Abstract
Atypical porcine pestivirus (APPV) had been detected in many countries. However, to date, a commercial detection kit is not available because of a lack of specific monoclonal antibodies (mAbs) to APPV. We generated 7 mAbs targeting the NS3 protein of APPV. Isotyping results indicated that all of these mAbs are IgG1 with a kappa light chain. We analyzed the epitopes recognized by mAbs 2B6, 6G11, 8D1, 8D3, and 8F12, which recognized the same linear epitope (GRIKSAYSDE); the 6H3 and 7E10 mAbs recognized 2 different conformational epitopes. Applications of these antibodies were verified by ELISA, western blot, indirect immunofluorescence assay, and flow cytometry. The antibodies were functionally workable for these immunoassays except for 8F12, which could not be used in flow cytometry.
Pestiviruses, which are single-stranded, positive-sense RNA viruses in the continuously growing family Flaviviridae, cause diseases in swine and ruminants. 9 Atypical porcine pestivirus (APPV) was identified as a novel and highly divergent porcine pestivirus distributed in swine herds in the United States. 5 This virus has also been detected in some European and Asian countries, and causes economic losses to the global pig-breeding industry.2,7-9,12,13 Similar to other pestiviruses, APPV contains 12 putative mature proteins: core protein, 3 envelope glycoproteins, P7 protein, and nonstructural proteins (NSs). 5 Among these proteins, NS3 is a multifunctional protein that has nucleoside triphosphatase enzymatic activity, as well as serine proteinase and RNA helicase activities. Uncleaved NS2-3 of pestiviruses is required for viral particle assembly; the release of NS3 is essential for viral RNA replication. 6
Even though APPV has been detected in animals without any clinical signs, many investigations have demonstrated that the presence of APPV genomes in newborn piglets was correlated with congenital tremor (CT) type A-II.1,3,9,10 CT, characterized by muscle spasms at birth, has been classified as type A (including 5 subtypes) and type B. 4 APPV-associated CT is closely related to preweaning mortality of piglets because CT may lead to severe growth retardation and starvation.
The economic loss caused by APPV in pig production worldwide remains undetermined because of asymptomatic infection in adult pigs and the lack of detection tools. Therefore, it is important to establish effective methods to detect APPV to reduce economic losses. To facilitate the study of APPV-NS3 and develop detection tests for APPV, we generated a series of monoclonal antibodies (mAbs) against APPV-NS3 and validated their use in various immunoassays.
To express APPV-NS3 protein, the synthesized APPV NS3 gene (GenBank accession KU041637.1), which is a classical virus strain, 9 was cloned into a pET-30a(+) vector, and the recombinant plasmid was named pET-NS3. pET-NS3 was verified by enzyme digestion and PCR amplification with NS3-specific primers (Suppl. Table 1), universal primers of pET-30a(+), or one of the universal primers with one of the specific primers. Enzyme digestion and PCR verification data suggested that the recombinant plasmid was constructed successfully (Fig. 1A). DNA sequencing analysis indicated that the pET-NS3 was constructed correctly without any mutation (data not shown). To express APPV-NS3 protein, the recombinant plasmid pET-NS3 was transformed into BL21(DE3) Escherichia coli. The transformed cells were cultured at 37°C in lysogeny broth medium containing 50 μg/mL of kanamycin and induced by 0.6 mM isopropyl-β-D-thiogalactopyranosidefor 6 h when the optical density at 600 nm reached 0.6–0.8. All cells were collected by centrifugation and sonicated, and the inclusion body was washed with phosphate-buffered saline (PBS) and centrifuged at 4°C. The pellets were dissolved in 8 M urea and subjected to dialysis in PBS with gradually reduced urea before use. The expressed product was analyzed by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE); NS3 protein was successfully expressed with high purity (Fig. 1B). The presence of recombinant APPV-NS3 protein was verified by western blot analysis by using rabbit anti–6×His-tag polyclonal antibody (Proteintech Group) as primary antibody (Fig. 1C). The recombinant APPV-NS3 protein was successfully expressed and purified, and could be used as an antigen to immunize mice for generation of mAbs.
