Abstract
To date, there is no clear standard to monitor drug treatment for canine Chagas disease. We used 2 real-time PCR (rtPCR) assays targeting Trypanosoma cruzi kinetoplast DNA (kDNA) and nuclear satellite DNA (nDNA) to detect T. cruzi in canine whole blood. Samples were collected randomly from 131 untreated dogs with unknown T. cruzi infection status in Texas. The kDNA-based rtPCR was slightly more sensitive (diagnostic sensitivity of kDNA = 49% vs. nDNA = 44%; p = 0.5732) but slightly less specific (diagnostic specificity of kDNA = 96% vs. nDNA = 97%; p > 0.9999) than the nDNA-based rtPCR. However, the differences in sensitivity and specificity between the nDNA- and kDNA-based rtPCR assays were not statistically significant. Using the nDNA- and kDNA-based qualitative rtPCR assays to monitor parasitemia from 137 itraconazole- and amiodarone-treated cases with nDNA- and kDNA-based PCR–positive baselines showed that the PCR positive rate decreased to 0% in 30 d. Using kDNA-based quantitative rtPCR to monitor normalized T. cruzi DNA copies in 4 representative dogs demonstrated that drug treatment could reduce parasite loads within 7–30 d. The kDNA-based qualitative rtPCR may be used for routine parasitemia screening of drug-treated Chagas-positive dogs, whereas nDNA-based qualitative rtPCR may be used for confirmation.
Chagas disease (American trypanosomiasis) is caused by the protozoan Trypanosoma cruzi, whose transmission can occur not only through the feces of infected triatomine vectors, but also through blood transfusions, organ transplantations, oral ingestion of infected vectors, contaminated food, transplacental routes, lactation, or laboratory accidents.3,8,19 More than 100 species of mammals can be infected by T. cruzi via >150 species of triatomine bugs. 19 The disease affects 6–9 million people worldwide, and is endemic in South America, Central America, Mexico, and the United States. 11 Dogs are a key enzootic host for domestic T. cruzi maintenance. 11 The prevalence of natural T. cruzi infection in dogs varies in endemic areas, but it can be higher than that in infected humans. 8
Infection of dogs with T. cruzi causes nonspecific clinical signs such as depression, lethargy, exercise intolerance, and arrhythmias. 8 Dogs that enter the indeterminate or latent phase after the acute phase may be asymptomatic, although some progress to heart failure. 3 Diagnosis of Chagas disease is generally based on serologic and parasitologic detection, clinical findings, and the likelihood of being infected. Traditional methods to detect parasites such as hemoculture, xenoculture, and microscopic examination of blood smears may be replaced by molecular approaches such as PCR. T. cruzi PCR methods developed for human samples showed heterogeneity in test performance when used with dog samples.2,9,13 We evaluated 2 real-time PCR (rtPCR) targets: 1) T. cruzi nuclear satellite DNA (nDNA) located within a conserved region of the 195-bp nuclear repetitive element in the trypanosome chromosome, 7 and 2) T. cruzi minicircle kinetoplast DNA (kDNA), which is conserved and abundant within a single mitochondrion. 17 Using these PCR methods, we attempted to detect T. cruzi DNA in circulating blood to monitor parasitemia during drug treatment.
EDTA whole blood samples were randomly collected, with animal owner consent, from 186 untreated and 157 drug-treated (itraconazole 10 mg/kg and amiodarone 7.5 mg/kg orally taken daily for 365 d 14 ) dogs in veterinary clinics or hospitals in Texas between March 2015 and February 2018 (Fig. 1). Blood was collected from the jugular vein, transferred to EDTA vacutainer tubes, and kept at 4°C for up to 1 wk. DNA was extracted from 1 mL of each blood sample (Genomic DNA from blood kit; Clontech, Mountain View, CA), and 8% of the DNA yield was used for T. cruzi nDNA and kDNA detection by rtPCR as described15,16 using dog β-actin gene as the internal control. For qualitative rtPCR, cycle threshold (Ct) values <39 and <36 were considered positive for nDNA and kDNA, respectively. For absolute quantitative PCR (qPCR; standard curve method), circulating parasite load was expressed as the ratio of log10 (T. cruzi copy number)/log10 (β-actin copy number), in which T. cruzi copy number was normalized by the β-actin copy number. Statistical analysis was performed using EpiTools (http://epitools.ausvet.com.au). Concordance between the 2 assays was analyzed using the Cohen kappa statistic: 0.00–0.20 = poor agreement; 0.21–0.40 = fair agreement; 0.41–0.60 = moderate agreement; 0.61–0.80 = substantial agreement; and 0.81–1.00 = almost perfect agreement. 12 Discordance between the 2 PCR results was assessed using the McNemar test, and a statistically significant difference was defined as p ≤0.05.
