Abstract
Clostridium perfringens type G is one of the pathogens involved in enteric diseases in poultry. NetB, a pore-forming toxin, is considered the main virulence factor responsible for necrotic enteritis during C. perfringens infection. We carried out a field study involving 14 farms to evaluate the occurrence of netB-positive C. perfringens and the impact of infection in Italian poultry flocks. Environmental samples (n = 117) and 50 carcasses were screened by microbiologic and molecular methods. Microbiologic investigations yielded 82 C. perfringens isolates. DNA was extracted from all samples and screened for α-toxin and NetB encoding genes by real-time PCR. The C. perfringens α-toxin gene was detected in 151 of 167 extracts (90.4%), and 31 of 151 (20.5%) were netB gene positive also. Sixteen isolates from a turkey flock with mild enteric disorders were also netB positive, demonstrating their occurrence not only in broiler but also in turkey flocks. A pulsed-field gel electrophoresis protocol was optimized to evaluate the diversity among isolates and revealed high genetic heterogeneity. The complete NetB toxin-coding gene of 2 C. perfringens isolates from turkey and broiler flocks were analyzed and showed very high relatedness with analogous sequences worldwide.
Keywords
Necrotic enteritis (NE) is an important disease, responsible for economic losses in the poultry industry. 20 Clostridium perfringens type G, a gram-positive, spore-forming, anaerobic microorganism, is the main causative agent of NE, although several predisposing factors play an important role in the pathogenesis of NE.19,21 In the past, α-toxin, a phospholipase C, was considered the main toxin responsible for NE, but this role has been questioned because other toxins have also been discovered to be involved.10,12 In particular, the NetB toxin, a pore-forming heptameric protein encoded on a plasmid, is required for the development of NE. 11 The prevalence of netB in poultry isolates has been evaluated in Sweden, Finland, and the United States, where the focus was on the genetic characterization of C. perfringens isolates circulating in broilers and turkeys by pulsed-field gel electrophoresis (PFGE), considered a useful tool to evaluate both relationships between genotypes and the epidemiology of C. perfringens infection.7,9,15,16
In Italy, an investigation carried out on 107 C. perfringens isolates recovered from broiler and commercial laying hen flocks revealed that 27% of isolates were netB positive, with 93% of these isolates coming from birds affected by intestinal disorders. 6 On the other hand, a similar study of 106 C. perfringens isolates from turkeys with enteric lesions failed to detect netB-positive isolates. 8
To verify the occurrence of netB-positive C. perfringens isolates under field conditions, we carried out an environmental survey in Italian broiler and turkey flocks from November 2015 until May 2017. Classical microbiologic methods, a real-time PCR (rtPCR) protocol, and PFGE analysis of the bacterial isolates were applied to litter samples and carcasses recovered during the whole production cycle.
The samples derived from 9 broiler and 5 fattening turkey flocks. In 6 broiler and 2 turkey flocks (flocks 1–6, 8, and 9) environmental samples were collected weekly by the boot sock method (Agritamp plus02; Biogenetics, Padova, Italy), performed by a single operator walking in every sector of the house (http://www.salute.gov.it/imgs/C_17_pubblicazioni_2453_allegato.pdf; Italian). For each flock, 3 separate sheds, with birds from 7 d old to near slaughter, were sampled. In addition, 15 carcasses were randomly obtained from healthy flocks 1 and 4–6. The remaining 6 flocks (broiler flocks 7, 10, and 12; turkey flocks 11, 13, and 14) were screened after veterinary notification of enteric disorders, and only carcasses were submitted for investigation. A total of 117 boot socks and 50 carcasses were collected.
Boot sock samples were diluted 1:10 w/v in sterile physiologic saline, and 1 mL of this suspension was inoculated into brain-heart infusion broth (1:10 v/v) and incubated at 37°C under anaerobic conditions for 24 h. Then, 0.1 mL from each tube was plated onto 5% sheep blood agar (SBA) plates that were incubated anaerobically for 24 h at 37°C. All carcasses, collected from both healthy and suspected infected flocks, were submitted for autopsy. The investigation focused mainly on the gastrointestinal tract (to determine the presence of typical necrotic gut lesions), and intestinal samples were selected for both microbiologic and histologic (hematoxylin–eosin and Gram staining) examinations. For microbiology, tissue fragments (5 cm proximal to and 3 cm distal to the Meckel diverticulum) 5 were incubated anaerobically at 37°C for 24 h in 10 mL of thioglycollate broth, and 0.1 mL of this broth was then inoculated onto SBA plates, as described previously. 3 The plates were incubated anaerobically for 24 h at 37°C.
