Abstract
Two Actinobacillus pleuropneumoniae isolates from clinical cases of porcine pleuropneumonia in Japan were positive in the capsular serovar 15–specific PCR assay, but nontypeable (NT) in the agar gel precipitation (AGP) test. Nucleotide sequence analysis of gene clusters involved in the biosynthesis of capsular polysaccharide (CPS) and lipopolysaccharide O-polysaccharide (O-PS) revealed that both clusters contained transposable element ISApl1 of A. pleuropneumoniae belonging to the IS30 family. Immunoblot analysis revealed that these 2 isolates could not produce O-PS. We conclude that the ISApl1 of A. pleuropneumoniae can interfere in the biosynthesis of both CPS and O-PS.
Keywords
Actinobacillus pleuropneumoniae is the causative agent of porcine pleuropneumonia, an economically important bacterial infection of swine. 5 To date, 18 serovars have been recognized, mainly on the basis of the antigenic diversity of capsular polysaccharides (CPS) and the O-polysaccharide (O-PS). 2 Characterization of the A. pleuropneumoniae serovar involved in an outbreak is useful for understanding the epidemiology of that outbreak, for preparing vaccines for the control of the disease, and for serologic monitoring of infected herds.4,5 Cross-reactions between some serovars (1, 9, and 11; 4 and 7; 3, 6, 8, and 15) and variable results between individual batches of test sera are usually observed in the conventional serotyping tests.4,5 To overcome these limitations, PCR assays based on the capsule loci have been developed for serotyping of A. pleuropneumoniae isolates.2,18
Sequencing the whole genome or CPS loci allows re-evaluation of previously nontypeable (NT) isolates,7,8 as well as those that had an unusual serologic or molecular test result. 6 Comparative sequencing analysis of the capsular loci of serovar 15 isolates revealed that the presence of insertion sequence ISApl1 in CPS loci could interfere with both the serologic and molecular typing methods. 8 Earlier studies have found that element ISApl1 in the reference and field isolates of serovar 7 interfered with biofilm formation, 10 and the production of ApxIV. 15
We report herein the isolation of serologically NT variants of A. pleuropneumoniae. We describe the genetic characteristics of the gene clusters involved in the biosynthesis of CPS and antigenic O-PS, and the defect in the production of CPS and O-PS in these isolates.
A. pleuropneumoniae–like organisms were isolated in January 2019 from the lung lesions of 2 fattening pigs at ~ 150 d of age suffering from acute pleuropneumonia. The lung specimens were inoculated on chocolate agar (Becton, Dickinson) and incubated at 37°C in the presence of 5% CO2. One colony from each of the 2 isolation plates was chosen for analysis. Species-specific identification based on partial 16S ribosomal DNA sequencing, performed as described previously,9,17 showed that the nucleotide sequence of a 919-bp hypervariable region of the 16S RNA gene of the 2 isolates had homology of 99.0–100% with those of other A. pleuropneumoniae isolates available in GenBank. Results of the PCR-based apx gene typing 9 revealed that the 2 field isolates contained apxIIA, apxIIIA, and apxIVA genes. The molecular work and the fact that the isolates were found to be dependent on nicotinamide adenine dinucleotide (NAD) in vitro, indicated that both isolates were A. pleuropneumoniae biovar 1. Further examination of the genes for the expression, activation, and secretion of Apx toxins by PCR1,17 revealed that these 2 isolates of A. pleuropneumoniae, named IB19-1 and IB19-2, contained the same apx gene combination as serovars 2, 4, 6, 8, and 15 (an apxIICA, apxIIICA, apxIBD, apxIIIBD, and apxIVA gene pattern). The serovar 15 reference strain HS143 1 and 3 representative field isolates (A–C) 14 of serovar 15 isolated from different geographic regions in Japan were used in the further characterization of these isolates as described below.
Serotyping of the 2 field isolates in our study, as well as the serovar 15 reference strain and the 3 earlier field isolates of serovar 15, was carried out with serovar-specific rabbit antisera prepared against serovar reference strains by using an agar gel precipitation (AGP) test as described previously. 17 All 3 earlier field isolates of serovar 15 and the reference serovar 15 strain, but not isolates IB19-1 and IB19-2, were serotyped as serovar 15 by the AGP test, based on the observation of serovar 15–specific precipitation lines (Fig. 1). The IB19-1 and IB19-2 isolates were identified as serovar 15 in a serovar 15–specific PCR assay. 18 Our results suggested that these 2 isolates were defective in CPS production.