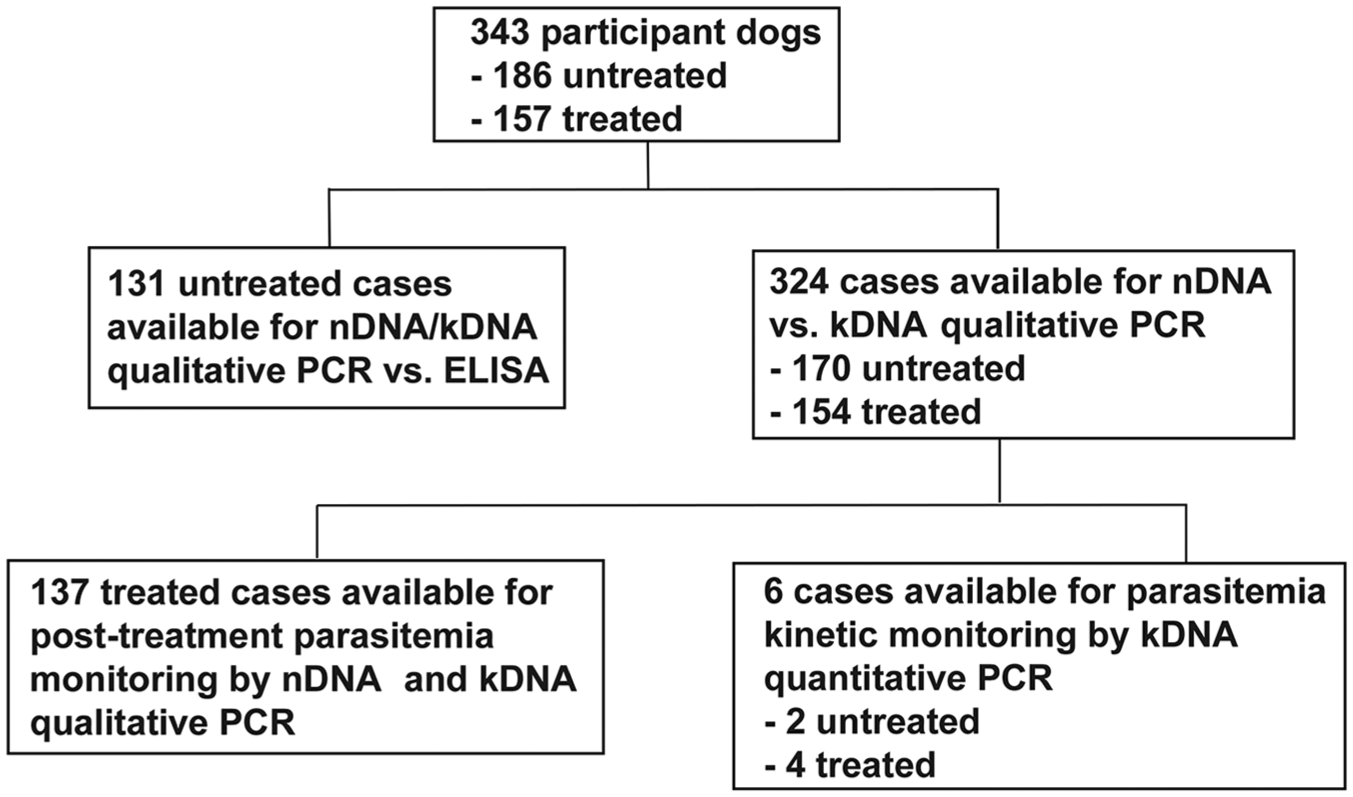
Flow diagram of participant dogs.
The analytical sensitivity of nDNA- and kDNA-based qualitative rtPCR was determined by testing serial dilutions of a positive control plasmid and cultured T. cruzi (TX Corpus Christi stocks, TcI, ATCC 30160). The lower detection limit for both nDNA- and kDNA-based PCR assays was 1 copy of plasmid and 1 parasite per PCR reaction. The dynamic ranges of both reactions were linear over 7 log units, spanning 101–107 copies of target genes.