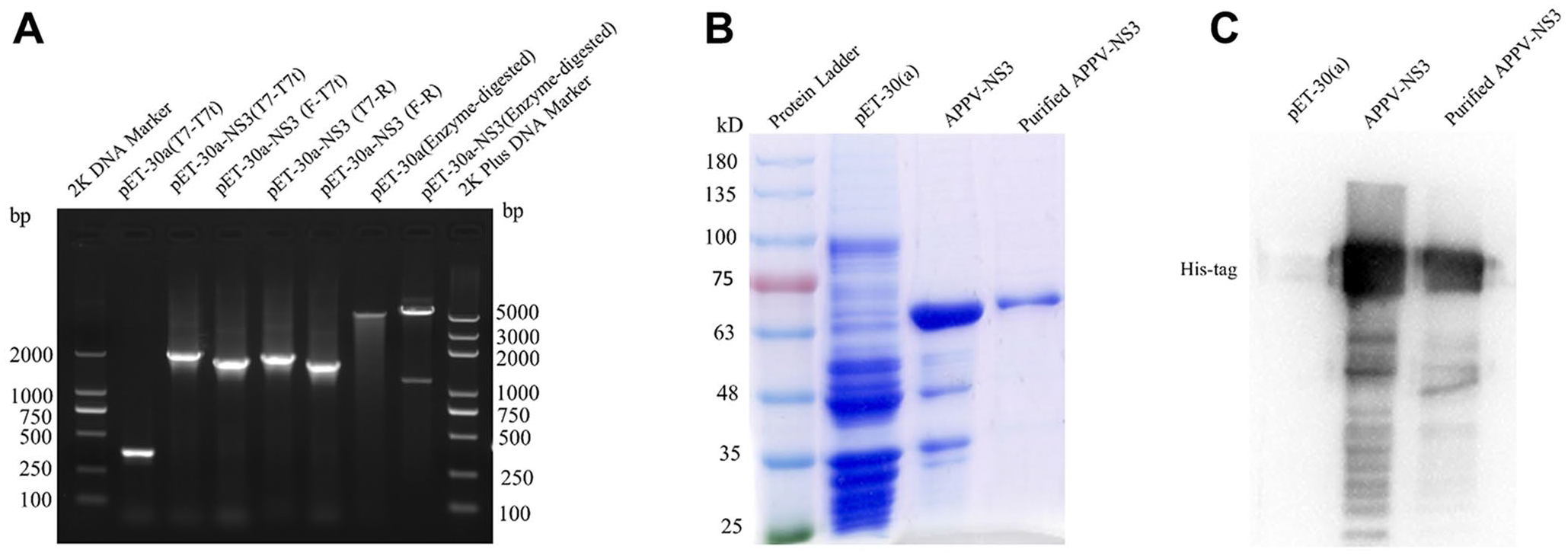
Expression and purification of recombinant atypical porcine pestivirus nonstructural protein 3 (APPV-NS3).
As described elsewhere, 11 BALB/c mice were immunized 3 times with recombinant APPV-NS3 protein with 30% ISA206 adjuvant at 2-wk intervals. Mice were euthanized when the serum antibody level against APPV-NS3 protein reached the plateau phase. The single splenocyte suspension was fused with SP2/0 cells (CRL-1581; American Type Culture Collection). To screen the antibody-secreting hybridoma cells, culture supernatant was collected and detected by indirect ELISA. Hybridoma cells in positive wells were subcloned with a limited dilution strategy. Ultimately, 7 antibody-secreting hybridoma cell colonies were obtained and named 2B6, 6H3, 6G11, 7E10, 8D1, 8D3, and 8F12. To generate ascites fluid with APPV-NS3 mAb, 10 6 hybridoma cells of each clone were injected into the mouse peritoneal cavity after a 2-wk treatment with paraffin oil. The ascites fluid was collected 5–7 d after injection of hybridoma cells. The isotype of prepared mAbs was analyzed by SBA clonotyping system–horseradish peroxidase (HRP; Southern Biotech). All prepared mAbs exhibited strong reactivity with HRP-conjugated goat anti-mouse IgG1 and HRP-conjugated goat anti-mouse kappa, which indicated that the isotypes of all mAb heavy chains were IgG1, whereas all light chains were kappa chains (data not shown).
