Abstract
Digital dermatitis (DD) is a major infectious foot disease of cattle worldwide. Some DD stages are associated with lameness, and the disease has significant economic and animal welfare consequences. The pathogenesis of the disease is not yet fully understood, but Treponema spp. have been associated consistently with clinical cases. Isolation of these fastidious bacteria is difficult and cumbersome. We describe an improved method enabling the culturing of the 3 Treponema spp. (T. pedis, T. phagedenis, and T. medium) from bovine foot specimens derived from DD lesions, using a combination of membrane filtering and subsequent growth on selective agar media. The entire procedure from sampling to verification of individual Treponema spp. takes up to 24 d. In addition, we established a MALDI-TOF MS–based identification method to be applied for confirmation of the different Treponema spp. This scheme provides an unambiguous, simple, and straightforward identification procedure for DD-associated Treponema spp.
Digital dermatitis (DD) is a multifactorial contagious foot disease associated with lameness in cattle that has major economic and animal welfare consequences.5,9 The etiology of DD is still in its infancy given the diverse microbial communities involved, but species of the genus Treponema are regularly associated with DD lesions. 13 Secondary infection with DD treponemes is also associated with so-called non-healing foot lesions in cattle.10,14 These lesions involve the corium, have a granular appearance, a pungent smell, are severely painful, and respond poorly to standard treatment such as topical administration of tetracycline or disinfecting solutions. 10
Treponema spp. are gram-negative, anaerobic bacteria that are generally very difficult to culture and isolate. 16 Besides the fastidious nature of treponemes, the high level of contamination of DD specimens makes isolation of treponemes challenging.11,12 Several studies have shown that, in particular, the 3 Treponema spp. (T. medium, T. pedis, and T. phagedenis) can be identified and isolated from DD lesions.2,3,11
Improving isolation protocols for these Treponema spp. will allow more insight into the role of these treponemes in pathogenicity and epidemiology of DD. The isolation and characterization of strains from different countries would in turn help to establish better molecular detection tools. We therefore aimed for an improved culture method for the fastidious Treponema spp.: T. pedis, T. phagedenis, and T. medium.
Lower hindlimbs of slaughtered cattle with signs of DD lesions were collected from 3 Swiss slaughterhouses in cantons Solothurn (n = 4), Bern (n = 4), and St. Gallen (n = 1). Three lower limbs of cattle that did not show any signs of DD or of any other foot disorder were used as negative-control samples originating from pathology examinations of non–foot-related diseases. Additionally, we used tissue samples surgically excised from 2 cattle for standard veterinary treatment of non-healing lesions of axial horn fissures. These tissues were transferred into oral treponeme enrichment broth (OTEB; Anaerobe Systems) with 5 μg/mL of rifampicin and 5 μg/mL of enrofloxacin, placed on ice, transported to the laboratory within 24 h, and processed as described below.
Hoofs were transported to the laboratory within 4 h after slaughter, and dry paper towels were used to remove visible organic debris and contaminants such as soil or feces. The DD lesions were photographed and scored according to protocols described previously4,8: M1 = a small (<2-cm cross-section) focal active lesion; M2 = a larger lesion (>2-cm cross-section) with an active ulcer; M3 = a healing, scabbed lesion; M4 = chronic stages of infection, characterized by hyperkeratosis or surface proliferation; M4.1 = the chronic stage with small reactive M1 lesions; M5 = specified healthy skin with no lesions. Specimens were taken either from the center of the lesions (M4), from the active area (M1 and M2), or from the plantar aspect of the foot between the heels (M5). We did not observe any M3 lesions. The surface was cauterized by using a heated metal spoon, and the specimen to be tested was then taken using a sterile biopsy punch (6-mm diameter biopsy punch, maximum depth of 7 mm; Stiefel).
