Abstract
Chronic wasting disease (CWD) is a prion-mediated, transmissible disease of cervids, including deer (Odocoileus spp.), which is characterized by spongiform encephalopathy and death of the prion-infected animals. Official surveillance in the United States using immunohistochemistry (IHC) and ELISA entails the laborious collection of lymphoid and/or brainstem tissue after death. New, highly sensitive prion detection methods, such as real-time quaking-induced conversion (RT-QuIC), have shown promise in detecting abnormal prions from both antemortem and postmortem specimens. We compared RT-QuIC with ELISA and IHC for CWD detection utilizing deer retropharyngeal lymph node (RLN) tissues in a diagnostic laboratory setting. The RLNs were collected postmortem from hunter-harvested animals. RT-QuIC showed 100% sensitivity and specificity for 50 deer RLN (35 positive by both IHC and ELISA, 15 negative) included in our study. All deer were also genotyped for PRNP polymorphism. Most deer were homozygous at codons 95, 96, 116, and 226 (QQ/GG/AA/QQ genotype, with frequency 0.86), which are the codons implicated in disease susceptibility. Heterozygosity was noticed in Pennsylvania deer, albeit at a very low frequency, for codons 95GS (0.06) and 96QH (0.08), but deer with these genotypes were still found to be CWD prion-infected.
Chronic wasting disease (CWD) is an infectious prion disease affecting cervid populations, including white-tailed deer (Odocoileus virginianus).5,13 As in other prion diseases, the course of CWD is characterized by the accumulation of abnormal prion protein (PrPSc) in lymphoid and nervous tissue, with clinical signs arising chiefly from neurodegeneration. 19 Detection of PrPSc has been reported from a variety of tissues in deer, of which the 2 tissues used for surveillance are primarily lymph node and obex.3,7–9,17
Wildlife disease surveillance in the United States entails submission of retropharyngeal lymph nodes (RLNs) from hunter-harvested deer. In captive animal disease management programs, the incorporation of new tissue types, such as recto-anal mucosa-associated lymphoid tissues, has made possible the desirable option of live-animal testing to be utilized in herd management based on deer genotypes in certain scenarios.10,23
The 2 official methods for CWD surveillance in the United States rely on the detection of proteinase K–resistant PrPSc, either by ELISA or immunohistochemistry (IHC), in neural or lymphoid tissue. Protein misfolding cyclic amplification (PMCA) 21 and real-time quaking-induced conversion (RT-QuIC)3,7–9 assays have been shown to be useful for sensitive detection of abnormal prions, including the CWD prion in a wide variety of tissues and body fluids.3,7–9,21 The methods are not yet approved for official surveillance and have only been used in a research setting. The promise of these new generation tests, potentially a very useful complement to current, harvest-dependent surveillance, warrants further investigation.
We chose to first assess the performance of RT-QuIC for lymph node tissues collected at postmortem in a diagnostic laboratory setting. The RT-QuIC assay offers some distinct advantages over the PMCA. First, it does not generate infectious prions during in vitro amplification; second, this analytical tool can be deployed comparatively easily in a diagnostic setting; and third, unlike PMCA, the result does not require additional analysis. In the RT-QuIC assay, recombinant PrP seed and thioflavin T (ThT) are added to the test sample. If the sample contains PrPSc, the seed becomes mis-folded and incorporates the fluorescent ThT into its fibrils. Multiple cycles of incubation and shaking amplify mis-folded seed, which increases the amount of incorporated ThT to a detectable level. Compared to IHC, the RT-QuIC assay is not impacted by low follicle count 12 or asymmetric prion distribution within the lymphoid tissue, and is similar to ELISA in this respect. The application of the RT-QuIC test is now being extended to antemortem samples, including feces and other tissues, where this method is likely to be of great value.3,7–9
In addition to analyzing the performance of the RT-QuIC assay, we performed a preliminary study of potential genetic linkages for CWD infections for the population under study. In addition to using a sensitive detection method for abnormal prions, a successful CWD control program might select a deer population that is less susceptible to CWD. One drawback, however, with this strategy of selecting resistant genotypes is that, in contrast to other prion-mediated neurodegenerative disorders in humans and sheep, resistant genotypes in deer thus far have been shown only to delay the onset and/or progression of the prion disease.4,18 Nevertheless, the codons at locations 95, 96, 116, and 226 within the PRNP gene have been associated with increased survival rates in CWD-infected white-tailed deer.15,16
Deer heads had been submitted for CWD surveillance during the 2019 Pennsylvania hunting season; retropharyngeal lymph nodes (RLNs) were collected and submitted for ELISA and IHC testing. Tissues were stored in plastic bags at −20°C for ELISA, or in 10% neutral-buffered formalin for IHC. We chose 50 specimens for this comparison from the 7,463 specimens tested at the laboratory during the 2019 season; 35 were from deer that were classified CWD positive by both IHC and ELISA, and 15 were from the non-detected category. The specimens were from 31 adults that were > 2.5 y, 2 fawns that were < 6 mo old, and 17 yearling deer (up to18 mo old), comprising 28 male and 22 female deer. All animals were aged based on dentition patterns.