To evaluate the diagnostic sensitivity (DSe) and specificity (DSp) of nDNA- and kDNA-based qualitative rtPCR, 131 dogs that had both DNA and serum samples available were randomly selected from 186 untreated cases with unknown T. cruzi infection status before PCR and serologic testing (Fig. 1). Anti–T. cruzi epimastigote IgG ELISA results were obtained as described previously 15 except that an anti-dog IgG peroxidase-labeled conjugate (Jackson ImmunoResearch Laboratories, West Grove, PA) was used. The kDNA-based qualitative rtPCR was slightly more sensitive (49% [28 of 57] DSe for kDNA rtPCR vs. 44% [25 of 57] for nDNA rtPCR; chi-square test: p = 0.5732) but slightly less specific (96% [71 of 74] DSp for kDNA rtPCR vs. 97% [72 of 74] for nDNA rtPCR; Fisher exact test: p > 0.9999) than the nDNA-based qualitative rtPCR (Tables 1, 2). However, the differences in DSe and DSp between nDNA- and kDNA-based rtPCRs were not statistically significant. The DSe of qualitative nDNA- and kDNA-based rtPCRs demonstrated that, within the ELISA-positive group (57 dogs), PCR-positive and -negative rates were 44–49% and 51–56%, respectively, suggesting that approximately half of these dogs were in the chronic phase, because parasitemia is usually high in the acute phase, but low or undetectable in the chronic phase. 3 The DSp of both nDNA- and kDNA-based rtPCRs was high (≥96%). Two PCR false-positive samples were consistently observed in the nDNA- and kDNA-based rtPCRs within the ELISA-negative group. We ruled out the possibility that these 2 dogs were in the acute phase before anti–T. cruzi IgG antibodies were generated because 1-mo follow-up ELISA and nDNA- and kDNA-based rtPCR results were all negative, indicating that these 2 dogs were not infected with T. cruzi. Another dog was also kDNA rtPCR–positive, but nDNA rtPCR–negative and ELISA-negative. Because follow-up samples were not available, these results are inconclusive. Periodic training of precautionary steps to lower the risk of contamination was implemented to minimize the occurrence of false-positive results.
Comparison of nDNA-based qualitative real-time PCR and ELISA results from untreated dogs with unknown Trypanosoma cruzi infection status.
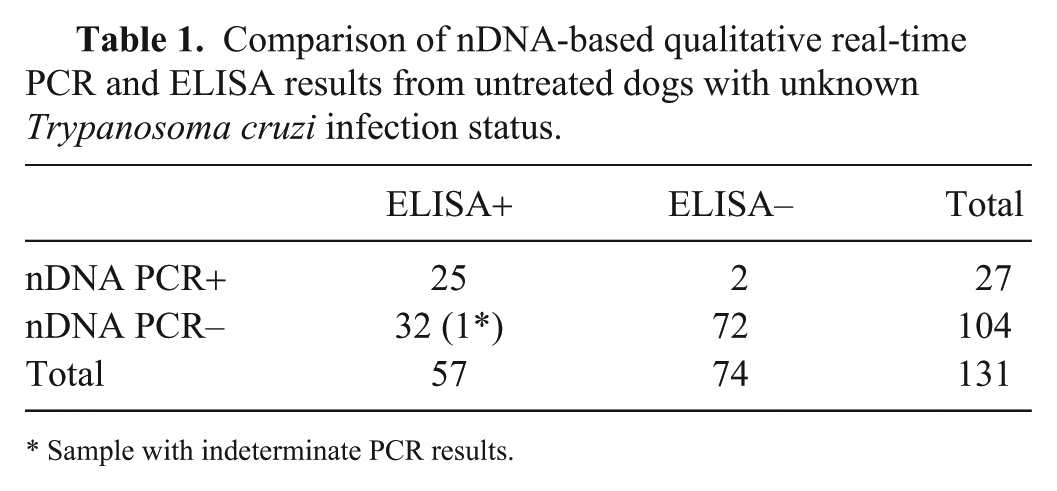
Sample with indeterminate PCR results.
Comparison of kDNA-based qualitative real-time PCR and ELISA results from untreated dogs with unknown Trypanosoma cruzi infection status.