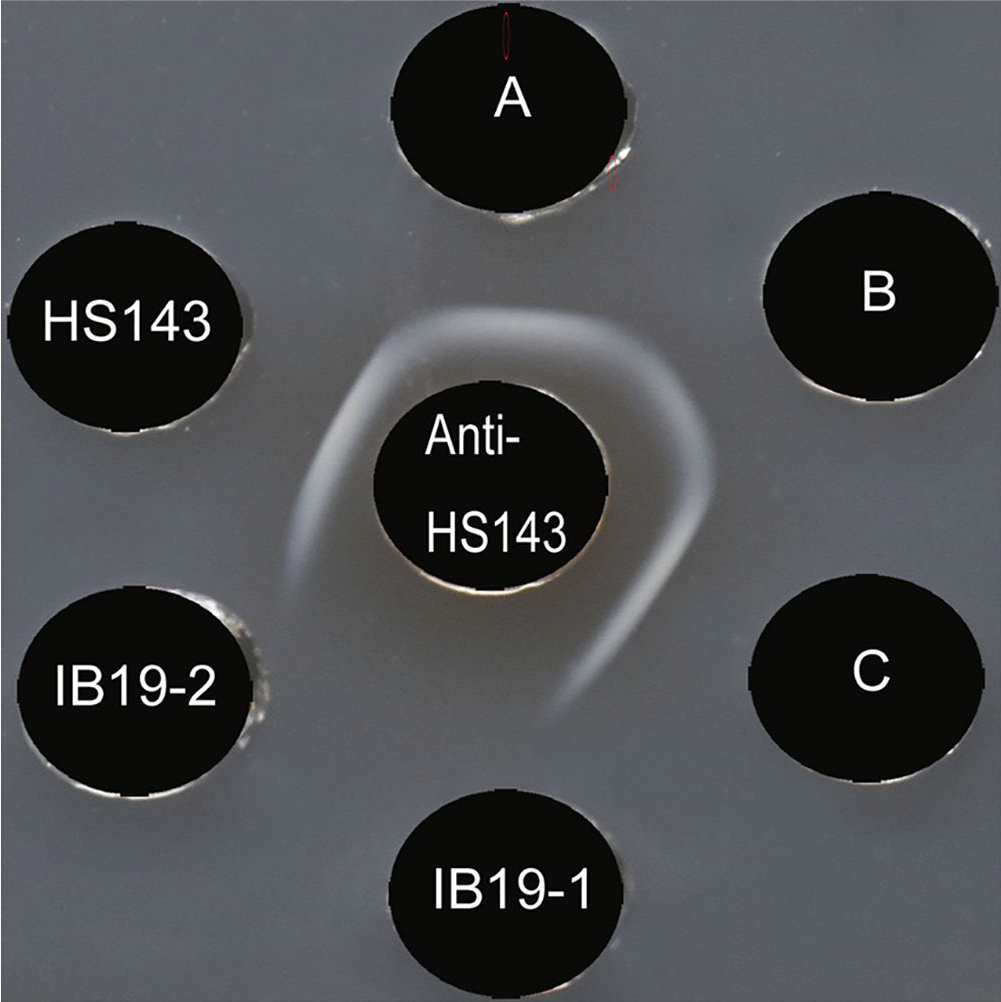

Agar gel precipitation test with rabbit antiserum raised against Actinobacillus pleuropneumoniae serovar 15 reference strain HS143. The antiserum against reference strain HS143 was placed in the center well (labeled anti-HS143). Antigens of the serovar 15 reference strain HS143 and the field isolates (labeled A, B, C, IB19-1, and IB19-2, respectively) were placed in the outer wells. Precipitation lines were not observed for the IB19-1 and IB19-2 isolates.
In order to elucidate the molecular basis of the serologic nontypeability of these 2 isolates, the gene clusters involved in the biosynthesis of CPS and O-PS were PCR-amplified from isolates IB19-1 and/or IB19-2. The primer pairs used for amplification of the cps 2 and o-ps6,20 genes were designed from previously published sequences (Suppl. Table 1). The PCR was performed using ingredients and conditions for PCR that were given in the protocol provided with the enzyme used, the Tks Gflex DNA polymerase (Takara Bio). Template DNA was prepared as described previously. 14 Briefly, the conditions for PCR were as follows: 94°C for 1 min (1 cycle), 98°C for 10 s, and 68°C for 3 min 30 s (30 cycles). The PCR products were purified (NucleoSpin gel, PCR clean-up kit; Macherey-Nagel), and the nucleotide sequences were determined using a primer-walking procedure as described previously. 16
The complete CPS loci (13,931 bp) of isolates IB19-1 and IB19-2 were found between the genes ydeN, encoding a predicted serine hydrolase, and modF, encoding a putative molybdenum transport ATP-binding protein (Fig. 2A). ISApl1 (ISApl1a) was found between the genes lysA and cps15A in the NT isolates, but not in the reference strain HS143 (Fig. 2A).
