high economic impact of PRRSV on pig farms, there is a great need for rapid and reliable PRRSV detection assays. We compared the performance of 2 commercial reverse-transcription real-time PCR (RT-rtPCR) assays, the VetMAX PRRSV NA and EU reagents (ABI assay) and the PRRSV general RT-rtPCR kit (Anheal assay), for the molecular detection of PRRSV in sera collected from pigs in China. Between June and September 2015, sera were collected from 219 healthy and 104 suspected PRRSV-infected pigs on 4 farms in China. Employing blinding, the 2 assays were run by 2 laboratories (Guangzhou Animal Health Inspection Institute [GAHII] and Sun Yat-sen University [SYSU] laboratories) and compared. Although both assays detected PRRSV with 100% specificity at both laboratories, the sensitivity (95% vs. 78% at GAHII; 94% vs. 72% at SYSU Laboratory) and the reproducibility (kappa value 0.933 vs. 0.931) were slightly better for the ABI assay compared to the Anheal assay.
Introduction
The worldwide demand for pork has dramatically increased in the past 2 decades, with China emerging as the world’s largest pork consumer. As of 2016, China was home to ~46% of the world’s pigs (~456 M; http://www.fao.org/faostat/en/#data/QA, using filters: China, World + (Total); Stocks; Pigs; 2016). As the demand for pork has increased, the trend in pork production has moved from many small farms to large industrial farming in China. Although industrial farming techniques have been embraced throughout China, increasing the productivity of swine farming, not all farms have embraced high levels of biosecurity. Large farms with dense populations of pigs have much opportunity for sustained respiratory virus transmission within herds. In particular, influenza A viruses, porcine reproductive and respiratory syndrome virus (PRRSV), and porcine circoviruses are often found in farms in China. 10
Porcine reproductive and respiratory syndrome (PRRS) is one of the most economically important and devastating viral diseases impacting pig production worldwide. Financial losses are chiefly the result of increased mortality and morbidity (e.g., poor reproductive performance) and increased use of vaccines, medications, and detection assays. Secondary respiratory complications following a PRRS outbreak on a farm can lead to additional costs. Comprehensive PRRS control strategies must include detection assays and herd monitoring after detection of PRRSV in a herd.
The high economic impact of PRRSV necessitates rapid and accurate detection so that herd management practices can achieve their highest impact. 1 The rapid evolution, genetic variability, 3 and high mutation rate of PRRSV strains confound the development of long-term, reliable detection assays, and mutations in PCR target areas may quickly render a detection assay obsolete. In 2006, a highly pathogenic subtype of PRRSV emerged in China and likely spread or co-emerged from a common ancestor in Vietnam where this strain remains highly prevalent.2,5 Given the diversity of viral strains and subtypes circulating in different countries, it is important to establish the “fit-for-purpose” of a proposed test in each location before it is used routinely.
Previous studies have compared the performance of 3 commercial reverse-transcription real-time PCR (RT-rtPCR) assays for PRRSV using boars experimentally infected with 6 different PRRSV isolates, none of which represented contemporary strains circulating in China. 6 A 2018 study describes the prevalence of PRRSV in China as consisting of PRRSV-2 lineages 1, 3, 5, and 8 as the primary genotypes, with sporadic epidemics caused by PRRSV-1. 7 The molecular epidemiology of PRRSV strains in China is diverse and complex, and comparison studies on the performance of commercial assays in this country have not been published, to our knowledge. We compared the performance of a relatively new product (Applied Biosystems VetMAX PRRSV NA and EU reagents, Thermo Fisher Scientific, Waltham, MA; hereafter ABI) with a commonly used molecular assay method (PRRSV general RT-rtPCR kit, Beijing Anheal Laboratories, Beijing, China; hereafter Anheal) in China.
Materials and methods
Ethical approval
All animal contact in this comparison of molecular assays was conducted under the Chinese governmental institutional animal ethical approvals for the Chinese institutions that partnered in this work.
Case definitions
A suspect case of PRRSV infection was loosely defined as a pig having clinical reproductive or respiratory illness signs consistent with PRRSV infection. Reproductive indicators of PRRSV infection often include poor conception rates, late-term abortions, increases in stillborn piglets, mummified fetuses, or weak or non-viable piglets. Respiratory signs of PRRSV infection often include fever, depression, lethargy, stunted growth, dyspnea, and pneumonia. PRRSV has often been associated with cyanosis of the ears (blue ear disease), abdomen, and vulva.
A confirmed case of PRRSV infection was defined as a pig that was suspected of PRRSV infection but also had confirmatory virus isolation or molecular detection of PRRSV by at least 1 of the 2 RT-rtPCR methods. If PRRSV test results in both laboratories were negative by both tests, then the animal status was classified as a confirmed PRRSV-negative case.
A healthy pig was defined as a pig without any signs of PRRSV infection (see as described in the suspect case). The selection of healthy pigs was made with a goal to compare sera assay results from healthy pigs with those from confirmed PRRSV-infected pigs.
Study sites
Between June and September 2015, serum samples were collected by veterinary staff from suspected PRRSV-infected pigs and healthy pigs at 4 swine farms located in Guangdong, Fujian, and He’nan Provinces in China (Fig. 1).