All presumptive C. perfringens colonies were confirmed by Gram staining, biochemical identification (Vitek; Biomerieux, Craponne, France), and by rtPCR. In addition, for each sample (boot socks and intestines), DNA was extracted from 300 µL of inoculated broth and bacterial colonies (Maxwell 16 tissue DNA purification kit; Promega, Milan, Italy). The detection of cpa and netB was carried out by a specific rtPCR assay, originally developed for quantitative study and herein used as an end-point PCR. 2 This protocol targets specific DNA sequences (402 bp for cpa and 75 bp for netB), with a single thermal cycle program. The specificity of both amplicons was confirmed by SYBR Green–generated melting curves, using a reference C. perfringens strain (CP56) as positive control. 2 The rtPCR results were analyzed (CFX Manager; Bio-Rad, Segrate, Italy).
C. perfringens field isolates were analyzed by PFGE, following a previously described protocol with some modifications. 15 Bacterial suspensions were treated with 1.6 mg/mL of lysozyme and, subsequently, proteinase K (0.665 mg/mL). Then, 1% agarose (SeaKem Gold; Lonza, Basel, Switzerland), containing 1% sodium dodecyl sulfate (SDS), was added and the suspension dispensed into plug molds. The plugs were stored in 10 mL of TEN buffer (10 mM Tris pH 7.5; 1 mM EDTA; 50 mM NaCl) containing 0.14 mg/mL of proteinase K, at 4°C for 24 h, in order to completely lyse the bacterial cells. After double digestion with SmaI (Promega), each plug was included into 1% agarose gel (SeaKem Gold; Lonza) containing 2% thiourea, and electrophoresed (CHEF Mapper XA system; Bio-Rad). The resultant patterns were analyzed, and a dendrogram was produced (BioNumerics v.6.6; Applied Maths, Sint-Martens-Latem, Belgium) using a 2% Dice correlation and optimization coefficient. Isolates that shared >90% identity were considered as a pulsotype.
Finally, the netB gene of 2 different C. perfringens isolates, 1 isolated from a broiler and 1 from a turkey, were amplified by 3 different PCR assays, using 1 published primer 2 and 5 primers designed in our study (Table 1). The amplicons were purified and sequenced by the Sanger method (Genetic analyzer 3500; Applied Biosystems, Foster City, CA). Nucleotide sequences were visualized (CHROMAS; http://technelysium.com.au/wp/chromas/), and the sequences were compared with similar ones available in GenBank by BLASTn (http://www.ncbi.nlm.nih.gov/blast/Blast.cgi) to confirm the presence of the netB coding sequence. The phylogenetic analysis was performed employing MAFFT (Multiple alignment program for amino acid or nucleotide sequences). The alignment was manually edited to create the final phylogenetic tree (MEGA software v.7; http://www.megasoftware.net/). The sequences obtained from a turkey and chicken were deposited in GenBank (accessions KY923245 and MF535104, respectively).
Primers designed for netB gene sequencing.
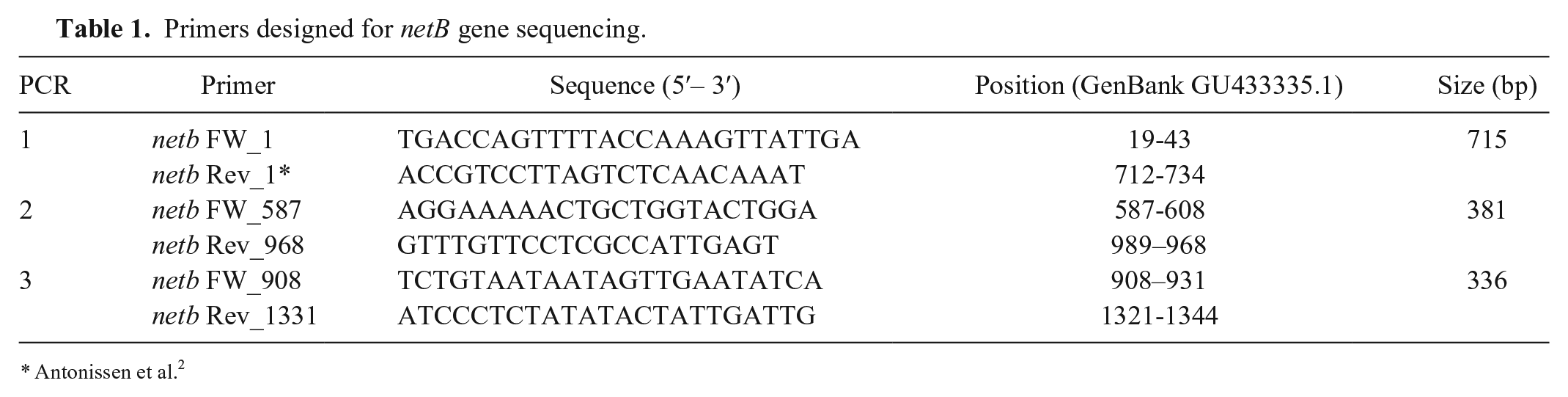
Antonissen et al. 2
We recovered 82 C. perfringens isolates from 7 of 9 broiler and 3 of 5 turkey farms (Table 2). The identity of the isolates was confirmed by rtPCR, and 22 isolates, from turkey flock 8 and broiler flock 10, were confirmed as netB positive. In flock 8, the birds had mild diarrhea and a moderate increase of mortality rate at 3 wk of age; netB-positive C. perfringens isolates were obtained, even if irregularly, in all 3 sampled sheds (from both boot socks and dead birds), from 24- to 115-d-old turkeys (Fig. 1). Broiler flock 10, in which increased mortality rate and enteric disorders were observed, was monitored once, after veterinary notification, and 45-d-old dead birds were sampled.