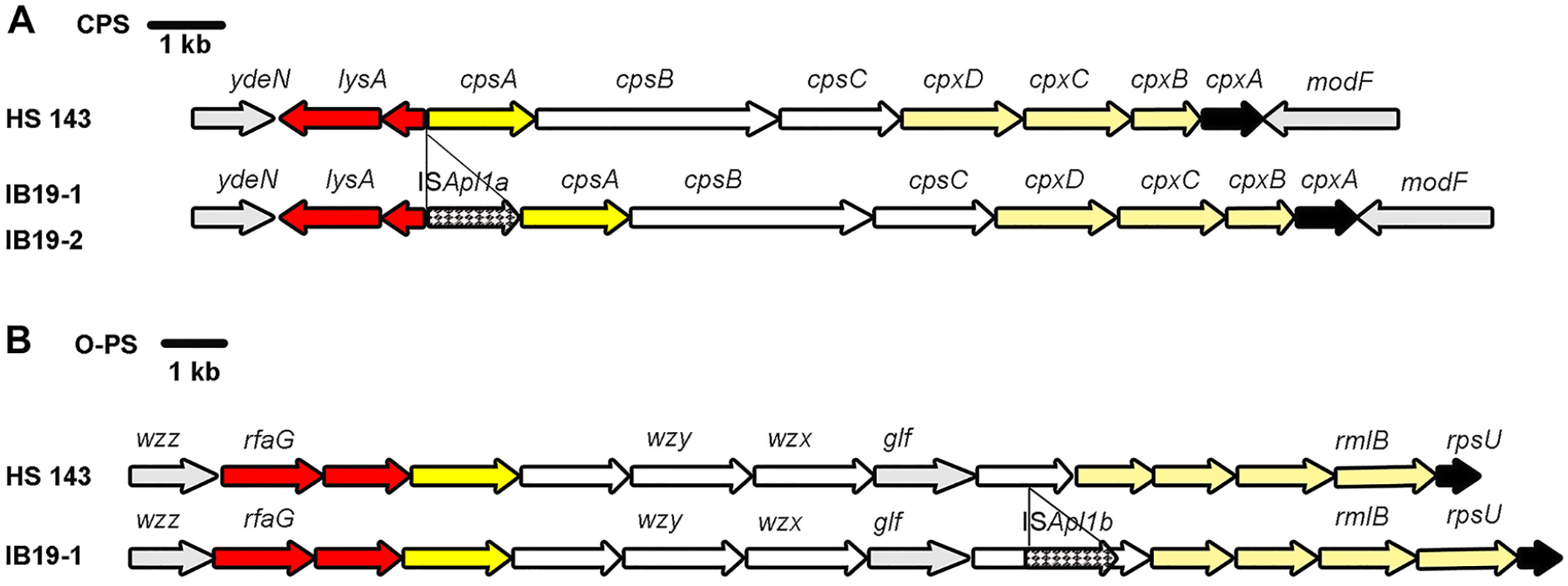
Schematic comparison of the complete capsular polysaccharide (CPS) biosynthesis loci (
The complete O-PS locus of isolate IB19-1 (15,257 bp) was found between the genes wzz, encoding a Wzz-like protein, and rmlB, encoding a dTDP-glucose 4,6-dehydratase (Fig. 2B). ISApl1 (ISApl1b) was found within the gene encoding undecaprenyl-phosphate galactose (Und-P Gal) phosphotransferase 19 (436 aa) in the NT isolate IB19-1, but not in the reference strain HS143. ISApl1 was located 792 bp downstream from, and in the same orientation as, the gene start codon. The nucleotide sequence of target site duplication (TSD) flanked by ISApl1b (GG and AT) was distinct from that flanked by ISApl1a (GC and GC; data not shown). Nucleotide sequences of the CPS and O-PS loci of isolate IB19-1 were deposited in the DNA Data Bank of Japan (DDBJ)/European Molecular Biology Laboratory (EMBL)/GenBank databases as accessions LC507611 and LC508021, respectively.
