Abstract
Uveitis is common in cats, and is often a feature of feline infectious peritonitis (FIP). We evaluated 3 tools for detection of feline coronavirus (FCoV) in aqueous humor: 1) a 7b gene reverse-transcription real-time PCR (7b-RT-rtPCR) assay to detect FCoV RNA, 2) a spike gene mutation RT-rtPCR (S-RT-rtPCR) assay to detect 2 point mutations in the spike gene of FCoV in cats positive by 7b-RT-rtPCR, and 3) immunocytochemistry (ICC) for detection of FCoV antigen in aqueous humor macrophages. We studied 58 cats, including 31 cats with FIP and 27 control cats. FIP was excluded by postmortem examination and negative immunohistochemistry (IHC). Aqueous humor samples obtained postmortem were assessed using 7b-RT-rtPCR in all cats, and positive samples were evaluated with S-RT-rtPCR. ICC evaluation of aqueous humor samples from 36 of the 58 cats was done using an avidin–biotin complex method and monoclonal anti-FCoV IgG 2A. Sensitivity, specificity, and negative and positive predictive values were calculated including 95% CIs. 7b-RT-rtPCR had a specificity of 100.0% (95% CI: 87.2–100.0) and sensitivity of 35.5% (95% CI: 19.2–54.6). Specificity of S-RT-rtPCR could not be determined because there were no FCoV 7b-RT-rtPCR–positive samples in the control group. Sensitivity of S-RT-rtPCR was 12.9% (95% CI 3.6–29.8). Sensitivity and specificity of ICC were 62.5% (95% CI: 40.6–81.2) and 80.0% (95% CI: 44.4–97.5), respectively. The combination of 7b-RT-rtPCR and IHC could be useful in diagnosing FIP; S-RT-rtPCR did not add value; and ICC of aqueous humor samples cannot be recommended for the diagnosis of FIP.
Keywords
Introduction
Feline infectious peritonitis (FIP) affects cats worldwide and is a fatal disease if left untreated. Although feline coronavirus (FCoV; Alphacoronavirus 1) is ubiquitous, it rarely causes disease. It is thought that mutation of the virus in infected cats leads to the development of FIP. 34 A study conducted in 2012 detected 2 nucleotide mutations in positions 23531 and 23537 of the FCoV spike (S) gene, leading to amino acid substitutions (methionine to leucine at position 1058; serine to alanine at position 1060) in the spike protein that correlated with FIP in >95% of cases. 6 However, the amino acid substitution M1058L has also been detected in tissues of cats with systemic FCoV infection but no signs of FIP.3,30 Therefore, it is still a matter of discussion whether these mutations are responsible for the development of FIP or are instead an indicator of systemic spread of FCoV. 30 Other mutations that might play a role in the pathogenesis of FIP have been discussed, such as substitutions in the furin cleavage site between receptor-binding (S1) and fusion (S2) domains of the spike protein, supporting systemic spread of FCoV. 23
Diagnosing FIP antemortem is challenging, especially in cats without effusion. 1 A definitive diagnosis can only be achieved by immunohistochemistry (IHC) for detection of FCoV antigen within lesions.2,33 However, false-negative IHC results can occur because of non-uniform distribution of lesions within tissues and non-uniform distribution of virus within lesions. 27
Therefore, a reliable test is needed for the diagnosis of FIP. Detection of FCoV RNA using reverse transcription PCR (RT-PCR) in effusions of cats with FIP and controls revealed a specificity of 100%, making a positive result highly indicative of FIP. 12 In cats without effusion, whole blood, serum, or plasma samples are commonly tested for viral RNA, but the results are inconsistent.8,9,17,31 In addition, healthy cats can have positive results in RT-PCR of blood samples.4,15 Thus, blood is not considered a good substrate for diagnosing FIP in cats.
RT-PCR of aqueous humor has not been used routinely for the diagnosis of FIP, even though ocular involvement, manifesting mainly as uveitis and chorioretinitis,13,32 occurs in ~29% of cats with confirmed FIP. 20 If aqueous humor is collected during a diagnostic work-up, detection of FCoV could be added. The aim of our study was to determine whether testing samples of aqueous humor for FCoV could be used in the diagnosis of FIP. The tests included 1) a 7b gene reverse-transcription real-time PCR (7b-RT-rtPCR) to detect FCoV RNA in aqueous humor, 2) a S gene mutation RT-rtPCR (S-RT-rtPCR) to detect mutated FCoV, and 3) immunocytochemistry (ICC) to detect FCoV antigen in aqueous humor macrophages.
Materials and methods
Cats
We enrolled 58 cats in our study, including 31 cats with FIP and 27 control cats with other diseases. Twenty-five of the 31 cats with FIP and 11 of the 27 control cats were part of a previous study. 10 All cats had naturally acquired diseases and were client owned. Our study was approved by the ethics committee of the Centre for Clinical Veterinary Medicine of the Ludwig-Maximilians-Universitaet Munich (reference 52-27-07-2015).