The ELISA (TeSeE SAP Combi kit; Bio-Rad) was conducted as per the manufacturer’s protocol. Briefly, homogenized tissues (0.20 g) from the RLN cortex were tested, and any samples with optical density (OD) > 0.035 plus the mean of the negative control were considered reactors and repeated in duplicate.
IHC staining was conducted as described previously.17,23 Briefly, tissues were embedded in modified paraffin (Paraplast; Millipore Sigma), sectioned at 5 μm, deparaffinized, treated with formic acid, and subjected to antigen retrieval. 23 Slides were stained (Discovery XT immunostainer; Ventana) using PrPres mAb F99, followed with biotinylated secondary antibody. After staining, slides were rinsed, dried, and coverslipped.
The RT-QuIC assay was performed using a truncated form of the Syrian hamster recombinant PrP (SHrPrP; residues 90–231), expressed and purified as described previously at CWD Evolution Inc. (Fort Collins, CO, USA), but with minor modifications.14,24 Sample homogenate (2% w/v) in Bio-Rad ELISA dilution buffer processed as per tissue extraction protocol were kept at −80°C after ELISA testing and until RT-QuIC analysis. The tissue homogenates were diluted 1:100 in 0.1% sodium dodecyl sulfate buffer, and 2 μL of this resultant homogenate was added to 98 μL of RT-QuIC reaction buffer (320 mM NaCl, 20 mM Na2HPO4, 1 mM EDTA), 1 mM ThT, and 0.1 mg/mL SHrPrPC, and incubated in a 96-well plate. Controls included tissues both from animals detected as CWD-positive by IHC and no template controls, as well as tissues non-detected with CWD by IHC from our routine CWD surveillance program. Plates were then subjected to 250 cycles of shaking with cycles of 1 min of shaking (700 rpm, double orbital) and 1 min of rest with incubation at 42°C (Omega-RT-QuIC/prion version-fluorescence base; BMG Labtech). ThT fluorescence readings (448-nm excitation and 482-nm emission, bottom read, 20 flashes per well at 4 mm setting) were taken following each 15-min cycle, using a gain setting of 1,700. The RT-QuIC assay requires ~62.5 h to complete. Positive results were those that crossed a fluorescence threshold determined by the mean fluorescence of all sample baselines plus 5 SD. The time to positivity was defined as the time at which sample fluorescence crossed the cycle threshold (Ct); time to reach maximum fluorescence (Cmax) was also recorded. Cmax is used mainly to assess the kinetic increase for seeding density once the Ct is crossed. The analyses were performed using MARS software (BMG Labtech).
Evaluation of the RT-QuIC test first involved assessing the seeding ability of RLN homogenates using recombinant PrP as a substrate. Dilutions of RLN homogenates up to 10−7 displayed positive amplification for duplicate specimens, except 1:10 as a result of sample inhibition (Fig. 1A). Samples that showed detection of prions by incorporation of ThT indicative of prion conversion were positive after 12.5 to 26.3 h at 1:100 dilution. The Ct and Cmax times were quite similar at the tested dilutions for all RLNs, validating strong seeding capacity in each of the samples tested, compared to ELISA and IHC that often showed only mild reactivity (Table 1). No spontaneous conversion was detected in any of the 15 animals from the non-detected category.