To investigate whether the IB19-1 and IB19-2 isolates were defective in O-PS production, the lipopolysaccharide (LPS) of the reference strain (HS143) and 5 field isolates (A, B, C, IB19-1, and IB19-2) was extracted and analyzed by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and immunoblot assay as described previously. 17 The results of immunoblot analysis showed that the rabbit antiserum raised against serovar 15 reference strain HS143 reacted with long-chain LPS, giving a ladder-like pattern typical of smooth LPS (O-PS), with 2 fast-migrating bands corresponding to the core and lipid A structures in the LPS preparations of the reference serovar 15 strain (Fig. 3). A similar pattern also occurred with the 3 earlier field isolates (A–C) but only 2 fast-migrating bands were seen in the LPS preparations of isolates IB19-1 and IB19-2 (Fig. 3). The results of the immunoblot analysis showed that the 2 serologically NT isolates IB19-1 and IB19-2 could not produce O-PS.
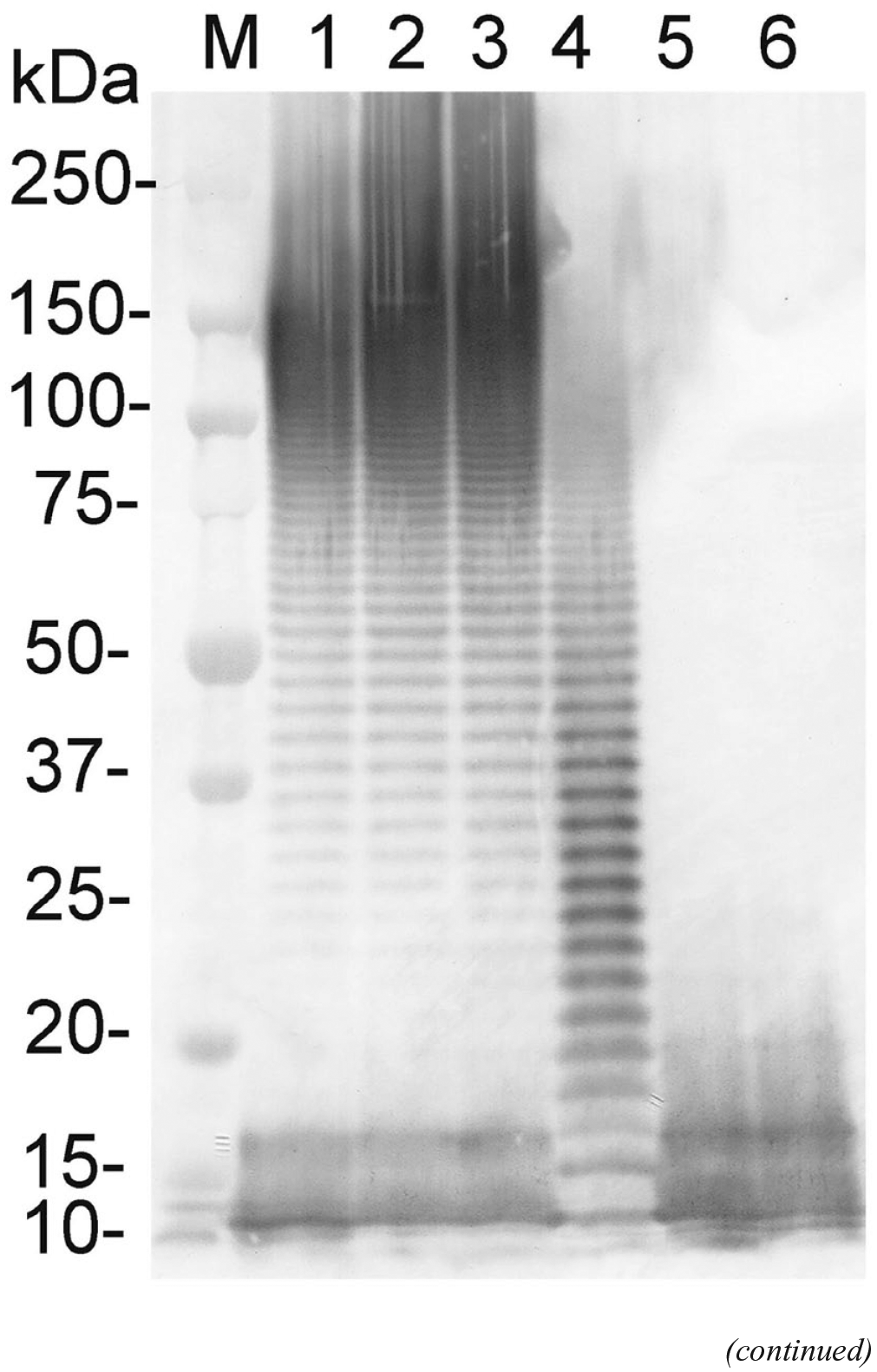
Immunoblot analysis of the purified lipopolysaccharide (LPS) preparations of Actinobacillus pleuropneumoniae serovar 15 reference strain (HS143) and 5 field isolates. The immunoblot was probed with rabbit antiserum against HS143.