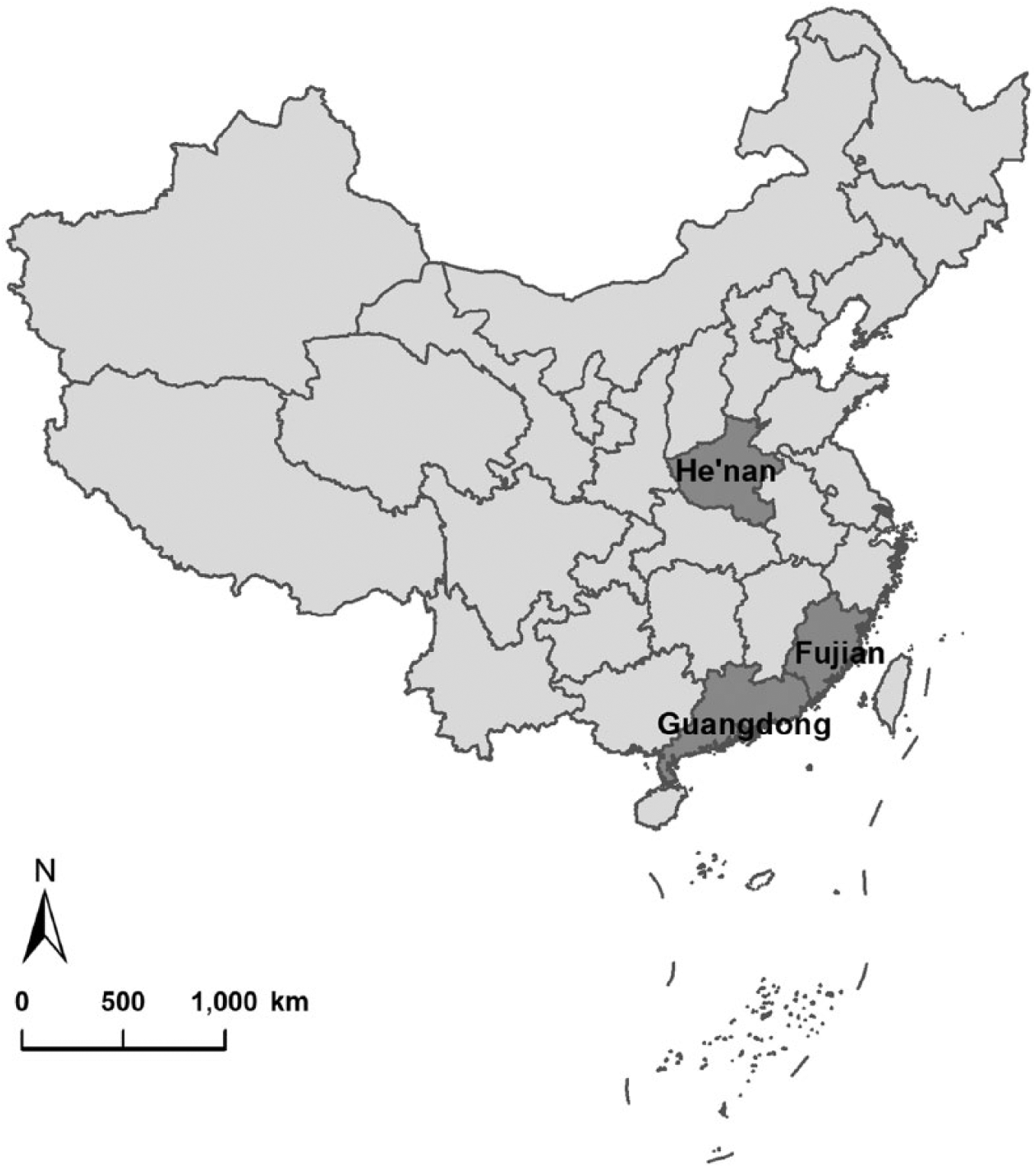
Specimens were collected from 4 farms in Guangdong, Fujian, and He’nan Provinces in China between June and September 2015.
Sample collection and allocation
Blood (8 mL) was collected from each pig in a serum separator collection tube (Vacutainer glass serum tubes with uncoated interior, Becton Dickinson, Franklin Lakes, NJ). Upon sera collection, demographic data were recorded for each pig including pig type, age, sex, and vaccination history. Blood samples were left at room temperature for up to 2 h until stored at 2–8°C for transport to Sun Yat-sen University (SYSU) laboratory (Guangzhou, China). Sera were then separated by centrifugation and aliquoted in a blinded fashion into 3 equal sets (by Y-T Zhang): 1 set was given to investigators at the Guangzhou Animal Health Inspection Institute (GAHII) laboratory (Guangzhou, China) for molecular study, 1 set was given to SYSU laboratorians for molecular study, and 1 set was stored at SYSU as a backup set. GAHII and SYSU laboratory staff then extracted nucleic acid and ran the 2 molecular PRRSV assays following the study procedures.
Nucleic acid extraction
At both the GAHII and SYSU laboratories, RNA was extracted and purified from serum samples (Applied Biosystems MagMAX pathogen RNA/DNA kit, Thermo Fisher Scientific) according to the manufacturer’s instructions. Internal positive control RNA (Applied Biosystems VetMAX Xeno, Thermo Fisher Scientific) was added to all samples during RNA extraction and purification to ensure recovery of RNA. After purification, RNA was tested promptly using the RT-rtPCR protocols.