Of the 58 cats, 5 had uveitis based on postmortem examination (4 in the FIP group, 1 in the control group); in the other cats, no ocular changes were noticed at clinical or postmortem examination. A diagnosis of FIP was based on the results of postmortem and histologic examinations as well as IHC of various organs including ocular tissue in all cats with uveitis. The procedure for IHC has been described previously. 11 Only cats with macroscopic and histologic lesions typical of FIP and positive IHC staining of affected tissue samples were included in the FIP group (Table 1).
Presence of uveitis and body cavity effusion and the results of 7b-RT-rtPCR for detection of FCoV, results of S-RT-rtPCR, and results of ICC in aqueous humor of 31 cats with FIP. In all cats, FIP was confirmed by histologic evaluation and positive IHC results.
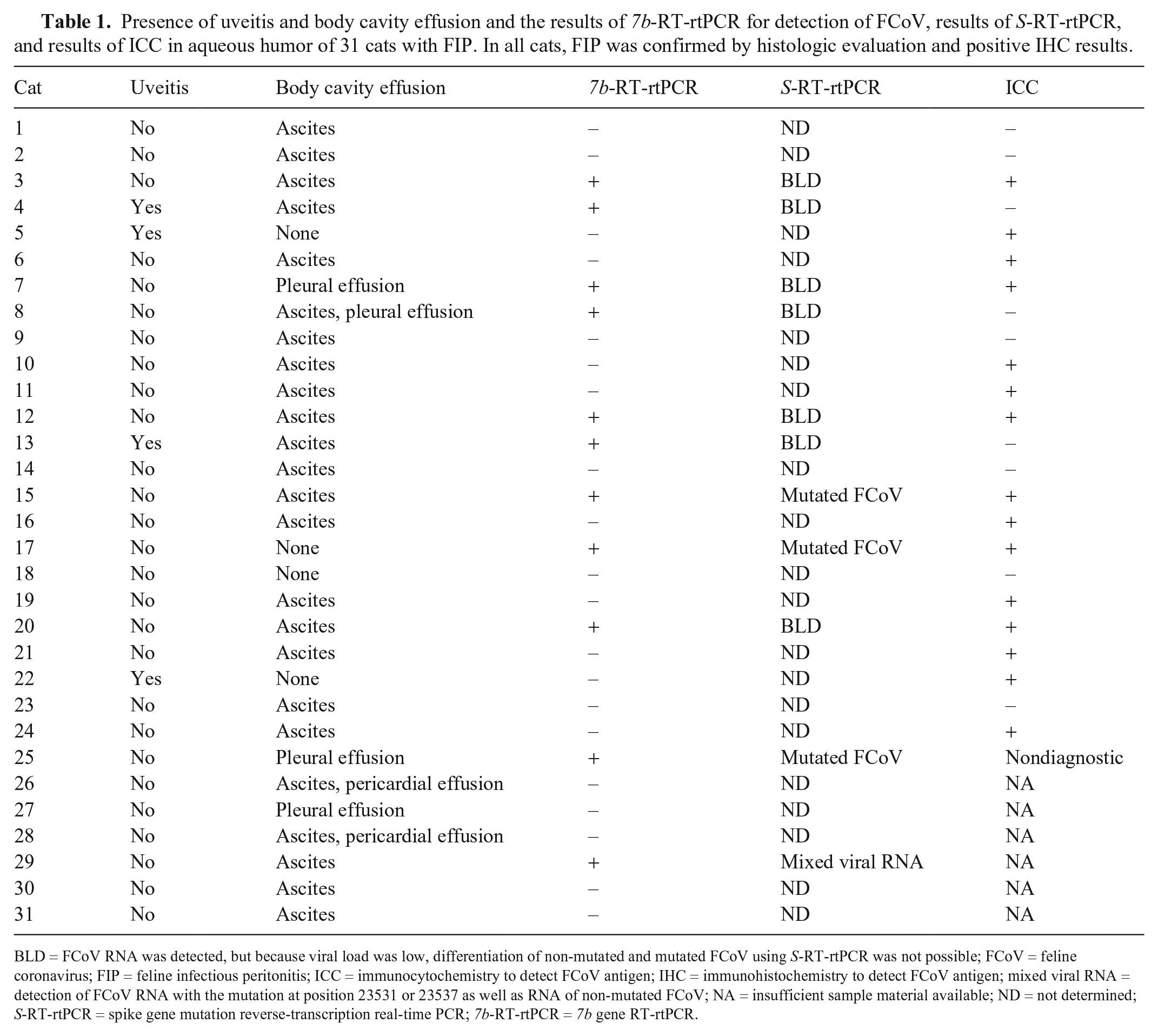
BLD = FCoV RNA was detected, but because viral load was low, differentiation of non-mutated and mutated FCoV using S-RT-rtPCR was not possible; FCoV = feline coronavirus; FIP = feline infectious peritonitis; ICC = immunocytochemistry to detect FCoV antigen; IHC = immunohistochemistry to detect FCoV antigen; mixed viral RNA = detection of FCoV RNA with the mutation at position 23531 or 23537 as well as RNA of non-mutated FCoV; NA = insufficient sample material available; ND = not determined; S-RT-rtPCR = spike gene mutation reverse-transcription real-time PCR; 7b-RT-rtPCR = 7b gene RT-rtPCR.
