Abstract
Veterinary pathology tissue banks are valuable resources for genetic studies. However, limited data exist as to whether quality DNA can be extracted from these tissues for use in canine genotyping studies. We extracted DNA from 44 formalin-fixed, paraffin-embedded (FFPE) tissue blocks from dogs; 9 of these dogs had DNA available from whole blood samples that had been banked. We genotyped DNA from 30 of 44 tissue blocks and 9 whole blood samples on the Illumina CanineHD BeadChip; DNA quality was insufficient in 14 of 44 samples from tissue blocks. There was significant correlation between the 260/280 ratio and single-nucleotide variation (SNV) call rate (
Genetic studies are gaining popularity in veterinary medicine with the development and application of advanced molecular techniques. Newer applications of molecular technology, such as genome-wide association studies and whole-genome sequencing, have provided new insight into the heritability of disease in veterinary species. However, because of the reliance of these techniques on high-quality DNA, specimens must generally be collected prospectively; alternatively, banked DNA that has been extracted from fresh specimens must be used. Veterinary pathology tissue block repositories provide a wealth of tissue material for evaluation using conventional pathology techniques but are undeveloped resources for the application of genome studies. Veterinary teaching institutions and commercial laboratories bank thousands of tissue blocks each year, and new opportunities for such studies exist if high-quality DNA can be recovered from these tissue blocks. Genome studies within a single breed can be performed with small sample sizes, 7 but analytical power is enhanced by large sample sizes. The availability of DNA from tissue banks would also allow investigation of uncommon diseases with potential translational application to human medicine. 5 Previous studies have documented the use of formalin-fixed, paraffin-embedded (FFPE) tissues for molecular techniques, such as PCR in veterinary medicine1–3 and medical studies of tumor-bearing people,8,9 but there is a lack of published data about the use of these tissues for veterinary single-nucleotide variation (SNV) array genotyping.
The optimal quality of DNA samples submitted for SNV array genotyping in canine studies is not well defined. SNV array genotyping generally requires 50 μg of DNA, and, anecdotally, 260/280 ratios > 1.7 result in adequate call rates. When DNA samples submitted for SNV genotyping result in call rates > 95%, they are generally accepted as high quality4,6,7; however, studies in human medicine have suggested that call rates > 65% might be adequate for SNV array genotyping from FFPE samples. 9
Herein we describe the methods used to extract DNA samples from canine FFPE samples. We hypothesized that DNA extracted from FFPE tissue blocks could be used for SNV analysis using the Illumina CanineHD BeadChip. Our project was undertaken to increase the sample size of a study of genetic susceptibility to disseminated fungal infections in German Shepherd dogs.
The veterinary medical record system at the University of California–Davis, Veterinary Medical Teaching Hospital (VMTH) was searched to find both autopsy and biopsy samples using the keywords “
Our data are presented with descriptive statistics using median and interquartile range (IQR) for nonparametric data or mean and SD, as appropriate. Comparisons between groups were made using the Mann–Whitney U or Student
From the 44 blocks (23 cases and 21 controls), six 5-μm thick scrolls of tissue sections were obtained from the VMTH pathology service. To prevent DNA cross-contamination of specimens, the microtome was cleaned (RNase away; Molecular Bioproducts) between each tissue block. When available, scrolls with spleen, liver, bone marrow, and lymph node were requested because of the suspicion that these organs would contribute higher DNA concentrations given their high leukocyte density. For biopsy specimens, any available tissue was requested. Given that most tissue blocks were obtained at autopsy, most contained ≥1 tissue (median: 2; range: 1–6). The date of acquisition and tissues present within the FFPE block were recorded.
Initially, DNA extraction was attempted on 6 tissue blocks of unaffected dogs (QIAamp DNA blood mini kit; Qiagen). Approximately 25 mg of tissue was manually trimmed from the embedding paraffin, and the scrolls were deparaffinized using 3 cycles of xylene wash. Following extraction, DNA concentration and quality were assessed using optical spectrophotometry (NanoDrop; Thermo Fisher Scientific). The mean concentration of DNA from these samples was 3.65 µg/µL (±3.57), and the 260/280 ratio was 1.61 (±0.38). These results are not included in the subsequent analysis.
Given marginal yields using this extraction technique, we then used another kit (Quick-DNA FFPE kit; Zymo Research). In brief, this kit uses a proprietary deparaffinization solution, followed by tissue digestion (proteinase and RNase) and DNA purification. This process resulted in 50 µL of eluted DNA in buffer solution. DNA for controls from whole blood was extracted using the QIAamp DNA blood mini kit. Extracted DNA was stored at −20°C until it was used for SNV array.