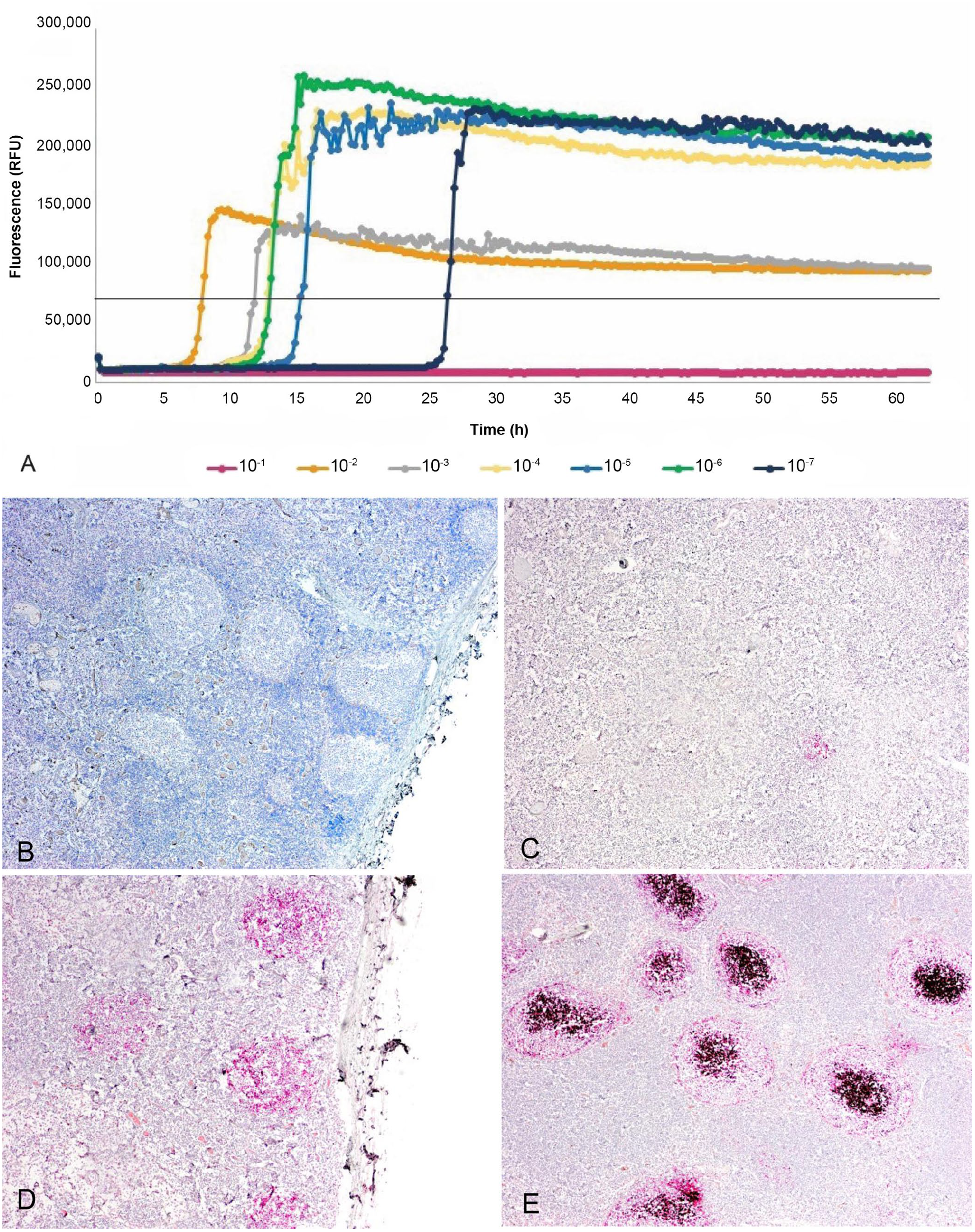
Chronic wasting disease detection in retropharyngeal lymph nodes (RLNs) from white-tailed deer by real-time quaking-induced conversion (RT-QuIC) testing and immunohistochemistry (IHC).
Comparison of real-time quaking-induced conversion (RT-QuIC), immunohistochemistry (IHC), and ELISA for chronic wasting disease PrPSc detection in lymph node tissues from Pennsylvania deer with specific PRNP genotype.