In all control cats, a postmortem examination, which included macroscopic and histologic examinations, was carried out to exclude FIP as the cause of death. Immunohistochemical staining of tissue samples showing histopathologic changes was negative for FCoV in all cats (Table 2). The individuals performing the laboratory tests were unaware of the underlying diagnoses.
Diagnosis, presence of uveitis, results of 7b-RT-rtPCR for detection of FCoV, results of S-RT-rtPCR, and results of ICC in aqueous humor of 27 control cats. In all cats, FIP was excluded by histopathology and negative IHC results.

See Table 1 notes for definitions of abbreviations.
Samples
Samples of aqueous humor were obtained postmortem via anterior chamber paracentesis using a 22-ga needle and a 2-mL syringe. All eyes were sampled by inserting the needle at the temporal limbus and positioning it parallel to the iris before advancing it into the anterior chamber for aspiration of aqueous humor. For 7b-RT-rtPCR and S-RT-rtPCR, samples of aqueous humor from both eyes of each cat were mixed and stored immediately after collection in cryogenic vials at –80°C. To determine the presence of FCoV RNA and to detect possible S gene mutations, 3 rtPCR assays were done (Idexx Laboratories). For ICC, 100 µL of aqueous humor was cytocentrifuged (Cytospin centrifuge Universal R; Hettich), and slides were stored at –20°C until analyzed. One slide was prepared for each cat.
7b-RT-rtPCR
Total nucleic acid was extracted from aqueous humor samples (QIAamp DNA blood BioRobot MDx kit; Qiagen) according to the manufacturer’s instructions with slight modifications (Idexx Laboratories). RT-rtPCR to detect FCoV was performed based on the occurrence of the 7b gene. 16
The 7b-RT-rtPCR was run with 5 quality controls that included: 1) PCR-positive controls (quantitatively; using synthetic DNA covering the RT-PCR target region [Integrated DNA Technologies]); 2) PCR-negative controls (PCR-grade, nuclease-free water); 3) negative extraction controls (extraction positions filled with nucleic acid–free water only); 4) a swab-based environmental contamination monitoring control; and 5) spike-in internal positive control (using lambda phage DNA). These controls assessed the functionality of the RT-PCR test protocols (control 1), for the absence of contamination in the reagents (2) and laboratory (4), absence of cross-contamination during the extraction process (3), and absence of RT-PCR inhibitory substances as a carryover from the sample matrix (5). The analytical as well as the clinical specificity of the 7b-RT-rtPCR was confirmed by resequencing amplification products with dedicated sequencing primers located 5’ and 3’ to those used in the routine RT-PCR protocol.
S-RT-rtPCR
All samples positive by 7b-RT-rtPCR were evaluated by the S-RT-rtPCR to discriminate between non-mutated and mutated virus, with 2 RT-rtPCRs carried out targeting the M1058L and S1060A single-nucleotide variations (SNVs) within the fusion peptide of the spike protein (Idexx Laboratories, unpublished data). The S-RT-rtPCRs allow the typing of an FCoV strain based on the presence (mutated FCoV) or absence (non-mutated) of 1 of 2 SNVs within the fusion peptide of the S gene. The paired S-RT-rtPCRs were previously validated analytically using synthetic DNA positive controls (IDT DNA Technologies) as well as clinically using samples collected from cats originally used to identify the 2 S gene mutations: cats 1) infected with FCoV and shedding but otherwise healthy, and 2) affected by FIP. 18 Briefly, highly specific hydrolysis probes were used to detect mutated and non-mutated sequences. The mutation detected at position 3174 (A→C/T) or 3180 (T→G) on the FCoV genome corresponded to amino acid positions 1058 and 1060; nucleotide 23531 and 23537; and M1058L and S1060A of reference sequence FJ938051, respectively. The non-mutated sequences were identified using an allelic discrimination approach (Idexx Laboratories, unpublished data). Probes for mutated and non-mutated S gene sequences were fluorophore-labeled (6-FAM and VIC, respectively). Results were analyzed to determine the 6-FAM:VIC (FIPV:FECV) fluorescence ratio emitted by the hydrolysis probes. S-RT-rtPCR was considered positive for either mutation when fluorescence in the mutation probe was at least 2-fold higher than in the non-mutated probe. S-RT-rtPCR was considered negative when 1) no FCoV was detected (negative), 2) FCoV without 1 of the 2 S gene mutations was detected (non-mutated), 3) FCoV load was below the cutoff of 1.5 million RNA equivalents per mL, which did not allow successful differentiation of the FCoV strains via S-RT-rtPCR (below limit of determination, BLD), or 4) no further differentiation via S-RT-rtPCR was possible despite a high FCoV load (>1.5 million RNA equivalents per mL of sample; indeterminate, IND). S-RT-rtPCR was considered positive when 1) FCoV with a mutated S gene (either mutation in nucleotide 23531 or 23537), or 2) both mutated and non-mutated S genes were detected in the same sample (mixed viral RNA; Table 3).