Isolation of Clostridium perfringens from boot socks and carcasses from broiler and turkey flocks.
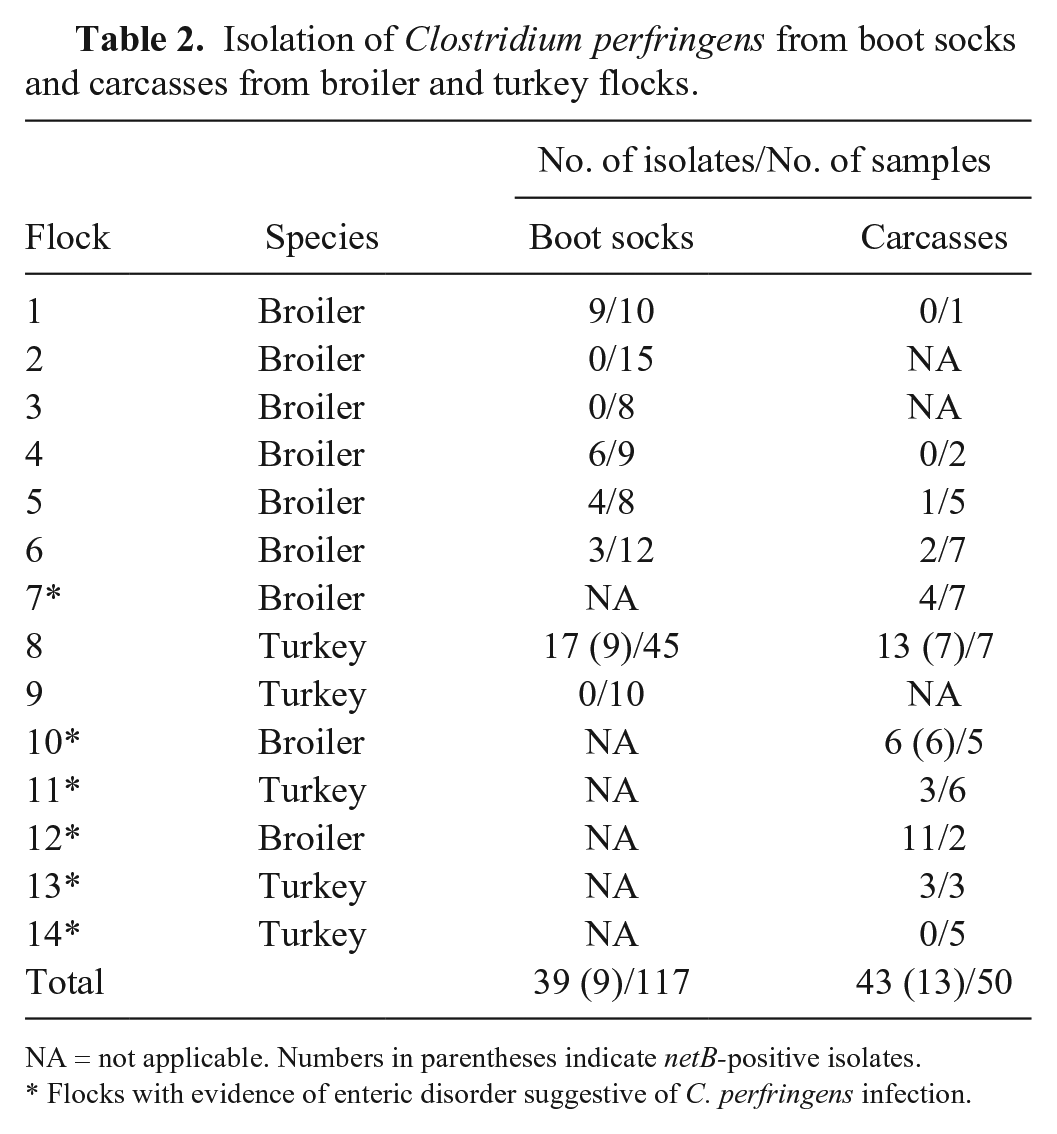
NA = not applicable. Numbers in parentheses indicate netB-positive isolates.
Flocks with evidence of enteric disorder suggestive of C. perfringens infection.