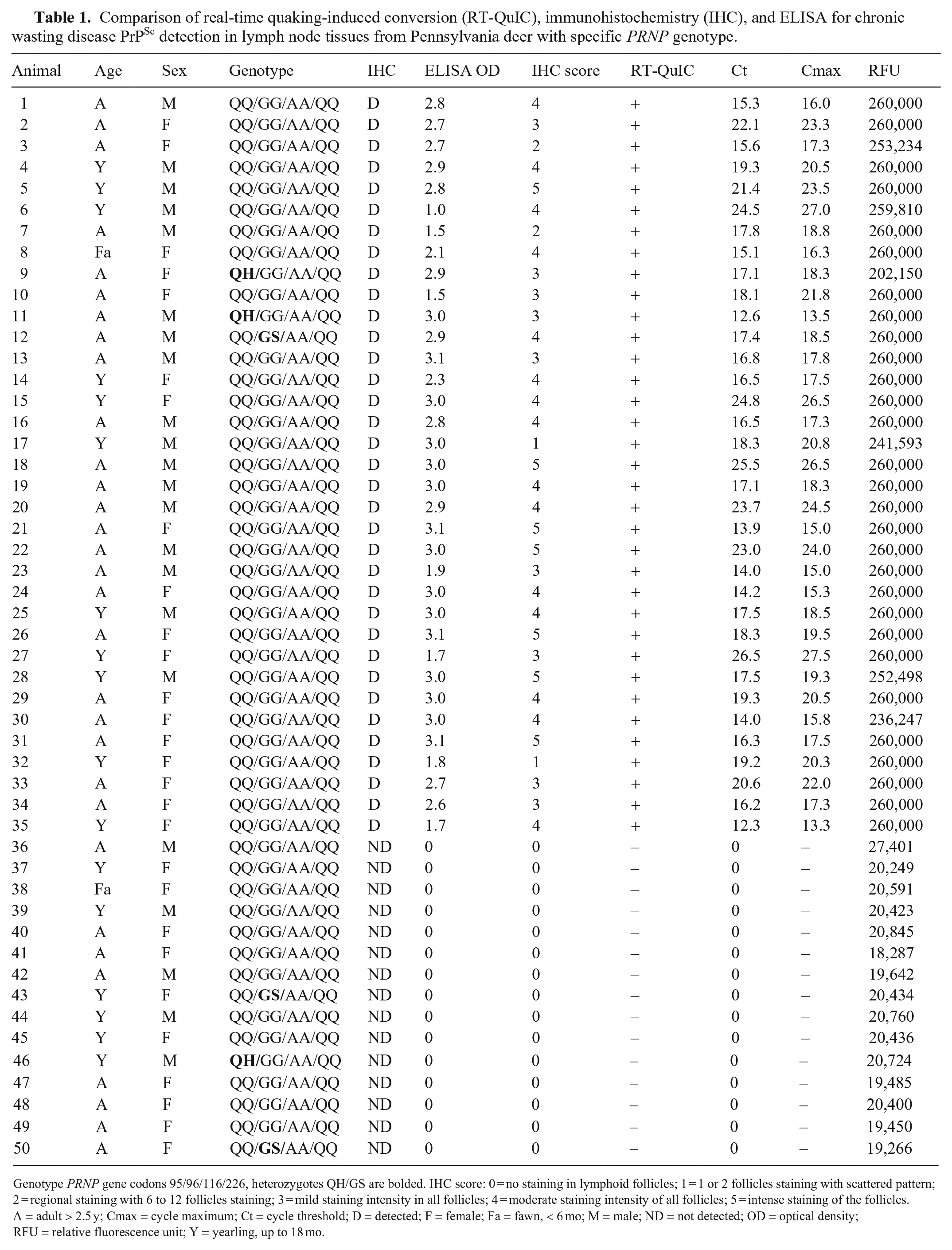
Genotype PRNP gene codons 95/96/116/226, heterozygotes QH/GS are bolded. IHC score: 0 = no staining in lymphoid follicles; 1 = 1 or 2 follicles staining with scattered pattern; 2 = regional staining with 6 to 12 follicles staining; 3 = mild staining intensity in all follicles; 4 = moderate staining intensity of all follicles; 5 = intense staining of the follicles.