RT-rtPCR
The Anheal and ABI assays differ in that the Anheal assay targets North American (NA) and European (EU) PRRSV, whereas the ABI assay simultaneously detects and distinguishes both NA and EU PRRSV in a multiplex assay. Both assays utilize hydrolysis probes. The specific gene targets of both commercial assays are proprietary and not publicly available. To test these assays, RNA was simultaneously examined using both the ABI and the Anheal assays at both the SYSU and the GAHII laboratories. Assays were performed per the manufacturer’s instructions (Applied Biosystems 7500 real-time PCR system, Thermo Fisher Scientific).
Anheal RT-rtPCR samples were run in 20-μL reactions consisting of 2.4 μL of nuclease-free water, 10 μL of RT-rtPCR reaction solution, 0.4 μL of enzyme mix, 2.2 μL of fluorescent probes, and 5 μL of extracted serum RNA run at 45°C for 15 min, 95°C for 1 min, followed by 40 cycles of 95°C for 5 s and 60°C for 35 s. The assay was considered valid if the positive control had a cycle threshold (Ct) ⩽30 and the negative control had a Ct value of zero. Samples were considered positive if the Ct ⩽30. If a sample produced a Ct value >30 and ⩽37, the RNA was re-isolated and re-tested by the Anheal RT-rtPCR.
The ABI RT-rtPCR employed 25-μL reactions consisting of 18 μL of master mix and 7 μL of extracted serum RNA run at 45°C for 10 min, 95°C for 10 min, followed by 40 cycles of amplification at 95°C for 15 s and 60°C for 70 s. The assay was considered valid if the Ct value of the RNA control was 32–35. Samples were considered positive if they had a Ct ⩽ 37. Samples were re-extracted and re-tested before confirmed if they had a Ct value >37 and ⩽40.
Data analysis
Tests results were classified into categories of true-positive, true-negative, false-positive, and false-negative compared to their final classifications of confirmed PRRSV-positive or -negative cases. For example, if a confirmed PRRSV-positive sample tested negative in the SYSU laboratory, then this result was regarded as false-negative.