DNA extracted from all FFPE tissues had a median concentration of 57.3 μg/µL (IQR: 21.3–144.9) and 260/280 ratio of 1.81 (IQR: 1.75–1.89; Fig. 1). Given the poor extraction from some FFPE-extracted samples, only DNA with a concentration > 15 μg/µL and 260/280 ratio > 1.6 was submitted for SNV analysis (
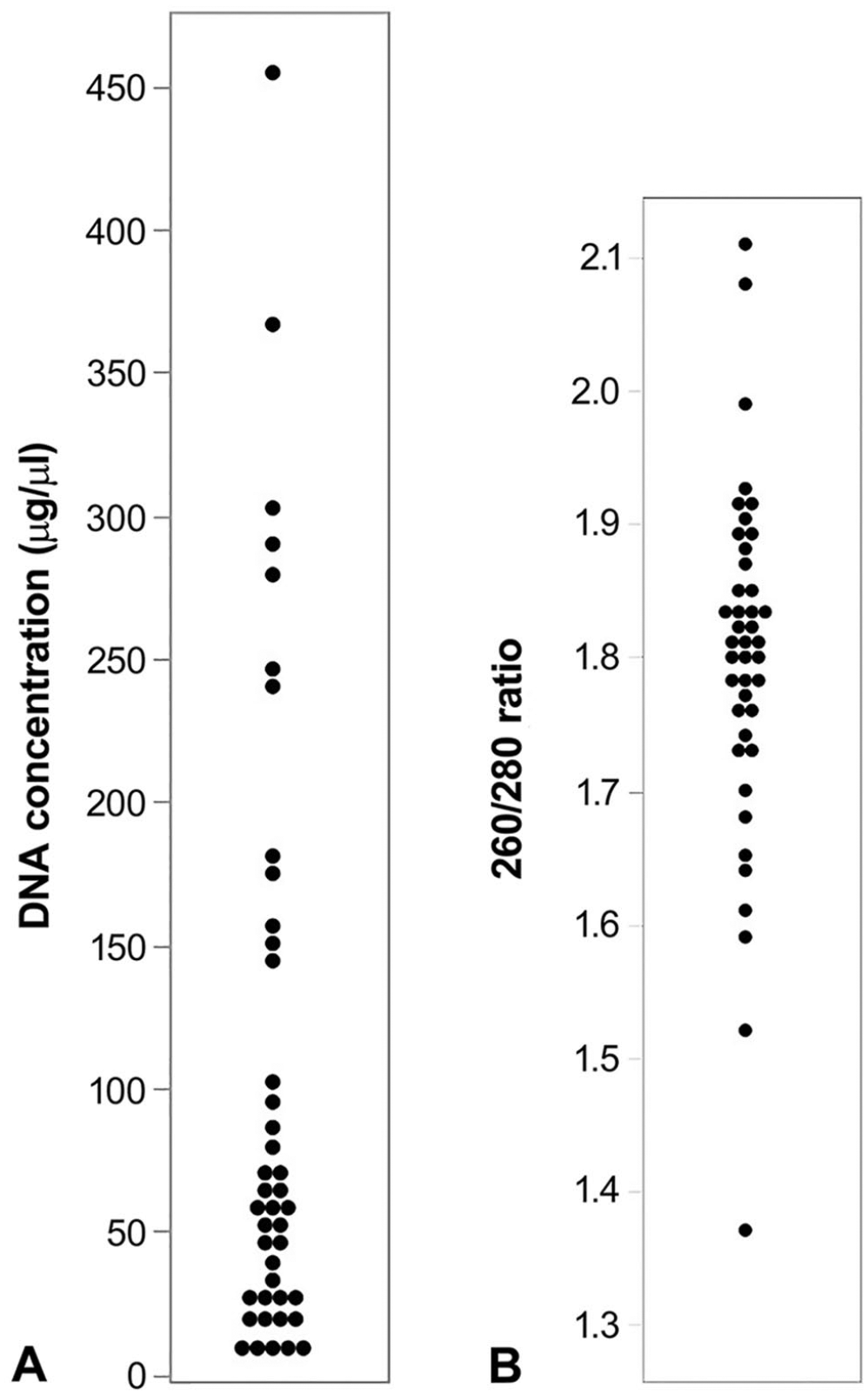
Results of DNA extraction from 44 formalin-fixed, paraffin-embedded (FFPE) tissue blocks.