To determine the epitopes recognized by these prepared mAbs, we constructed 4 truncated NS3 proteins: NS3(1–144), NS3(129–275), NS3(259–402), and NS3(386–528) (Suppl. Fig. 1A). They were all cloned into a pET-30a(+) vector using primer sets (Suppl. Table 1) and transformed into BL21(DE3) competent cells; all 4 truncated NS3 proteins were successfully expressed (Suppl. Fig. 1B, 1C). Then, the 4 truncated APPV-NS3 proteins were used as coating antigen (200 ng) in an indirect ELISA to identify the epitope regions recognized by these mAbs. The results demonstrated that 6 of 7 mAbs had strong binding activity to APPV-NS3(129–275); 7E10 showed reactivity to APPV-NS3(259–402) (Suppl. Fig. 1D). To further locate the epitopes recognized by these mAbs, a series of peptides covering the APPV-NS3(130–402) fragment was synthesized with 20 amino acid overlap between neighboring peptides (Gill Biochemical Biotechnology; Suppl. Fig. 2A). The synthesized peptides were used as coating antigens to measure the reactivity of mAbs by indirect ELISA: 8D1, 8D3, and 8F12 exhibited strong reactivity to both NS3(169–208) and NS3(189–228) fragments, indicating that their epitopes may locate in the NS3(189–208) segment (Suppl. Fig. 2B); 6H3 and 7E10 had strong reactivity to NS3(209–248) and NS3(358–402), respectively; both 2B6 and 6G11 showed strong reactivity to NS3(169–208), NS3(189–228), and NS3(209–248) (Suppl. Fig. 2B). To map the epitopes recognized by these mAbs, another set of peptides covering NS3(189–248) and NS3(358–402) was synthesized (Suppl. Fig. 2C): 2B6, 6G11, 8D1, 8D3, and 8F12 recognized the same linear epitope, GRIKSAYSDE; 6H3 had some reactivity to NS3(219–228) and NS3(224–233) [Suppl. Fig. 2D; i.e., 6H3 might recognize a conformational epitope formed by NS3(209–248)]. Likewise, 7E10 may also recognize a conformational epitope formed by NS3(373–402). The linear epitope, GRIKSAYSDE, recognized by 5 mAbs, and the simple conformational epitope recognized by 6H3, are located in the peptidase_S31 region (Fig. 2); the conformational epitope recognized by 7E10 belongs to the DEXDc superfamily.