The following diagnostic procedures (Fig. 1) were applied for isolation of Treponema spp. Briefly, tissue samples were transferred to a sterile petri dish and minced into small pieces (~1-mm2 cubes) using a sterile scalpel blade in an anaerobic workstation (DG250; Don Whitley Scientific) with mixed gas (85% N2, 10% H2, 5% CO2) at 37°C. The minced tissue was then transferred to 7 mL of OTEB supplemented with 10% rabbit serum (Thermo Fisher), 5 μg/mL of rifampicin, and 5 μg/mL of enrofloxacin, then incubated anaerobically at 37°C.11,20 After 4 h, the primary cultures were subcultured on anaerobic enrichment agar (MTGE; Anaerobe Systems) supplemented with 1% rabbit serum without antibiotics; nitrocellulose membrane filters (25-mm diameter, 0.45-µm pore size; MF-Millipore) were placed on the surface of the agar plate (Fig. 2A). 6 Approximately 100 µL of the primary OTEB culture were spotted onto the membrane filter, which was removed after 6 h with sterile forceps, and plates were incubated anaerobically at 37°C for 10 d.

Process of culture isolation and identification of Treponema spp. associated with bovine digital dermatitis. MTGE = anaerobe enrichment agar; OTEB = oral treponeme enrichment broth. * = performed under strict anaerobic conditions (anaerobic workstation); ++ = supplemented with 10% rabbit serum and selective antibiotics; + = supplemented with 10% rabbit serum (1% for MTGE).
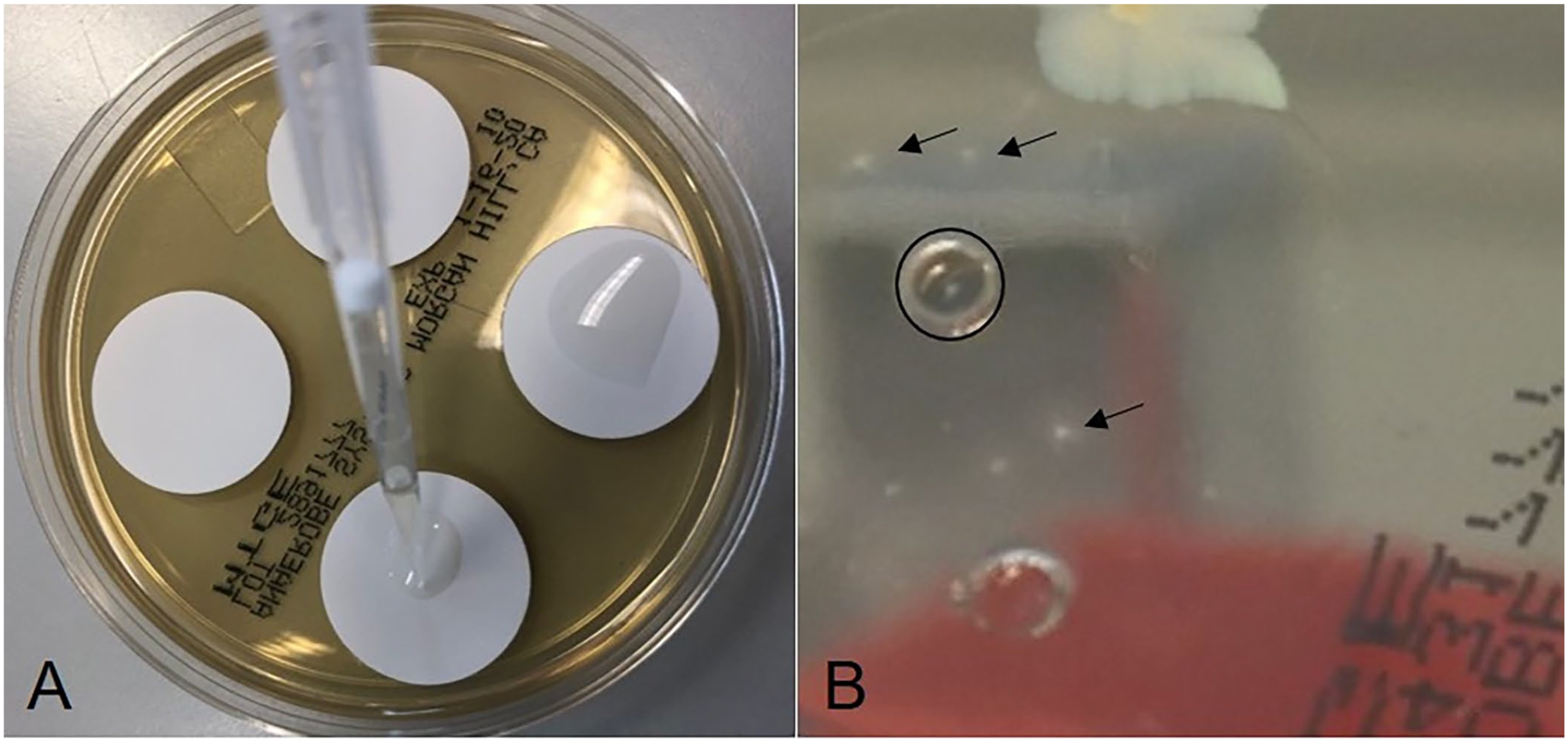
Filter selection and growth of Treponema species on anaerobe enrichment agar (MTGE) agar.