Statistical analyses were performed (SPSS v.21.0, SPSS, Chicago, IL). Comparison of demographic characteristics between confirmed PRRSV-infected pigs and PRRSV-negative pigs (healthy pigs and suspected PRRSV-infected pigs with negative RT-rtPCR results) were carried out using chi-square or Fisher exact tests. Variables with a p ⩽ 0.05 were considered statistically significant. Sensitivity, specificity, and positive predictive and negative predictive values for each RT-rtPCR assay were separately measured for both the GAHII and SYSU laboratories, comparing each individual assay against a gold standard of either assay being positive. Concordance between the 2 assays in each laboratory was estimated using a kappa statistic. The reproducibility of each assay (only one assay result per specimen per laboratory was compared) between the 2 laboratories was also assessed using a kappa statistic.
Results
Study samples
Between June and September 2015, 323 serum samples were collected from a convenience sample of 4 pig farms in China where PRRSV was said to be causing illness. Seventy-two specimens were collected from farm 1 and 99 from farm 2, in Guangdong Province; 76 specimens were collected from Fujian Province; and 76 samples were collected from He’nan Province (Fig. 2). Ninety-one (88%) of 104 suspected PRRSV-infected pigs and 9 (4%) of 219 healthy pigs were molecularly confirmed as PRRSV-infected by at least 1 RT-rtPCR assay in at least 1 of the 2 laboratories.
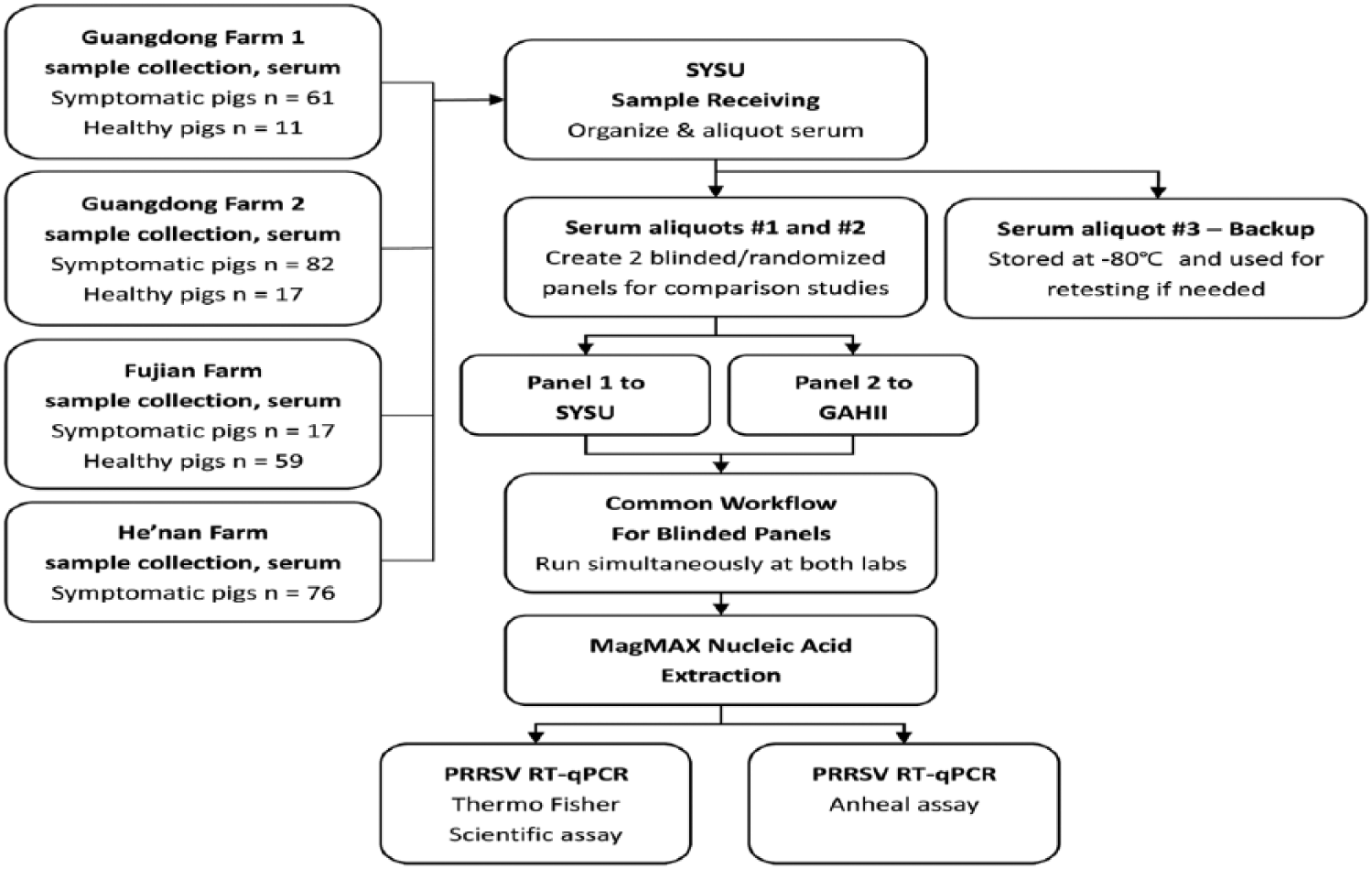
Sera were collected from suspected porcine reproductive and respiratory syndrome virus (PRRSV)-infected and healthy pigs. Two panels of original sera were created for Sun Yat-sen University (SYSU) and Guangzhou Animal Health Inspection Institute (GAHII) laboratories. SYSU and GAHII laboratorians were blinded with respect to the pigs’ signs of PRRSV infection. Nucleic acid extractions were performed separately by each laboratory. Each laboratory performed both ABI and Anheal RT-rtPCR assays.