Results of ICC, 7b-RT-rtPCR, and S-RT-rtPCR in aqueous humor samples.
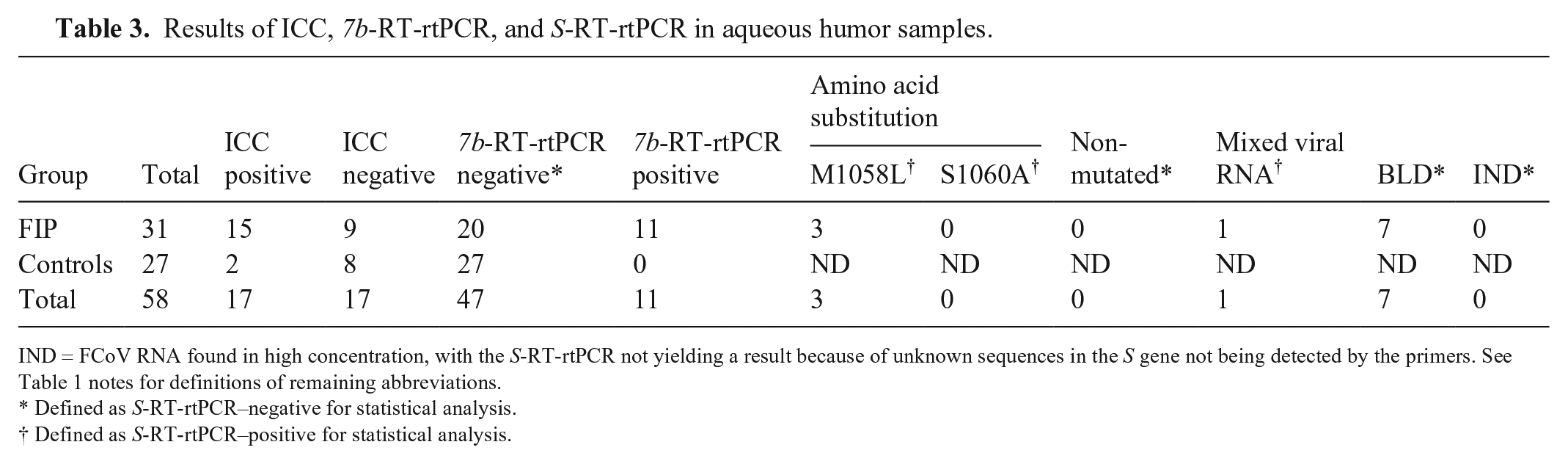
IND = FCoV RNA found in high concentration, with the S-RT-rtPCR not yielding a result because of unknown sequences in the S gene not being detected by the primers. See Table 1 notes for definitions of remaining abbreviations.
Defined as S-RT-rtPCR–negative for statistical analysis.
Defined as S-RT-rtPCR–positive for statistical analysis.
Immunocytochemistry
ICC was carried out in 25 samples from 31 cats with FIP and in 11 samples from 27 controls, as described previously. 11 In the remaining 22 cats, the yield of material was insufficient to carry out ICC. Stained slides were evaluated for cellularity and positive staining using light microscopy; only samples containing >50 nucleated cells were included in the analysis. Samples were considered positive when macrophages had dark brown granulated cytoplasm, and negative when the cytoplasm of macrophages was not dark brown or there were cells with nonspecific immunostaining, which included lymphocytes, neutrophils, erythrocytes, and plasma cells. Samples with a lack of cellular material on the slides were categorized as nondiagnostic and excluded from statistical analysis.
Statistical evaluation
Sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV), overall accuracy, and 95% CIs were determined (Table 4). When evaluating S-RT-rtPCR results, samples revealing BLD or IND were considered negative; samples containing mixed viral RNA were considered positive. Microsoft Excel and an online program calculating a non-asymptotic binomial CI (https://stattools.crab.org/Calculators/binomialConfidence.htm) were used.