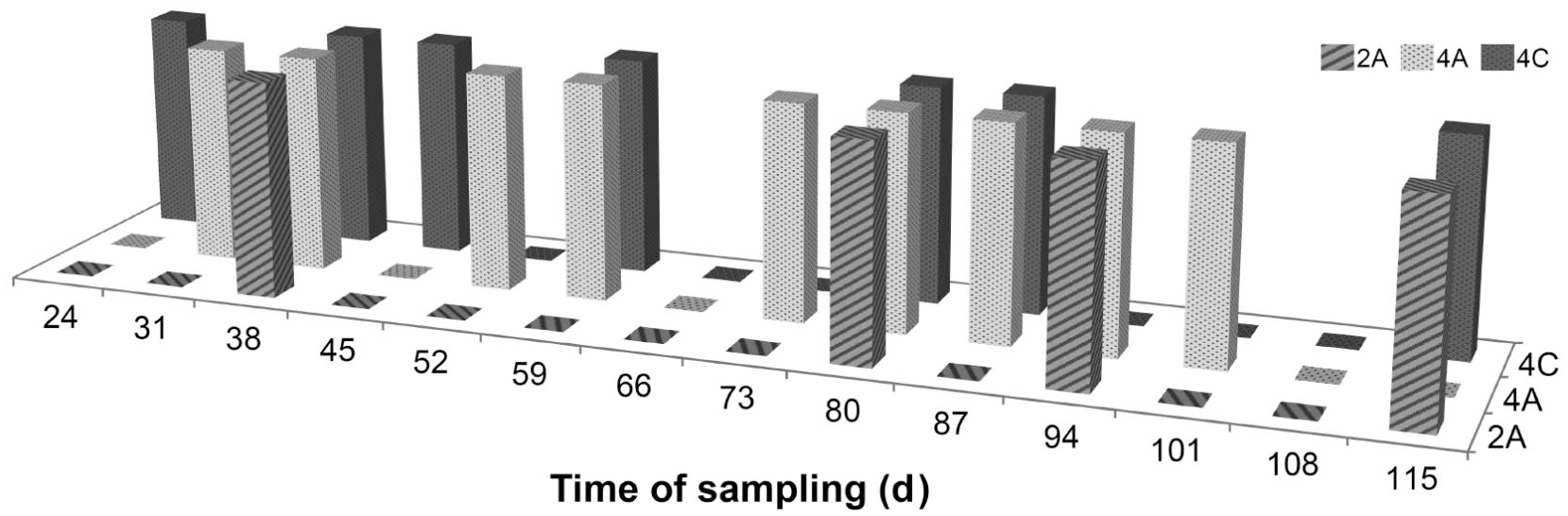
Detection of netB-positive Clostridium perfringens for each monitored shed (2A, 4A, 4C) during the weekly sampling of turkey flock 8. Time of sampling is age (d) of the birds.
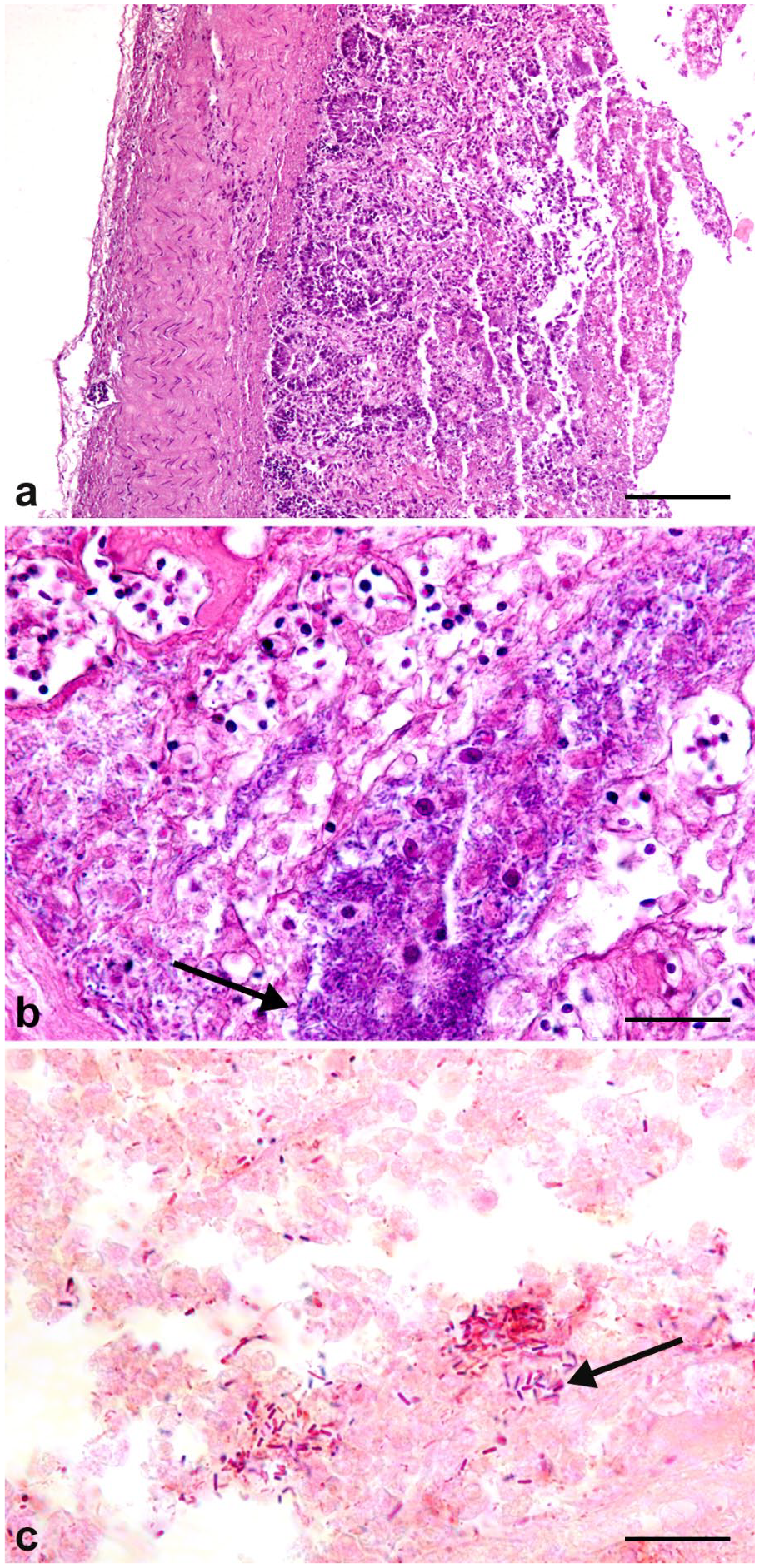
Of 50 examined carcasses, 7 turkeys (2 from flock 11, 3 from flock 13, and 2 from flock 14) and 1 broiler (flock 12) had necrotic lesions, with typical pseudomembranes. The intestinal lumen was filled with yellow liquid that contained necrotic material detached from the mucosal surface. Gross lesions were in the small intestine, especially in the duodenum (wall was thickened and covered by extensive yellow-brown pseudomembranes). Histology confirmed the occurrence of severe necrosis of the mucosa. In addition, cellular debris, fibrinous exudate, and gram-positive bacillus-like aggregates were found (Fig. 2a–c).
The rtPCR screening, carried out on all 167 samples, revealed cpa-positive C. perfringens in 151 extracts and, among them, 31 (20.5%) also carried the netB gene (Table 3). The netB-positive extracts came from flocks 8 (turkey) and 10 (broiler), in which only mild clinical signs of enteric disorder were noted, but no typical necrotic lesions were observed. Curiously, C. perfringens isolates recovered from carcasses with severe intestinal lesions suggestive of necrotic enteritis (flocks 11–14) were netB negative.
Detection of the Clostridium perfringens cpa gene in boot socks and carcasses for broiler and turkey flocks.
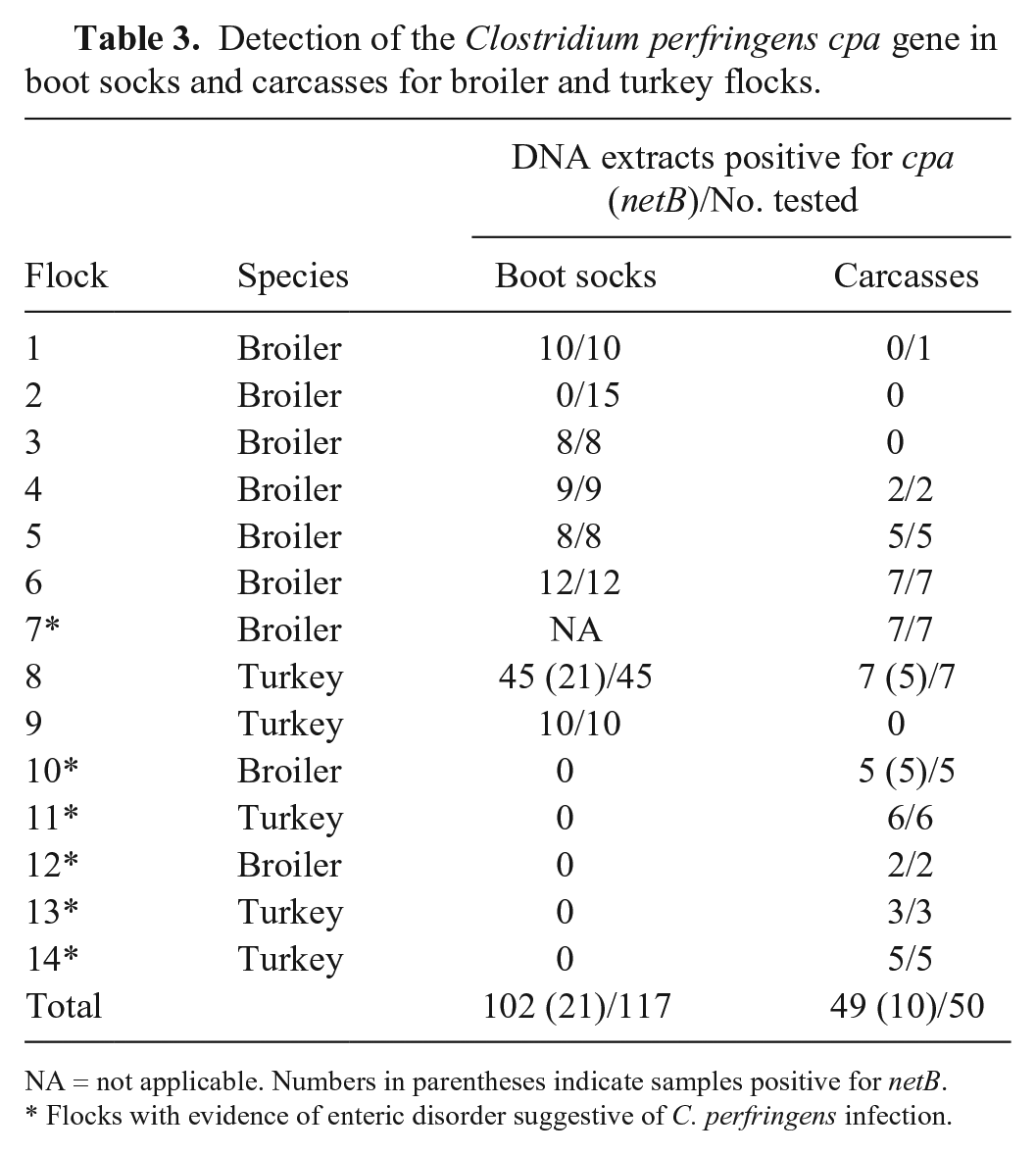
NA = not applicable. Numbers in parentheses indicate samples positive for netB.