The prevalence of several potential risk factors differed with respect to PRRSV detection (Table 1). The He’nan farm had the highest prevalence (71%) of PRRSV-confirmed pigs. Similarly, pigs ⩽2 mo in age also had the highest prevalence of PRRSV-confirmed specimens (38%) compared to pigs of other ages. Interestingly, there was no statistically significant difference in PRRSV-confirmed prevalence for influenza vaccine history or for sex.
Characteristics of confirmed porcine reproductive and respiratory syndrome virus (PRRSV)-infected pigs versus PRRSV-negative pigs.
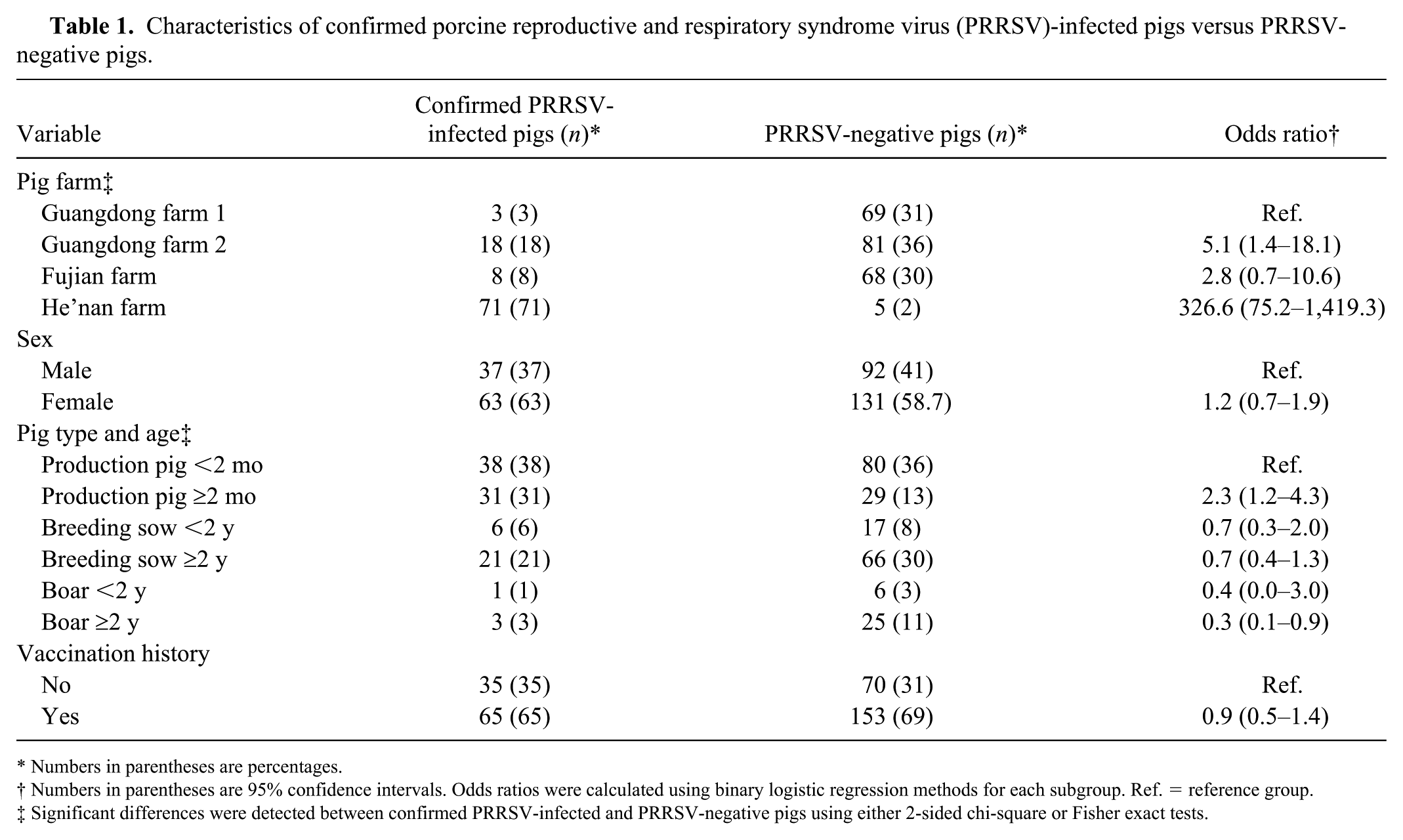
Numbers in parentheses are percentages.
Numbers in parentheses are 95% confidence intervals. Odds ratios were calculated using binary logistic regression methods for each subgroup. Ref. = reference group.
Significant differences were detected between confirmed PRRSV-infected and PRRSV-negative pigs using either 2-sided chi-square or Fisher exact tests.
Overall performance of assays
At GAHII, 95 (30%) of 323 samples tested positive using the ABI PRRSV assay. The sensitivity and specificity of this assay was 95% and 100%, respectively; the positive and negative predictive values were 100% and 98%, respectively (Table 2). Similarly, at GAHII, 78 (24%) of 323 samples tested positive using the Anheal assay. The sensitivity and specificity of the Anheal assay was 78% and 100%, respectively; the positive and negative predictive values were 100% and 91%, respectively (Table 2).
Overall performance of ABI and Anheal RT-rtPCR assays.
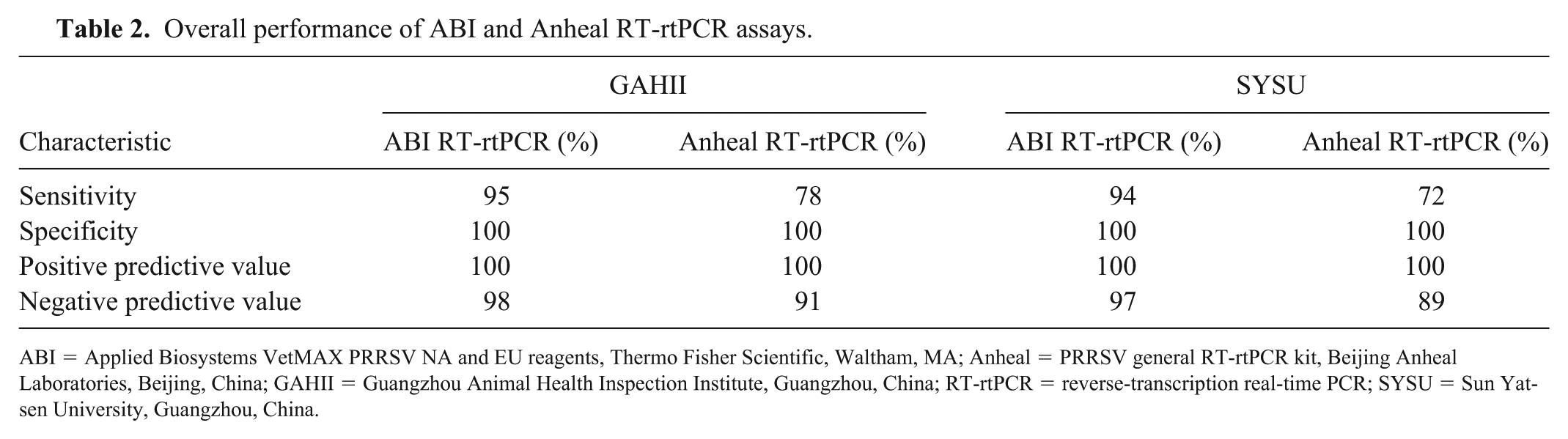
ABI = Applied Biosystems VetMAX PRRSV NA and EU reagents, Thermo Fisher Scientific, Waltham, MA; Anheal = PRRSV general RT-rtPCR kit, Beijing Anheal Laboratories, Beijing, China; GAHII = Guangzhou Animal Health Inspection Institute, Guangzhou, China; RT-rtPCR = reverse-transcription real-time PCR; SYSU = Sun Yat-sen University, Guangzhou, China.