Individual treponemes, which are motile, migrated through the membrane filters and grew on the MTGE agar plate. Typically, the colonies were small (1–2 mm diameter) after 10 d of incubation, appearing similar to spherical cotton balls with a white-to-hazy structure and with irregular or well-defined peripheries (Fig. 2B). The areas of suspicious colonies were picked using a sterile 3-mm biopsy punch; the agar slice was cultured in 1 mL of liquid medium (mycoplasma broth base; Oxoid) supplemented with 10% rabbit serum and 5 µg/mL of cocarboxylase, and 1 mg/mL of L-cysteine hydrochloride but without antibiotics, then incubated anaerobically at 37°C for 10 d. A few quality checks were included in the procedure. From the primary cultures after 4 h of incubation, an aliquot was taken for analysis for the putative presence of T. pedis, T. phagedenis, and T. medium by specific nested PCR (nPCR) assays. 12 Another aliquot was used for dark-field microscopy at 20–40× magnification to check for the presence of viable treponemes based on morphologic features (motile spirochetes). The presence of contaminating microorganisms was checked by plating a 100-µL aliquot of the culture on trypticase soy agar with 5% sheep blood (Becton Dickinson) for 48 h (in both aerobic and anaerobic conditions); colonies indicated contaminating microbes.
Each isolate was confirmed by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS; Microflex LT; Bruker) and an in-house database for identification. 15 The corresponding reference spectra of the T. pedis, T. phagedenis, and T. medium type and reference strains have been deposited (https://maldi-up.ua-bw.de/index_en.asp#EN) and can be shared with interested users. 17 A 500-µL aliquot of the liquid culture was centrifuged at 10,000 × g for 5 min; the pellet was washed with 1 mL of distilled water and centrifuged at 10,000 × g for 2 min. The pellet was suspended in 300 µL of distilled water and then mixed with 900 µL of absolute ethanol. After centrifugation at 10,000 × g for 2 min, the supernatant was discarded. Following an additional centrifugation step, the residual liquid was removed by pipetting. The remaining pellet was air dried, suspended in 50 µL of 70% formic acid, and then mixed with 50 µL of acetonitrile. After centrifugation at 10,000 × g for 2 min, 1 µL of the supernatant was spotted in duplicate on a steel target plate and air dried. Each spot was then overlaid with 1 µL of HCCA matrix (α-cyano-4-hydroxycinnamic acid; Bruker), air dried, and subsequently analyzed. In each case, the Treponema spp. was unambiguously identified with scores ≥2.0; scores of <1.7 were assigned to the next species. The individual cultures were stored at –80°C.
Highly motile spirochetes indicating Treponema spp. were observed in all primary cultures derived from DD and non-healing lesions, but not in the negative-control samples (Table 1). Nested PCR indicated the presence of T. pedis, T. phagedenis, and T. medium in 8, 10, and 5 of the 11 primary cultures derived from lesions, respectively. At least 1 of the 3 DD-associated Treponema spp. was successfully isolated from all DD specimens and non-healing lesions. On 3 occasions, >1 species was isolated. As expected, none of the 3 DD-associated Treponema spp. was cultured from healthy foot tissue, and Treponema-specific DNA could not be amplified using the specific nPCR assay.