A previous investigation showed that the insertion of ISApl1 into one of the genes involved in biosynthesis of CPS as well as the deletion of these genes are responsible for the serologic nontypeability of 2 A. pleuropneumoniae field isolates. 8 The location and orientation of the ISApl1a insertion in the CPS loci of isolates IB19-1 and IB-2 were the same as those of previously identified serovar 15 isolates QAS68, QAS83, and QAS102, 8 which were identified as serovar 15 strains by the AGP test. 8 The reasons for the discrepancy between the previous results 8 and our result is not clear, but may reflect genetic variations in the O-PS biosynthesis gene clusters between the different serovar 15 isolates investigated in the 2 studies. In our study, the gene encoding Und-P Gal phosphotransferase at the O-PS locus of isolate IB19-1 was disrupted by the insertion of ISApl1b. It is reasonable to assume that disruption in the gene encoding Und-P Gal phosphotransferase impairs biosynthesis of both O-PS and CPS.
The Und-P Gal phosphotransferase of A. pleuropneumoniae is homologous to another well-described Und-P Gal phosphotransferase, WbaP. 19 It is well known that WbaP is required to catalyze the transfer of Gal-1-P from uridine-P to the lipid carrier Und-P to yield Und-P-P-Gal, which primes the biosynthesis of both O-PS and CPS in gram-negative bacteria such as group 1 capsule-producing Escherichia coli, 3 Klebsiella pneumoniae, 3 Kerstersia gyiorum, 11 Providencia spp., 13 Salmonella enterica, 3 and Vibrio vulnificus. 12 Genes that contribute to O-PS biosynthesis may also contribute to CPS biosynthesis, as both antigens are comprised of repeating polysaccharide units, and that genes involved in polysaccharide biosynthesis, polymerization, and transport may naturally be shared among both processes.3,11–13,21 However, little is known regarding the contribution of A. pleuropneumoniae O-PS biosynthesis genes to the biosynthesis of CPS. Therefore, further studies would be needed to elucidate whether the O-PS biosynthesis genes of A. pleuropneumoniae contribute to the biosynthesis of CPS.
Results from the AGP test and immunoblot analysis suggest that the NT isolates IB19-1 and IB19-2 were defective in both CPS and O-PS production. Our results, taken with the previously described observations, 8 support the idea that ISApl1 insertions in the CPS and/or O-PS loci of isolates with serovar 15–specific capsule genes can render them serologically NT. A previous study has shown that there were 4 kinds of nucleotide sequences of the TSD (TG, TT, GG, and AA) for ISApl1. 15 Two new TSD sequences (GC and TA) have been identified in serovar 15 in Japan, 8 where isolation of serovar 15 has recently increased.9,17 In our study, one new TSD sequence (AT), as well as the previously reported TSD sequences (GC and GG; data not shown), were identified.
Our antigenic and genetic analyses of 2 serologically NT isolates revealed ISApl1 insertions in the CPS and O-PS loci, resulting in the absence of both K and O antigens. Veterinary diagnostic laboratories should be vigilant, given that infections caused by such an isolate will not be detected by serologic tests based on long-chain LPS.
Supplemental Material
Supplemental_material – Supplemental material for Characterization of nontypeable Actinobacillus pleuropneumoniae isolates
Supplemental material, Supplemental_material for Characterization of nontypeable Actinobacillus pleuropneumoniae isolates by Ho To, Kaho Teshima, Michiha Kon, Saori Yasuda, Yuta Akaike, Kazumoto Shibuya, Shinya Nagai and Chihiro Sasakawa in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Dr. P. Blackall (The University of Queensland, Australia) for providing serovar 15 reference strain HS143, and Dr. F. Koike (Toyoura Veterinary Clinic, Japan) for arranging the collection of the bacterial isolates. We thank Dr. T. Sato and Ms. H. Kojima (Nippon Institute for Biological Science) for their advice during this work.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