Similar results were found from SYSU analyses, with 94 (29%) of 323 samples testing positive using the ABI assay. This assay had a sensitivity and specificity of 94% and 100%, respectively; the positive and negative predictive values were 100% and 97%, respectively (Table 3). At SYSU, 72 (22%) of 323 samples tested positive using the Anheal assay. The sensitivity and specificity were 72% and 100%, respectively; the positive and negative predictive values were 100% and 89%, respectively (Table 2).
Concordance between the ABI RT-rtPCR and Anheal RT-rtPCR assays, by laboratory.
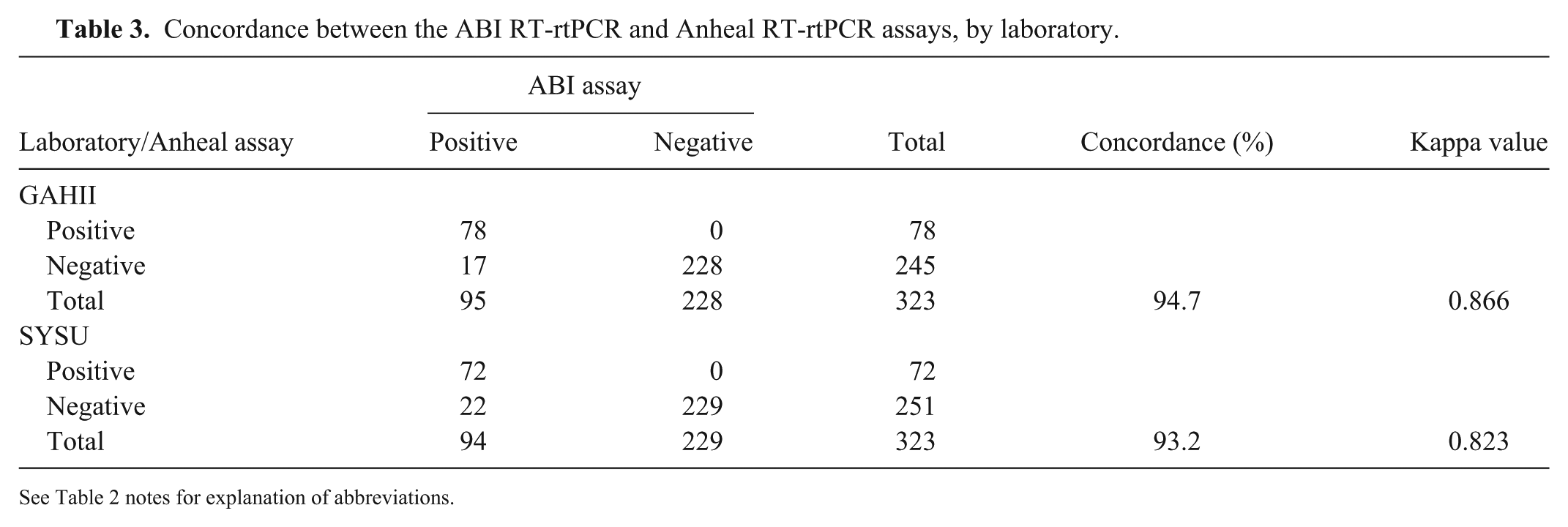
See Table 2 notes for explanation of abbreviations.
Specifically, for PRRSV subtype detection, 94 of the 100 confirmed samples were positive for NA subtype by ABI assay, whereas only 78 were positive by Anheal assay. The shift of Ct values away from the y = x isoline (Fig. 3) indicated that the ABI amplification signals came up earlier than did those of the Anheal; the Anheal assay tended to show negative results for those with higher Ct values in the ABI assay. Hence, the ABI assay was more sensitive than the Anheal assay. The shift of the data is more obvious in the GAHII laboratory than in SYSU; such a phenomenon may be explained by operation bias (from nucleic acid extraction to reaction system preparation before RT-rtPCR running) between the 2 laboratories.