Results including dark-field microscopy, nested PCR, and cultivation of Treponema spp. from specimens derived from lesions typical of digital dermatitis (DD) from 3 different Swiss slaughterhouses (1–4 = slaughterhouse of Oensingen, 5–8 = slaughterhouse of Langnau, 9 = slaughterhouse of St. Gallen), non-healing lesions (10, 11), and negative-control samples (12–14).

NA = score not applicable.
Dark-field microscopy of primary culture (oral treponeme enrichment broth [OTEB] inoculated with minced specimen and incubated at 37°C for 4 h). + = highly motile spirochetes present; − = no highly motile spirochetes present.
Specific nested PCR assays as described, 12 to indicate the presence of T. pedis, T. phagedenis, or T. medium.
Nested PCR from primary culture (OTEB inoculated with minced specimen and incubated at 37°C for 4 h).
Pure single colonies obtained after 10 d of incubation at 37°C using the combination of membrane filter and selective media. The final species identification was achieved by MALDI-TOF MS, which confirmed nested PCR findings.
The fastidious nature of treponemes and the high level of contamination of DD specimens require a selective method for their cultivation. Based on the motility of the treponemes, we applied a filter membrane and selective media to foster the isolation of Treponema spp. from DD-derived lesions. Dark-field microscopy aided at several steps of the cultivation procedure to ascertain the enrichment of viable Treponema spp. (Fig. 1). T. phagedenis was the most abundant and readily isolated species, followed by T. pedis and T. medium, in agreement with previous studies demonstrating that T. phagedenis is the most frequent Treponema species found in DD lesions.3,12,19 We isolated T. medium successfully from 1 specimen only, despite its presence in 5 primary cultures as indicated by nPCR. Similarly, T. pedis was isolated successfully from only 4 of the 8 primary cultures positive for this species by nPCR; T. phagedenis was isolated from all of the nPCR-positive primary cultures. This could indicate that T. medium was less likely to survive subculturing. In fact, T. medium did grow more slowly in solid media as compared to T. phagedenis and T. pedis. Additionally, T. phagedenis could outgrow the other 2 species, in particular T. medium.
A strong association has been reported between non-healing foot lesions and Treponema spp. involved in DD lesions. 10 Attempts to isolate Treponema from non-healing lesions have so far failed, despite the occurrence of spirochetes verified by phase-contrast microscopy in such lesions.10,18 Our method enabled the isolation of 3 pure isolates from non-healing lesions (2 T. pedis; 1 T. phagedenis), probably because of the selective power introduced by the filter, which favored the motile bacteria and contributed to fewer contaminating non-motile bacteria.
We did not isolate T. pedis, T. medium, or T. phagedenis from healthy hoof tissue, which supports the association of these Treponema spp. and DD. This was further corroborated by the absence of such Treponema-specific DNA in these samples. A lower abundance of Treponema DNA has been observed in healthy tissue originating from farms affected with DD using swab sampling. 1 This supports the association of Treponema spp. at the onset of DD, 7 but does not indicate cause and effect.
Isolation of Treponema species remains a time- and cost-intensive challenge and needs further improvement and optimization. This could be achieved at the level of tissue processing (e.g., grinding, enzymatic) or selective media including the use of species-specific antibiotics (particularly to suppress growth of T. phagedenis).
Footnotes
Acknowledgements
We thank Drs. Karl Nuss and David Devaux (Clinic for Ruminants, Vetsuisse Faculty, University of Zurich, Switzerland) for collecting the samples of non-healing lesions. We thank the veterinarians in slaughterhouses for their support during sample collection.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our work was supported financially by the Institute of Veterinary Bacteriology and the Clinic for Ruminants at the Vetsuisse Faculty, University of Bern, Switzerland.