Sensitivity, specificity, NPV, PPV, and overall accuracy of different methods to diagnose FIP and prevalence of FIP. Sensitivity of S-RT-rtPCR was calculated for all 31 cats with FIP. Two ICC samples were excluded from statistical analysis.
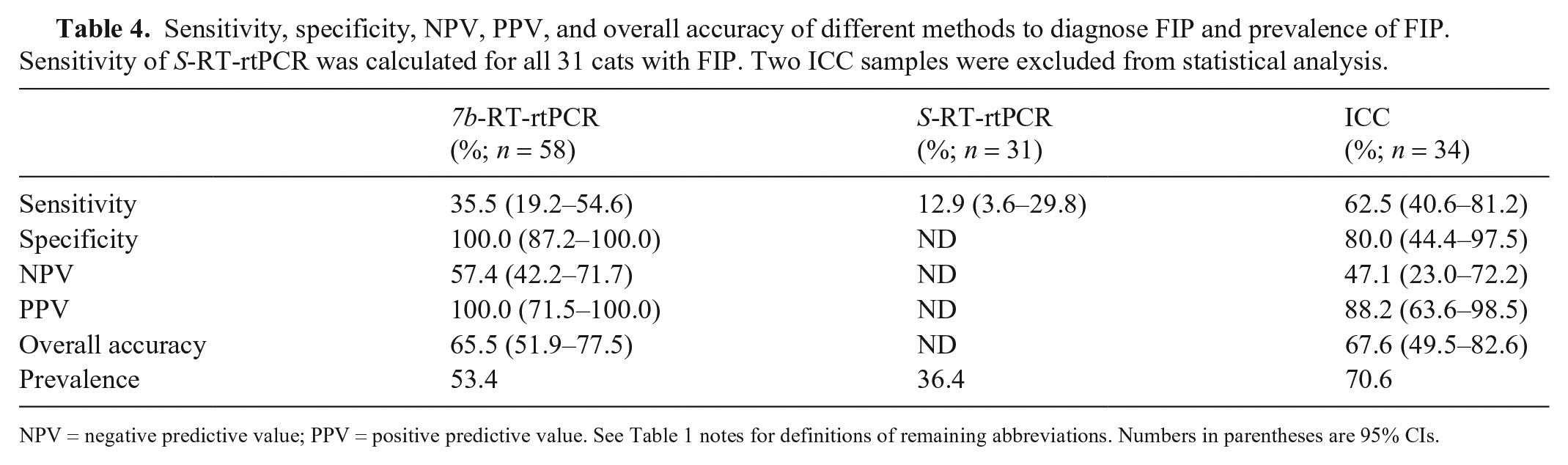
NPV = negative predictive value; PPV = positive predictive value. See Table 1 notes for definitions of remaining abbreviations. Numbers in parentheses are 95% CIs.
Results
All 27 samples from control cats were negative for FCoV RNA (Tables 2, 3). In cats with FIP, FCoV RNA was detected in 11 of 31 aqueous humor samples using 7b-RT-rtPCR. Of the 11 positive samples, 3 had a mutation corresponding to the amino acid substitution M1058L (Table 3) in S-RT-rtPCR, and in one sample, mixed viral RNA was detected (Table 1). In the remaining seven 7b-RT-rtPCR–positive samples, S-RT-rtPCR could not be performed because of low viral load. The specificity of S-RT-rtPCR could not be determined because none of the samples from control cats were positive by 7b-RT-rtPCR.
Thirty-six samples (25 FIP, 11 controls) were available for ICC. In cats with FIP, 15 samples were positive, and in control cats, 2 were positive (Tables 1–3). In 2 control cats, the ICC result was unclear because of dark brown immunostaining in macrophages; these were considered negative given that they lacked the typical granular pattern seen in positive ICC staining. Two samples (1 in a cat with FIP, 1 in a control cat) were excluded from statistical analysis because of the lack of cellular material on the slides.
Four cats with FIP and 1 control cat had ocular signs, which included lymphoplasmacellular endophthalmitis, keratitis, iridocyclitis, uveitis, and retinitis (Table 5). Of the 4 cats with FIP, IHC of ocular tissue was positive in 3 cases and negative in 1; the latter cat was diagnosed with FIP based on positive results of IHC of organ tissue samples.
Results of histologic evaluation, IHC of ocular tissue, 7b-RT-rtPCR, S-RT-rtPCR, and ICC of aqueous humor in cats with uveitis and FIP, as well as control cats with uveitis.