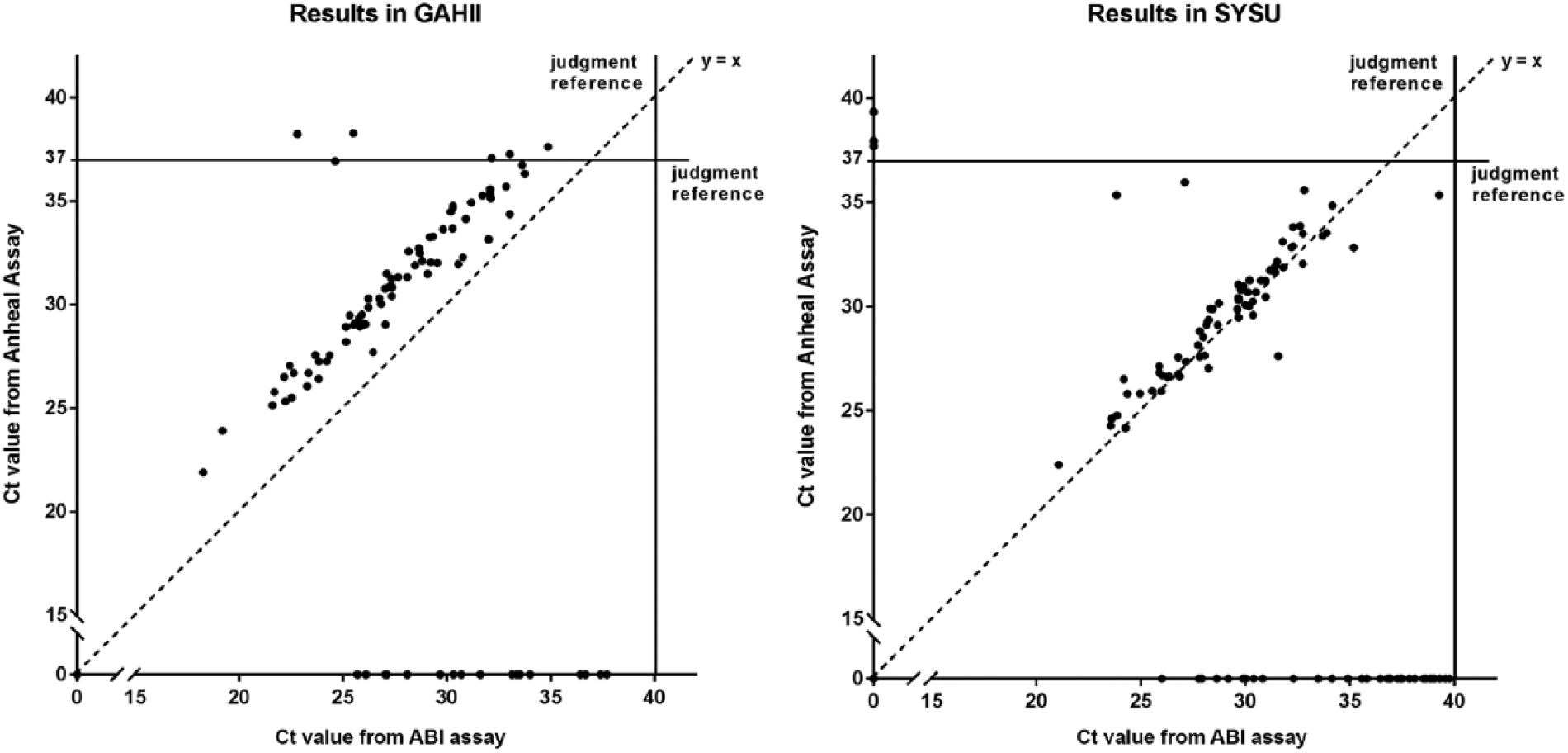
Comparison of cycle threshold (Ct) values from 2 porcine reproductive and respiratory syndrome virus (PRRSV) RT-rtPCR tests at 2 laboratories. “No Ct” indicates that no PCR amplification signal was observed by the endpoint of a test, which was 37 cycles for the Anheal assay or 40 cycles for the ABI assay. Where one of the 2 assays had “No Ct,” the results were plotted on either the x- or the y-axis.
As well, 9 samples were detected as both NA and EU PRRSV positive and 1 sample as EU PRRSV positive by ABI assay in both laboratories. The Anheal assay does not differentiate between subtypes, although it covers both subtypes. Moreover, we consider that these EU-positives may largely be the result of routine vaccination of most animals for EU PRRSV with a live-attenuated vaccine, given that the Ct values were consistently higher than expected (29.2–36.5). Therefore, we were not able to directly compare the EU detection efficacy between these 2 assays.
Concordance and reproducibility of assays
The percent concordance between the 2 commercial assays in the GAHII and SYSU laboratories were similar at 95% and 93%, respectively (Table 3). The kappa values of reproducibility for each of the assays were again similar, at 0.866 (ABI assay) and 0.823 (Anheal assay; Table 3). The overall reproducibility of the ABI RT-rtPCR assay (κ = 0.933) was slightly higher than that of the Anheal RT-rtPCR assay (κ = 0.931; Table 4).
Reproducibility of Anheal RT-rtPCR and ABI RT-rtPCR results across GAHII and SYSU laboratories.