Flocks with evidence of enteric disorder suggestive of C. perfringens infection.
The PFGE analysis returned good quality images, suitable for BioNumerics analysis, for 45 C. perfringens isolates. A total of 32 PFGE genotypes were identified, and the dendrogram showed that 6 of them included isolates reaching 100% identity and coming from flocks 1, 5, 6, 8, and 10 (Fig. 3). The netB-positive C. perfringens from turkey flock 8 seemed to be widely distributed throughout the dendrogram, clustering in 3 different groups (groups A–C in Fig. 3) with identity values of 80.8–93.3%. The netB-positive isolates, belonging to broiler flock 10, were recovered from 5 intestines and produced a cluster with an identity of 92.3% (group D in Fig. 3).
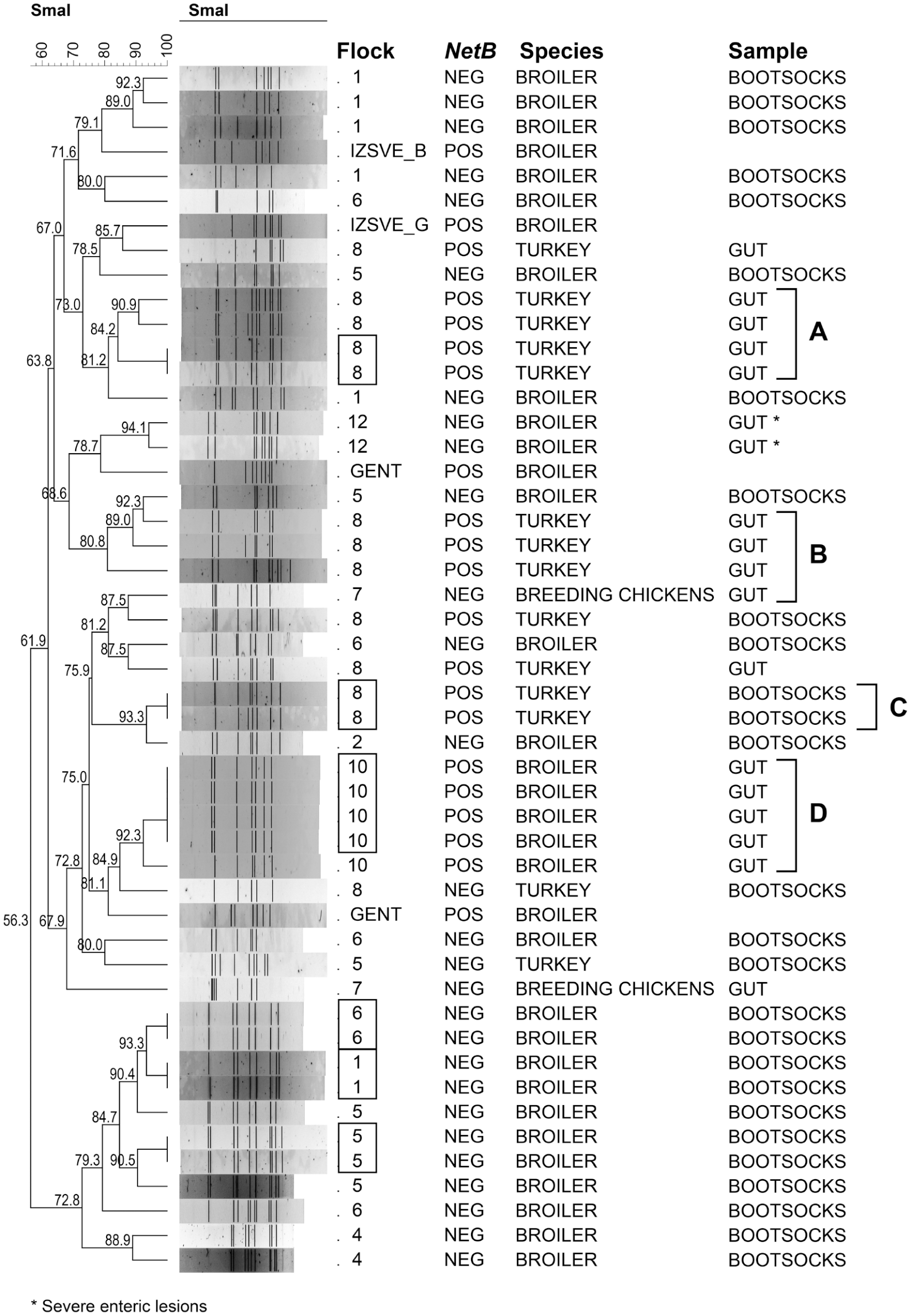
Dendrogram produced by BioNumerics software and generated after pulsed-field gel electrophoresis analysis of Clostridium perfringens isolates. POS = C. perfringens netB–positive isolate; NEG = C. perfringens netB–negative isolate. The dendrogram includes 4 C. perfringens strains, kindly provided by Prof. Filip Van Immerseel (Gent University, Belgium) and Dr. Luca Bano (Istituto Zooprofilattico Sperimentale delle Venezie, Italy) for comparison with the field isolates. The netB C. perfringens isolates, coming from turkey and broiler flocks, were grouped (groups A–D). Numbers within boxes indicate flocks with isolates reaching 100% identity.