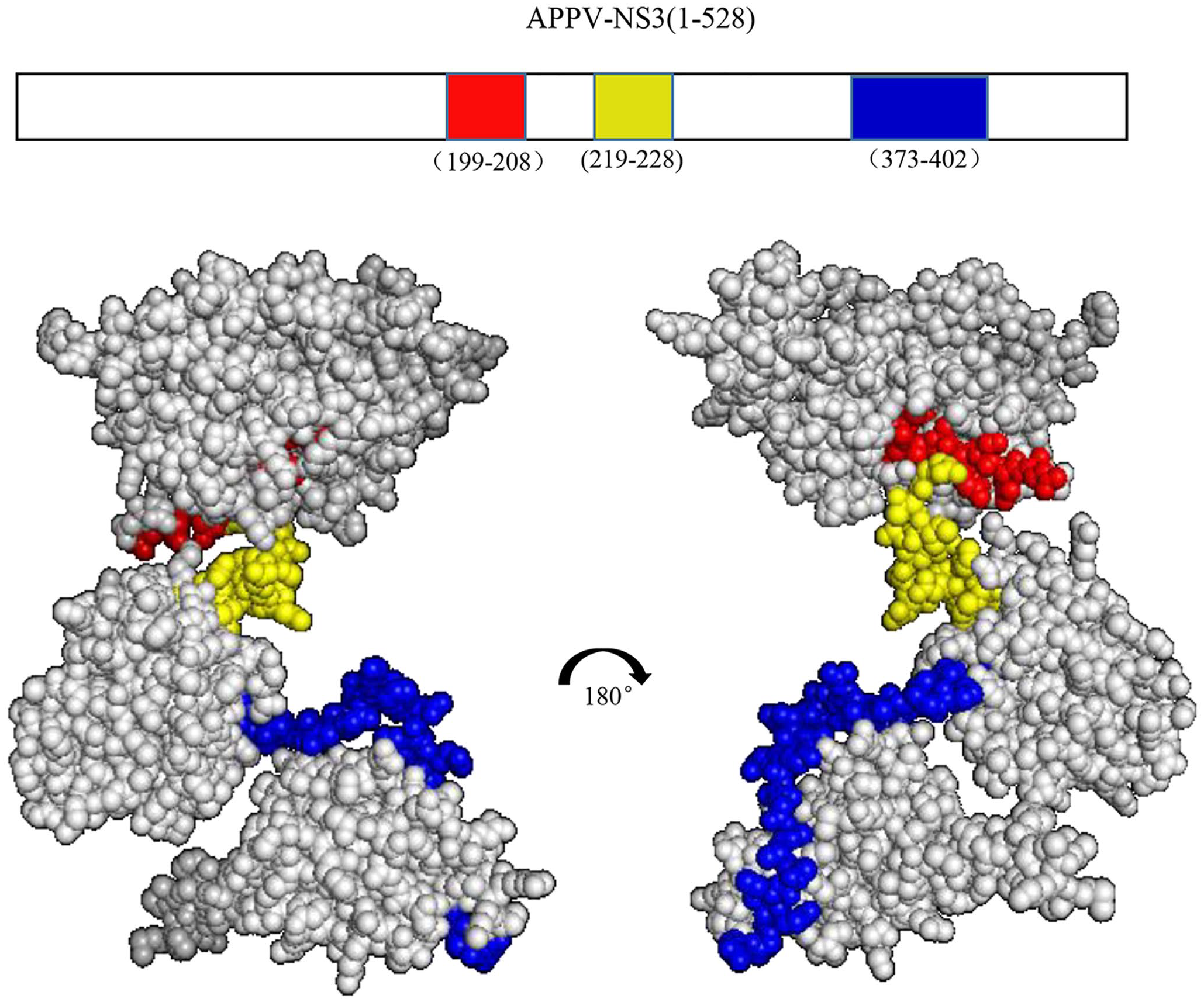
Mapping of epitopes recognized by nonstructural protein 3 (NS3) monoclonal antibodies (mAbs). Positions of antibody epitopes are displayed on an atypical porcine pestivirus (APPV)-NS3 protein structure model. The red region represents APPV-NS3(199–208) recognized by 5 mAbs. The yellow region represents APPV NS3(219–228) recognized by 6H3. The blue region represents APPV NS3(373–402) recognized by 7E10.
We next tested these prepared mAbs in immunoassays. For ELISA, the purified APPV-NS3 protein (concentration of coating antigens was 2 μg/mL, and the mass is 200 ng) was coated on the ELISA plate (Corning) then incubated with all prepared mAbs as the primary antibody. HRP-conjugated goat anti-mouse IgG (Thermo Fisher Scientific) was added to each well as secondary antibody and developed in tetramethylbenzidine chromogenic solution. The readout of the ELISA plate indicated that 6 of the 7 mAbs that we prepared in our study can be used at 1:8,000 dilution, and the 6G11 can be diluted at least 1,000-fold (Fig. 3A).