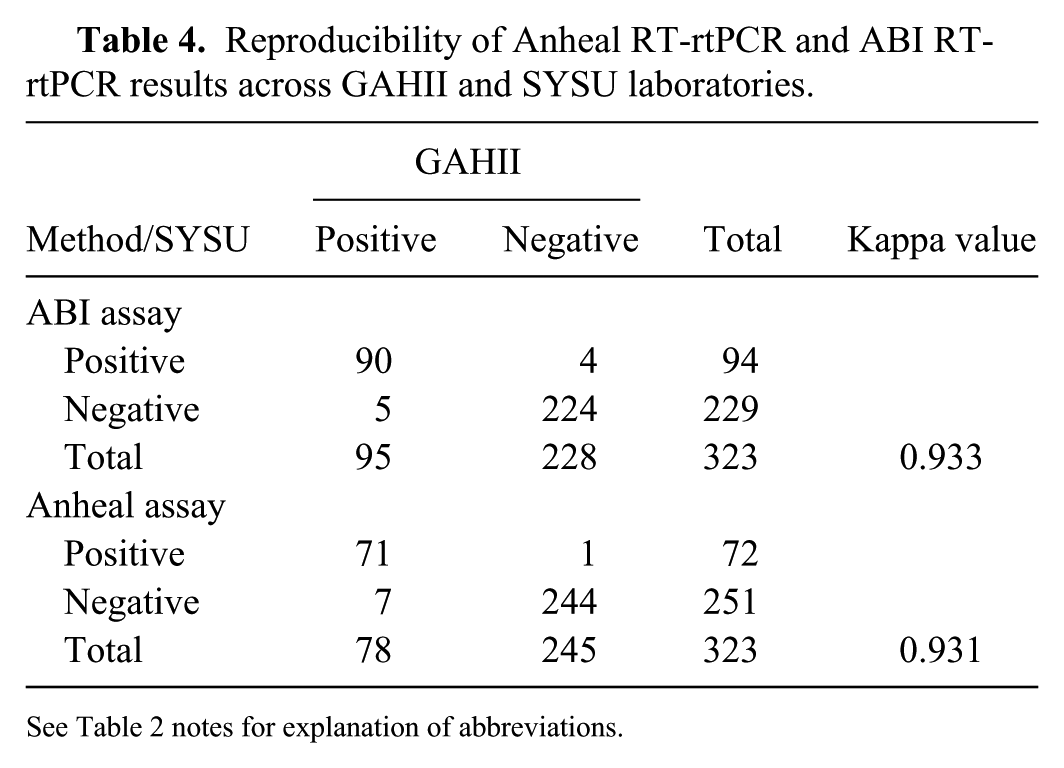
See Table 2 notes for explanation of abbreviations.
Discussion
Outbreaks of PRRS-like disease were first recognized on intensive pig farms near Beijing in 1995 and marked the first documented appearance of PRRS in China. 11 From the initial viral introduction, the disease spread rapidly to most of the provinces of China, leaving the majority of pig farms in China endemically infected. 11 In 2006, large-scale outbreaks of a virulent or highly pathogenic PRRSV (HP-PRRSV) subtype emerged in the Jiangxi Province and within a few months spread to >10 provinces, affecting over 2 million pigs. 9 Since then, HP-PRRSV has become the dominant epidemic strain circulating in swine farms, and HP-PRRS is now one of the most severe disease threats to the Chinese swine industry in terms of swine health and the economic losses resulting from production failures.
In 2018, there seems to be a gap in knowledge regarding the performance characteristics of commercial RT-rtPCR assays for the detection of PRRSV RNA in pigs in China. Lack of understanding of the basic performance characteristics of available tests confounds test result interpretations and weakens disease control efforts if the disease status of an animal is improperly categorized due to a false test result. We demonstrated that 2 molecular assays are fit-for-purpose and can detect PRRSV RNA in swine serum samples from pigs produced in the dense swine production area in south-central China.
In our study, there was generally good concordance, with kappa scores of 0.866 and 0.823 indicating excellent agreement 8 between the assays for both laboratories. Although the test specificity of the 2 tests was 100% as measured by the 2 laboratories, the sensitivity was less for the Anheal test compared to the ABI test. With sensitivity of the Anheal assay of 72–78%, the negative predictive values were 89–91%, demonstrating that the false-negative rate for the Anheal test was greater than for the ABI test. The measured sensitivity of the ABI test of 97–98% is considered excellent. The reproducibility of both tests had high kappa scores that were very close (0.933 and 0.931), indicating that the results from both tests in the 2 laboratories were highly reproducible (Table 4).
There were several limitations in our study. Test results were categorized as true-positive, true-negative, false-positive, and false-negative, with no tie-breaking reference method to referee discordant results. The PRSSV case definition also had limitations. A confirmed laboratory result required the pig to have both clinical signs of infection and a positive RT-rtPCR assay result, which could have misclassified animals with early infection or asymptomatic infection and no signs of disease. RT-rtPCR assays are capable of detecting pre-clinical PPRS;4,12 however, we did not collect serial blood specimens from pigs, therefore we were unable to detect such subclinical infections. Hence, with these shortcomings, the true sensitivity of both assays is likely to have been underestimated.
Both RT-rtPCR assays seem fit-for-purpose in detecting PRRSV, although the more sensitive ABI assay would yield fewer false-negative RT-rtPCR results compared to the Anheal assay. Controlling PRRSV spread in China, where virus strains are highly prevalent and diverse, requires the use of laboratory tests with well-understood strengths and limitations. Our data help to highlight the performance characteristics of the 2 commercial tests. Further studies utilizing the 2 commercial tests should be undertaken to evaluate the performance of the tests in other locations within China and on other sample matrices such as oral fluids and tonsil swabs in order to understand the use of these tests for the optimal detection of PRRSV in field samples.
Footnotes
Acknowledgements
Our study was supported by the participating institutions as well as by a grant to Duke University and Sun Yet-Sen University by Thermo Fisher Scientific. At the time specimens were collected and the assays were performed, JD Callahan was employed by Thermo Fisher Scientific. The authors acknowledge the much-appreciated collaborative support of Meng-Meng Ma from Guangzhou Center for Disease Control and Prevention; Chang-Xu Song and his team from South China Agricultural University; and the various veterinary staff at the study sites.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was supported by contracts to Duke University and Sun Yet-Sen University from Thermo Fisher Scientific.