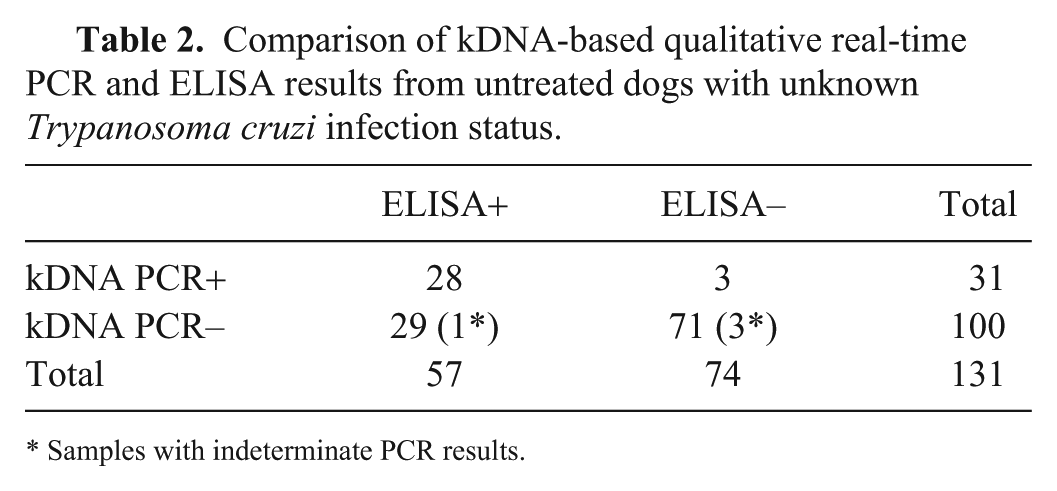
Samples with indeterminate PCR results.
In our nDNA- and kDNA-based qualitative rtPCRs, positive or negative results were based on the Ct cutoff value established, and no indeterminate results (including 1 nDNA- and 4 kDNA-based rtPCR indeterminate results; Tables 1, 2) were included for data analysis. However, clinicians may consider indeterminate results informative for Chagas disease diagnosis and treatment consideration; therefore, any nDNA rtPCR Ct values of 39–40 and kDNA rtPCR Ct values of 36–40 could be called indeterminate instead of negative. Indeterminate results could be false-positive or false-negative. Here, the indeterminate rate was 1% (1 of 104) for nDNA rtPCR and 4% (4 of 100) for kDNA rtPCR.
Comparing nDNA- and kDNA-based qualitative rtPCR results from testing 324 samples from untreated and treated dogs (Table 3), the proportion positive agreement rate was 96%, the proportion negative agreement rate was 99%, and the overall proportion agreement was 98% with Cohen κ = 0.95 (95% CI: 0.91–0.99), indicating almost perfect agreement between these 2 PCRs. However, the difference between the 2 PCR methods was significant (McNemar test: p = 0.041), which resulted from 6 of 324 samples tested with inconsistent kDNA- and nDNA-based rtPCR results.
Agreement between nDNA- and kDNA-based qualitative real-time PCR results from untreated and treated dogs.
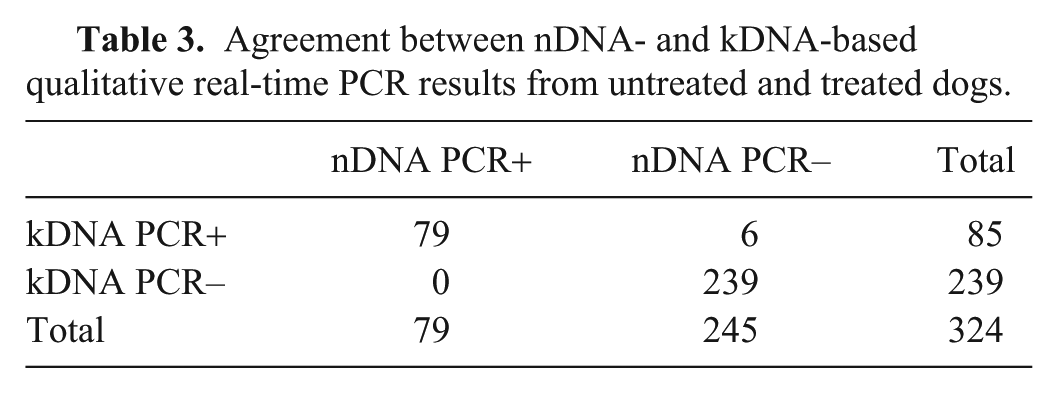
To apply nDNA- and kDNA-based qualitative rtPCR to detect parasitemia during itraconazole and amiodarone treatment, 137 blood samples with positive nDNA rtPCR and kDNA rtPCR baselines were collected within 1 y of treatment and divided into the following 5 groups (Table 4):
1) 1–7 d of treatment: 15 of 17 samples were positive using both nDNA and kDNA rtPCRs. Two samples collected at day 7 were negative in both rtPCRs.