A = adult > 2.5 y; Cmax = cycle maximum; Ct = cycle threshold; D = detected; F = female; Fa = fawn, < 6 mo; M = male; ND = not detected; OD = optical density; RFU = relative fluorescence unit; Y = yearling, up to 18 mo.
For genotyping, DNA was extracted from frozen RLNs (EZ1 DNA investigator kit; Qiagen), following the manufacturer’s instructions. The PRNP gene was amplified (Platinum 2X universal mix; Invitrogen) with 0.4 µM each of primers CWD-13 (5′-TTTTGCAGATAAGTCATGGTGAAA-3′) and CWD-LA (5′-AGAAGATAATGAAAACAGGAAGGTTGC-3′). PCR conditions were as follows: 95°C for 5 min, 10 cycles of denaturation at 95°C for 45 s, annealing at 58°C for 45 s, and extension at 72°C for 90 s, followed by 35 cycles of 95°C for 45 s, 57°C for 45 s, and 72°C for 90 s, with final extension at 72°C for 5 min, as described previously. 15 PCR products were sequenced using amplification primers with Sanger sequencing, as described previously. 22 All sequences were individually analyzed for conflicts and secondary peaks and aligned to the O. virginianus reference sequence, AF156185. Specifically, the nucleotides and corresponding amino acid polymorphisms were examined at codons 95 (glutamine Q or histidine H), 96 (glycine G or serine S), 116 (alanine A or glycine G), and 226 (glutamine Q or lysine K).
All 3 methods, ELISA, IHC, and RT-QuIC, detected PrPSc in RLNs from the 35 positive deer chosen for our study (Table 1; Fig. 1B–E). For positive tissues, ELISA OD values were 1.0 to 3.5; although subjective, IHC staining was scored 1 to 5 based on staining intensity and distribution of PrPSc in lymphoid follicles, with 5 representing the highest intensity and number of lymphoid follicles stained.
Confirming a previous study, we demonstrated that a RT-QuIC test was able to detect all deer (100% sensitivity and specificity) that were positive for PrPSc by ELISA and IHC (Table 1), but our detection of PrPSc in a wider variety of subjects such as a fawn increases confidence in the range of the test compared to a report that focused only on adults. 8 Also, the previous study did not include as many positive white-tailed deer, and none of them were from Pennsylvania. 8 In the samples that we tested, all animals exhibited strong signals for PrPSc with the RT-QuIC test as indicated by reaction intensity and by Ct and/or Cmax.
One limitation of our study is that it was not a population study. Despite small numbers included in our study, we included deer from fawn to adult, comprising both sexes. We found no predilection for PrPSc in either sex in our study. All deer were homozygous for 2 codon positions (116 [AA] and 226 [QQ]) within the PRNP gene. For codons 95 and 96, there was low-level heterozygosity in all examined deer at codon 95H/Q and 96G/S with frequencies of 0.06 and 0.08, respectively. Genotyping showed deer were largely homozygous at codons 95, 96, 116, and 226 (QQ/GG/AA/QQ; frequency 0.86) of the PRNP gene, which indicates the existence of CWD-susceptible deer genotypes in Pennsylvania. Genetic heterozygosity said to confer protection (95H/Q or 96G/S) against CWD was associated with positive adult deer in our study group, suggesting that such genotypes did not afford complete protection.
Polymorphism in the PRNP gene has been shown to be important in affording protection against developing disease in several animal species including cervids.2,11,23 Several studies of PRNP genetics in free-ranging and captive white-tailed deer have reported lower infection rates associated with certain genotypes. In particular, the 96S allele has been associated with lower CWD prevalence compared with 96G.2,11,23 Lower CWD infection was suggested in another study for animals having alleles 95H, 116G, and 226K. 20 This area remains under intense investigation, and our data contribute to these studies. For successful CWD control, a combined approach of early detection together with breeding or selecting for resistance, once such markers are clearly identified, is likely to play a significant role, as has been the case for scrapie.1,6
Footnotes
Acknowledgements
We thank Pennsylvania Game Commission staff for tissue collection; Anna Martin Hays, Donna Krouse, Tammy Hocker, and Charles Oboh from Pennsylvania Veterinary Laboratory for technical help; and Dr. Kevin Brightbill of Pennsylvania Department of Agriculture and Dr. Davin Henderson from CWD Evolution Inc. for their support.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