Comparison of netB locus sequences revealed substantial identity of our 2 sequences, with an exception in the turkey sequence of a nucleotide variation (G/A) in position 509, leading to an alanine to threonine shift, a feature that has been described previously. 1 Moreover, the turkey sequence had a nucleotide deletion in the intergenic region downstream of the netB gene. Homolog searches returned 34 sequences that were used to build a multiple alignment, revealing an identity value of 99.8–100% between our sequences and other netB genes, limited to the coding region, mainly coming from Australia, Canada, Denmark, and some partial sequences from Egypt and Iran.
Phylogenetic inference, rooting the tree on the KC460332 sequence from an ostrich, highlighted the distribution of netB sequences in 2 main clusters, although poorly supported by bootstrap values (Fig. 4). Cluster A included NetB_Chicken_2016_ITA sequence from our study along with complete sequences from Australia, Canada, and Denmark, and partial sequences from Egypt and Iran isolates. Cluster B grouped the NetB_Turkey_2016_ITA from our study with sequences from isolates around the world.
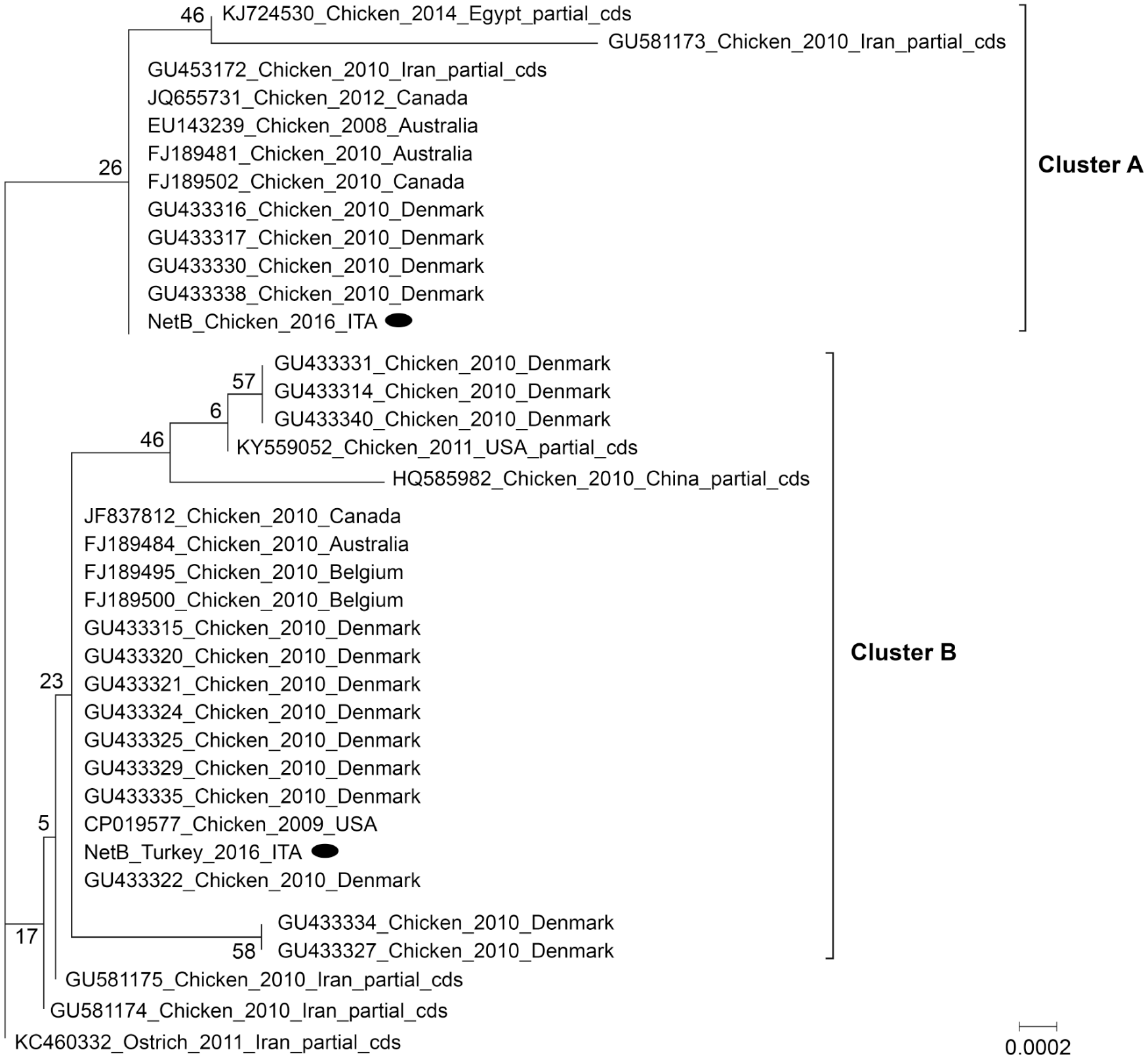
Phylogenetic tree based on the complete 960-bp netB gene comparison, including the 2 netB complete sequences obtained from broiler chickens and turkeys, named NetB_Turkey_ITA and NetB_Chicken_ITA, respectively. Maximum likelihood method, bootstrapping (1,000 replicates), with 40% site coverage cutoff.
Our study offers new insight regarding the prevalence of C. perfringens in Italian poultry flocks. As reported previously, 6 we confirmed the occurrence of netB-positive C. perfringens isolates in broiler flocks, with or without an enteric disorder, suggesting the need for additional environmental factors for disease development. Regarding turkey flocks, although we detected netB-positive C. perfringens isolates, the role of this organism in the pathogenesis of NE is still unclear.