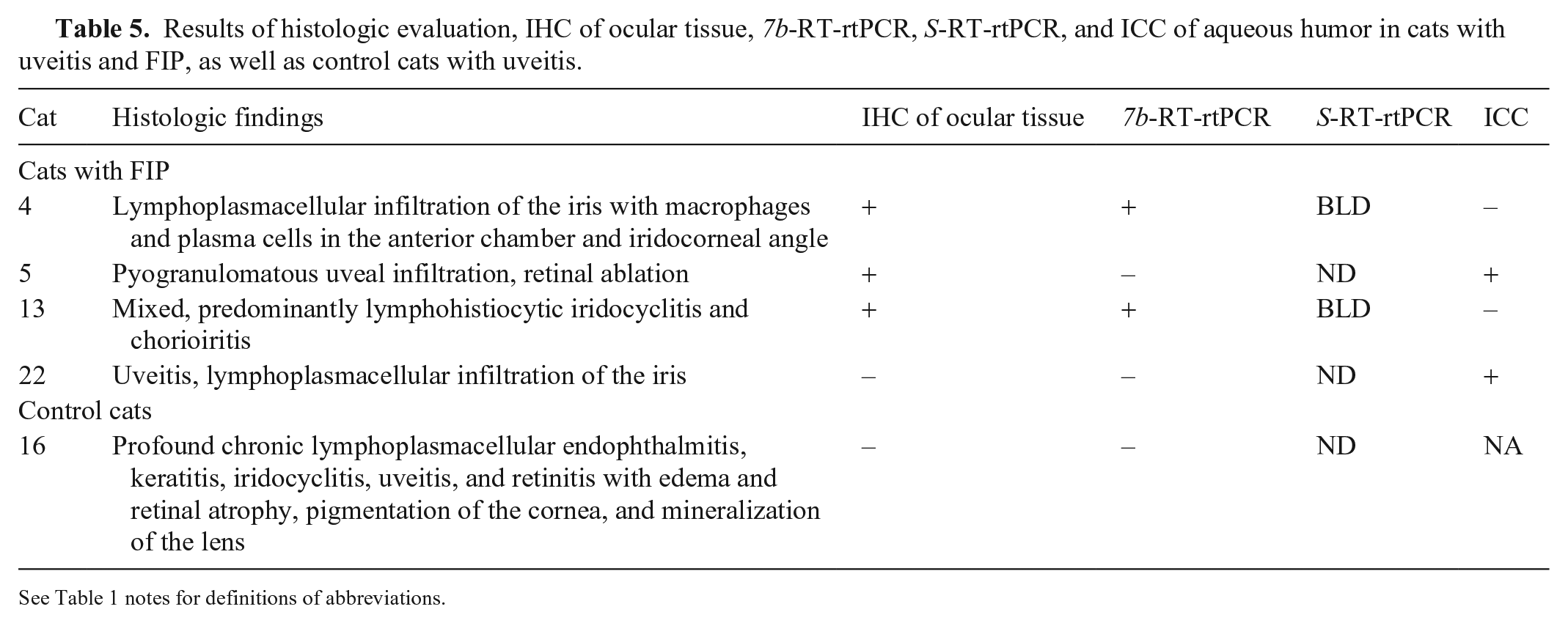

See Table 1 notes for definitions of abbreviations.
Discussion
Specificity of 7b-RT-rtPCR in aqueous humor was 100% (95% CI: 87.2–100.0). Although promising results have been achieved using the nucleoside analog GS-441524,26,29 as well as the protease inhibitor GC376, 28 for treatment of cats with FIP, a commercial drug is not yet available. Therefore, specificity of a test for FIP remains more important than sensitivity because misdiagnosis would likely lead to euthanasia of many cats. The high specificity of 7b-RT-rtPCR in aqueous humor was unexpected because FCoV is widespread in the cat population and FCoV RNA can be detected in blood and tissue samples of cats that do not have FIP.3,4,21,30
The absence of FCoV RNA in aqueous humor samples of the control group might be explained because the cats were not infected with FCoV or an intact blood-aqueous barrier prevented spillover of infected macrophages from the blood. The role of the blood-brain barrier (BBB) for FCoV entry into cerebrospinal fluid (CSF) has been discussed. 7 FCoV RNA was detected in CSF of cats with FIP indicating that macrophage-bound FCoV is able to enter the CSF via an impaired BBB. 7 Further studies showed positive immunostaining of macrophages in CSF in cats with FIP.14,19 However, because only one of the control cats had ocular involvement, absence of FCoV infection and/or an intact blood-aqueous barrier are the most likely reasons for the negative results.
The sensitivity of 7b-RT-rtPCR in aqueous humor was 35.5% (95% CI: 19.2–54.6). The relatively low sensitivity in cats with FIP might have been because only a small number of cats had signs of uveitis. Previous studies have used PCR to detect other pathogens in samples of aqueous humor. Felid herpesvirus 1 (FHV-1) DNA was detected by PCR in 11 cats with uveitis, 25 and Toxoplasma gondii DNA was found in 8.7% of healthy cats and 18.6% of cats with uveitis 22 ; hence, viruses and protozoa are more likely to enter the aqueous humor when infection leads to a compromised blood-aqueous barrier. Interestingly, only 2 of 4 cats with FIP and uveitis in our study had positive 7b-RT-rtPCR results. The results of IHC of ocular tissue were positive in 3 of the 4 cats with FIP.