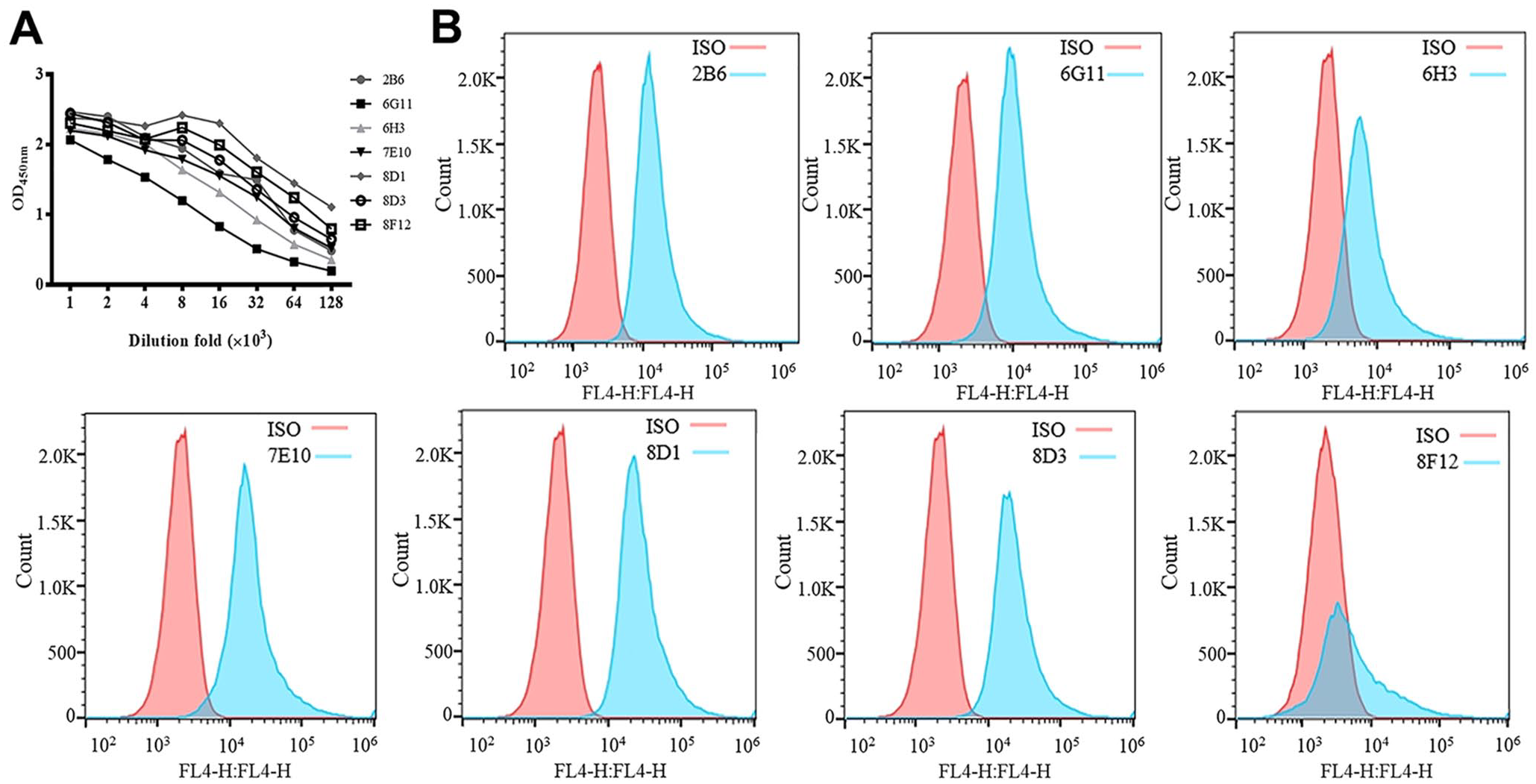
Applications of monoclonal antibodies (mAbs) in immunoassays.