Interestingly, all netB-positive isolates were from flocks with the subclinical form of NE, with a moderate increase of mortality rate but an absence of typical pathologic findings of enteritis. In contrast, when severe NE was observed during autopsy of deceased birds (flocks 11–15), no netB gene was detected. We cannot rule out that intense dysbiosis could affect the intestinal microbiota, resulting in a decrease of the netB-positive clostridia initially responsible for the infection. The role of additional toxins, such as beta2 or TpeL, cannot be ruled out, although their contribution to the pathogenesis of NE in poultry should be further investigated.4,13,14 A 2017 study demonstrated that C. perfringens isolates recovered from turkeys do not harbor the same pathogenicity loci associated with isolates associated with the disease in chickens. 18
The PFGE analysis showed a high genetic diversity among C. perfringens isolates, all coming from flocks in which severe NE was not reported. These results are consistent with previous reports in which mild clinical cases of NE were observed.7,9,16 However, although many different C. perfringens isolates were fingerprinted, our analysis revealed that most of the netB-positive isolates, coming from flocks 8 and 10, produced distinct PFGE patterns within the dendrogram (groups A–D, Fig. 3).
The netB gene sequences obtained in our study were identical in the CDS region to each other, except for the G509A mutation, and highly similar to analogous netB sequences already available in GenBank.
No netB genes from turkey isolates of C. perfringens have been deposited prior to our study. It has been reported that both nucleotide and amino acid sequences of several netB-positive isolates appeared to all be highly conserved, and only one amino acid change, A168T, was observed in a few isolates, but no difference in cytotoxicity was shown. 12 This mutation, at CDS position 509, was present in the turkey sequence (KY923245) obtained in our study; the broiler netB sequence (MF535104) from our study did not show any difference. However, although our work confirmed strong conservation in the netB gene sequence, it could be useful to investigate any variation in those genes that could have an influence over netB gene expression and, therefore, in toxin production, as in the VirS-VirR system. 17
Overall, the laboratory protocol applied in our study, based on both classical and biomolecular methods, is a rapid and suitable tool for field investigations, and could contribute to better understanding of the role of C. perfringens in avian diseases.
Footnotes
Acknowledgements
We thank Filip Van Immerseel and his staff at the Department of Pathology, Bacteriology and Avian Diseases, Faculty of Veterinary Medicine, University of Gent, Belgium, and Luca Bano, Istituto Zooprofilattico Sperimentale delle Venezie, Italy, for kindly providing netB-positive C. perfringens strains.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