In total, 30 DNA extracts from FFPE tissue blocks were submitted for genotyping (Illumina CanineHD BeadChip; Neogen Genomics). Of these, 9 extracts were matched with DNA extracts from whole blood. Overall, the FFPE extracts resulted in a median call number of 166,647 (IQR: 144,227–200,539) with a rate of 75.5% (IQR: 65.3–90.8%; Suppl. Table 1). Of the 30 total FFPE extracts submitted for SNV analysis, 23 resulted in a call rate > 65%. There was significant weak correlation between 260/280 ratio and call rate (
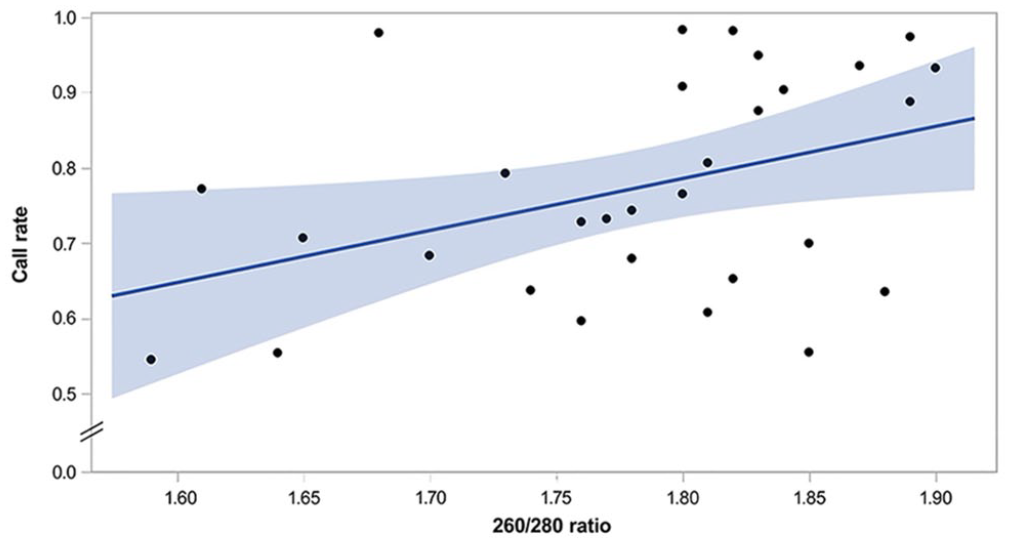
Results of DNA extraction from 30 formalin-fixed, paraffin-embedded tissue blocks. This scatterplot depicts a linear regression modeling single-nucleotide variation call rate on the y-axis as predicted by DNA 260/280 ratio on the x-axis, with slope of 0.6903 and intercept of −0.457 (
Of the paired FFPE and whole blood DNA extracts, DNA concentrations and 260/280 ratios were consistently higher in extracts from whole blood than those from FFPE (median difference: 83.5 µg/µL and 0.1, respectively). All DNA extracts from whole blood had call rates > 95% (median: 98.3%; IQR: 98.3–98.6%) and 260/280 ratios greater than our arbitrary cutoff of 1.6 (mean: 1.89 ± 0.027; Suppl. Table 1). Eight of 9 FFPE extracts had call rates > 65%. The mean Δ homozygosity for all matched paired extracts was 17,807 (±16,919). Delta-homozygosity was lower in samples with call rates > 65% (13,957 ± 13,217) when compared to the matched pair with a call rate < 65% (48,607;
In total, 25 FFPE blocks had tissue from organs of interest (liver 13, spleen 22, bone marrow 1, lymph node 5). The presence of none of the tissues was associated with significant differences in DNA concentration, 260/280 ratio, and call rates. The age of FFPE blocks was 1–24 y (median: 7.5). There was no significant relationship between age of tissue blocks and DNA concentration (
We demonstrated that DNA extracted from FFPE tissue blocks can be used for veterinary genomic studies such as SNV array genotyping, although genotyping rates are much lower than those from blood samples. This result is similar to results of studies of FFPE DNA extraction and SNV array genotyping from human samples. In previous studies, mean call rates from FFPE samples were 69.4–83.4%, lower than call rates from fresh frozen (FF) tissue samples (89.4–93.6%).8,9 These studies demonstrated a high rate of agreement in genotyping between FF and FFPE samples with a positive relationship between call rate and genotyping agreement. The quality of DNA extracted cannot be predicted based on tissue type or specimen age, but higher 260/280 ratios are weakly but positively correlated with SNV call rate. The clinical implications of this association are unclear but suggest that protein contamination, along with other factors, influences call rates in DNA extracts from FFPE tissue blocks.
A limitation of genotyping from FFPE tissues is the intrinsic degradation of DNA caused by formalin. This might also be exacerbated by tissue digestion leading to DNA fragmentation. In our study, we were only able to extract DNA of sufficient concentration and quality for SNV analysis in 30 of 44 tissue block samples. Despite this screening, only 23 of these 30 samples resulted in call rates > 65%, resulting in an overall genotyping rate of 52%. We only evaluated 2 methods of DNA extraction from FFPE tissues; a broader comparison of kits and methods should be performed to assess the ability to genotype animals using different approaches. Although whole blood and FFPE extract SNV calls were not directly compared, comparable levels of homozygosity (indicated by lower Δ homozygosity) suggest that extracts with call rates > 65% provide similar results when analyzed on the Illumina CanineHD BeadChip.
Supplemental Material
Supplemental_material – Supplemental material for Quality of DNA extracted from formalin-fixed, paraffin-embedded canine tissues
Supplemental material, Supplemental_material for Quality of DNA extracted from formalin-fixed, paraffin-embedded canine tissues by Jonathan D. Dear, Jane E. Sykes and Danika L. Bannasch in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Ms. M. Aguilar and Ms. M. Singer-Berk for their technical assistance in preparing samples for analysis.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Funding for this project was provided by the Center for Companion Animal Health, School of Veterinary Medicine, University of California–Davis. The project was also supported by the National Center for Advancing Translational Sciences, National Institutes of Health (NIH), through grant UL1 TR001860. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