To validate their use in western blotting, immunofluorescence assay (IFA), and flow cytometry assays, the full-length APPV NS3 gene was amplified (Suppl. Table 2) and cloned into pEGFP-c1 and pCMV-HA vectors, named pEGFP-c1-NS3 and pCMV-HA-NS3, respectively. For western blotting, cell lysate transfected with pEGFP-c1-NS3 containing 6×His-tag was detected with 6×His-tag antibody or prepared mAbs as the primary antibody. HRP-conjugated secondary antibodies were used to detect the primary antibodies, and proteins were visualized by electrochemiluminescence. The APPV-NS3 protein was successfully expressed in the pEGFP-c1-NS3–transfected HEK-293T cells (Suppl. Fig. 3A). The results from prepared mAbs showed that all 7 mAbs could be used to detect NS3 protein in pEGFP-c1-NS3–transfected cells; no nonspecific bands were found in the mock-transfected cells (Suppl. Fig. 3B).
To evaluate whether these mAbs could be used to detect NS3 protein by IFA, HEK-293T cells seeded onto coverslips in 24-well plates were transfected with pCMV-HA-APPV-NS3 or empty vector pCMV-HA. The expression of APPV-NS3 was verified by western blotting using anti-HA antibody or mAb 8D3; the target protein was successfully expressed in HEK-293T cells (Suppl. Fig. 3C). After confirmation of NS3 expression, the prepared mAbs were used as primary antibody to react with transfected 293T cells, followed by incubation with AF488-labeled goat anti-mouse IgG secondary antibody (Southern Biotech). The cells were observed under a fluorescence microscope. Most of the cells transfected with pCMV-HA-NS3 showed strong fluorescence (Suppl. Fig. 3D), indicating that all prepared mAbs were usable in IFA for detection of APPV-NS3 protein.
To determinate whether these mAbs were usable in flow cytometry analysis, HEK-293T cells were transfected with pCMV-HA-NS3 then permeated and fixed. Prepared mAbs or mouse IgG1 isotype control (Abcam) were used as primary antibodies, and AF647-labeled goat anti-mouse IgG (Abcam) was used as a secondary antibody for flow cytometry staining. Six of these prepared mAbs, except 8F12, could be used as a primary antibody to detect APPV-NS3 protein by flow cytometry (Fig. 3B). During the experimental process, we found that mAb 8F12 decreased cell numbers significantly (data not shown), implying that it might be toxic to HEK-293T cells.
We selected the APPV-NS3 protein as an antigen to generate mAbs, given that NS proteins are not part of the virions but play a critical role in virion formation and genome replication. A hybridoma clone can only recognize one epitope. Antigen epitopes can be divided into linear epitopes and conformational epitopes. In our study, some mAbs recognized linear epitopes but others recognized conformational epitopes. These mAbs will significantly improve the applied and basic research of APPV and facilitate the surveillance investigation, prevention, and control of APPV.
Supplemental Material
Supplemental_material – Supplemental material for Preparation, identification, and functional analysis of monoclonal antibodies against atypical porcine pestivirus NS3 protein
Supplemental material, Supplemental_material for Preparation, identification, and functional analysis of monoclonal antibodies against atypical porcine pestivirus NS3 protein by Miaomiao Yan, Jingling Huang, Jianing Chen, Wenting Yang and Guangliang Liu in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Dr. Manita Aryal for proofreading this manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the Elite Youth Program of the Chinese Academy of Agricultural Sciences, and partly by China Central Public-interest Scientific Institution Basal Research Fund 1610312020020.
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