Even though FCoV RNA was detected in aqueous humor in 11 of 31 cats with FIP, only 4 had 1 of the 2 S gene mutations. Thus, sensitivity of S-RT-rtPCR for the 31 cats with FIP was low (12.9%; 95% CI: 3.6–29.8). Three of the 4 cats had the S gene mutation leading to the amino acid substitution M1058L. The fourth sample was positive for non-mutated as well as mutated FCoV, indicating the existence of different FCoV strains within a cat. The samples of the other 7 cats contained FCoV in such low quantities that a differentiation was not possible using our assay.
ICC evaluation of aqueous humor revealed positive staining for FCoV antigen in 2 of 10 cats of the control group, which yielded a specificity of only 80.0% (95% CI: 44.4–97.5) for this technique. Of the 2 false-positive cats, one had pulmonary adenocarcinoma and the other lymphoma, and both had no ocular signs at macroscopic postmortem examination. However, cytologic evaluation of aqueous humor of these 2 control cats revealed a moderate number of inflammatory cells (Table 6). It is plausible that false-positive results were attributable to nonspecific positive staining of target proteins inside macrophages, 24 although the use of normal goat serum to block nonspecific antibody binding during ICC should have minimized this risk. In addition, both samples were positive for melanin, which might have been confused with the 3,3’-diaminobenzidine tetrahydrochloride (DAB) used in the ICC staining process. Another possibility is that these 2 cats were in an early subclinical stage of FIP because of immunosuppression associated with their primary illness. However, IHC of tissues with lesions was negative. It has been shown that healthy cats can harbor FCoV in tissue macrophages of various organs. 21 Thus, the positive results could have been staining of non-mutated virus, although this is unlikely given that 7b-RT-rtPCR in these 2 cats was negative.
Presence of uveitis and results of 7b-RT-rtPCR, ICC, cytology, and presence of protein in aqueous humor samples in 24 cats with FIP and 10 control cats in which ICC was performed (2 samples were excluded because of a lack of material on the slides).
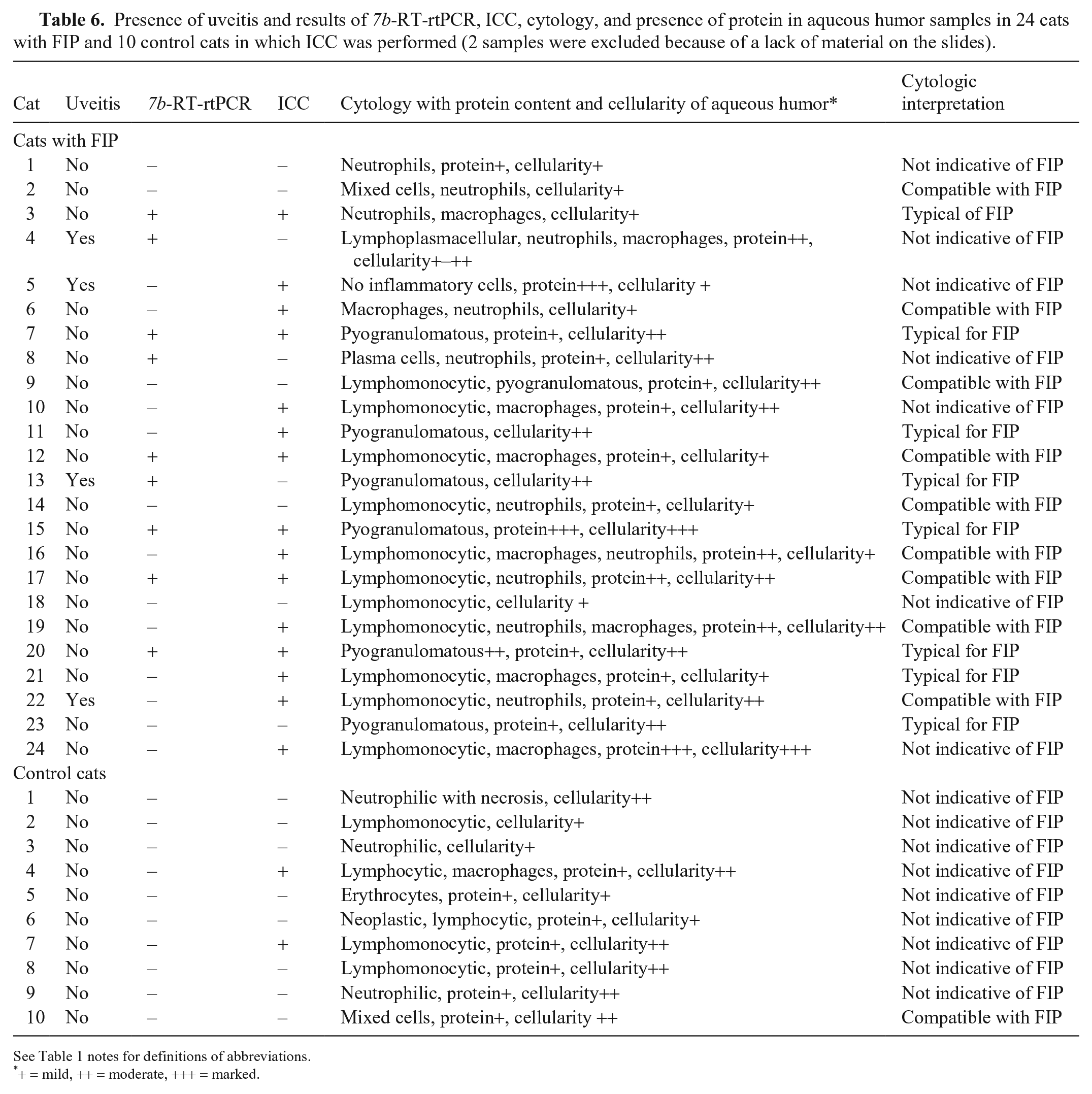
See Table 1 notes for definitions of abbreviations.
+ = mild, ++ = moderate, +++ = marked.
Sensitivity of ICC of aqueous humor was 62.5% (95% CI: 40.6–81.2) and thus higher than that of 7b-RT-rtPCR. The higher sensitivity was unexpected because ICC likely needs more viral particles than RT-rtPCR for a positive result. It is possible that intracellular FCoV antigen is better conserved, and viral RNA in aqueous humor is more easily destroyed during storage and transport. On the other hand, based on findings in control cats, false-positive ICC staining in cats with FIP might have contributed to spuriously high sensitivity.
Nine cats with FIP had false-negative results in ICC of aqueous humor samples. All 9 of these aqueous humor samples had signs of inflammation on cytologic evaluation, but FCoV antigen was not present, which indicates that perhaps the inflammatory process was not severe enough for the virus to cross the aqueous-blood barrier (Table 6). In dogs experimentally infected with canine infectious hepatitis virus, the absence of virus in aqueous humor coincided with the presence of specific antibodies at the time of iridocyclitis. 5 Thus, the false-negative ICC results might have been the result of local antibody production, which would bind and mask the antigen in ICC. Alternatively, antibodies entering aqueous humor via inflamed vessels could bind to existing antigen, leading to a failure of detection of the virus by ICC. To our knowledge, the presence of antibodies against FCoV in aqueous humor has not been assessed.
Our study has a few limitations. Samples were obtained postmortem, which might have led to a reduction in the amount of viral RNA and antigen because of degradation. Histologic evaluation and IHC of ocular tissue were done in only a few cases and not routinely in all cats. Finally, S-RT-rtPCR was not carried out on the samples of all cats. It is theoretically possible, albeit unlikely, that there was a sequence mutation at PCR primer sites, which would have resulted in samples being negative for 7b-RT-rtPCR but positive for S-RT-rtPCR. Such false-negative results in 7b-RT-rtPCR might have contributed to the low sensitivity of the S-RT-rtPCR.
Although 7b-RT-rtPCR was specific in the diagnosis of FIP in our study, it had a low sensitivity of 35.5%, and the use of S-RT-rtPCR did not lead to an increase in specificity. ICC had the best sensitivity (62.5%), but its specificity was unacceptably low. Only 5 of 58 cats had uveitis and, thus, the sensitivity of the laboratory tests might have been higher if only cats with uveitis had been evaluated. Based on the results of our study, the combination of 7b-RT-rtPCR and IHC could be useful in diagnosing FIP; S-RT-rtPCR did not add value, and ICC of aqueous humor samples cannot be recommended for the diagnosis of FIP.
Footnotes
Acknowledgements
We thank Karin Stingl, Institute of Veterinary Pathology, for her valuable technical assistance in performing the IHC.
Declaration of conflicting interests
C. Leutenegger was the Head of Molecular Diagnostics at Idexx Laboratories, at the time the study was performed. H.-J. Balzer and N. Pantchev are employed at Idexx Laboratories, Ludwigsburg; this laboratory offers the FCoV and FIP virus RT-rtPCR on a commercial basis and performed the testing in our study. Idexx played no role in the study design, in the collection and interpretation of data, or in the decision to submit the manuscript for publication. K. Hartmann has collaborated with and given talks for various companies, including Idexx Laboratories. There is no commercial conflict of interest given that the information generated here is solely for scientific dissemination. The authors declare that they have no competing interests.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