2) 8–14 d of treatment: 1 sample collected at day 8 was positive. However, this dog became PCR-negative at day 16 in both nDNA and kDNA rtPCRs.
3) 15–30 d of treatment: only 1 of 42 samples had a very weak nDNA-positive rtPCR result at day 30. This dog’s serum had itraconazole levels in the optimal range of 1.5–3.0 μg/mL (measured using liquid chromatography–mass spectrometry at Texas A&M Veterinary Medical Diagnostic Laboratory, College Station, TX; unpublished data), but had unknown amiodarone levels, and thus the cause of its unexpected positive PCR is unclear.
4) 31–180 d of treatment: 1 sample collected at 3 mo was positive in both nDNA and kDNA rtPCRs. However, this dog had received drug treatment intermittently. Poor treatment compliance or re-infection might have caused relapse.
5) 181–365 d of treatment: 6 dogs were positive in both nDNA and kDNA rtPCRs. Two dogs were kDNA rtPCR–positive, but nDNA rtPCR–negative for samples collected at 8 mo. All 8 dogs had received drug treatment intermittently. Poor treatment compliance or re-infection might have caused relapse.
6) After completing the 1-y treatment regimen: 1 of 12 samples converted from nDNA and kDNA rtPCR–negative to strong positive at 12 mo after treatment.
Positive PCR detection rates of Trypanosoma cruzi during drug treatment of Chagas- positive dogs.
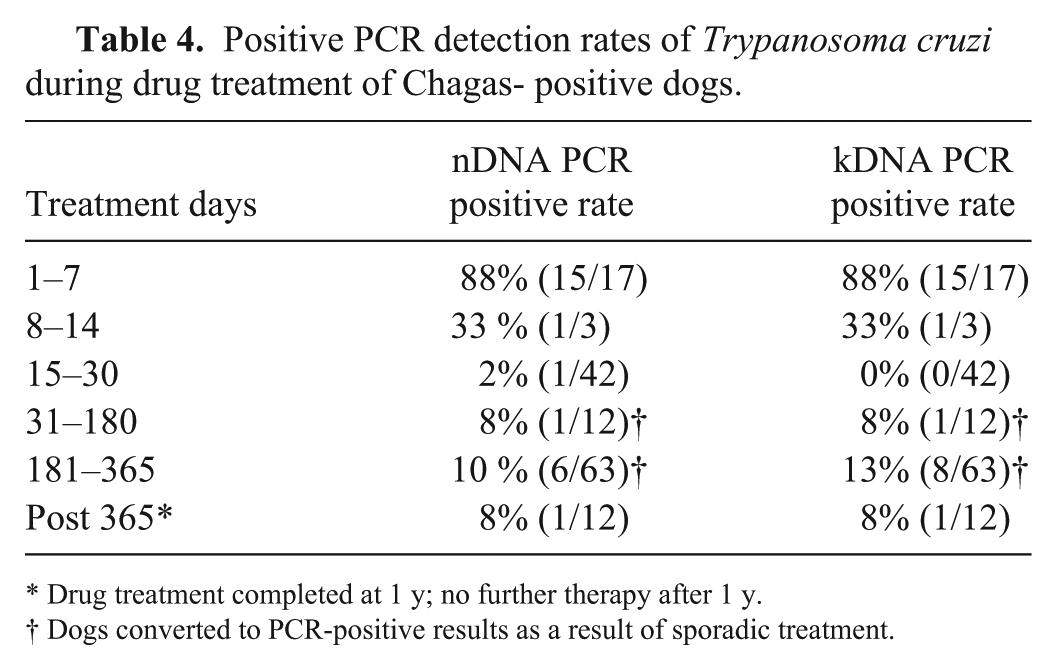
Drug treatment completed at 1 y; no further therapy after 1 y.
Dogs converted to PCR-positive results as a result of sporadic treatment.
Among the other 11 samples, consistent PCR-negative results were observed up to 36 mo after treatment. The 1 case with a sudden strong PCR-positive result occurring 1 y post-treatment suggests an exogenous re-infection, although endogenous reactivation cannot be ruled out. The second round of treatment was initiated once relapse was confirmed by positive PCR results, and parasite DNA was undetectable in the blood after 8 d of treatment (Fig. 2, dog D), indicating that no drug resistance occurred. Descriptive statistics was applied to the PCR positive rate decrease during drug treatment (Table 4) because not every dog had complete follow-up data at every time, and the times were too few for time series analysis.
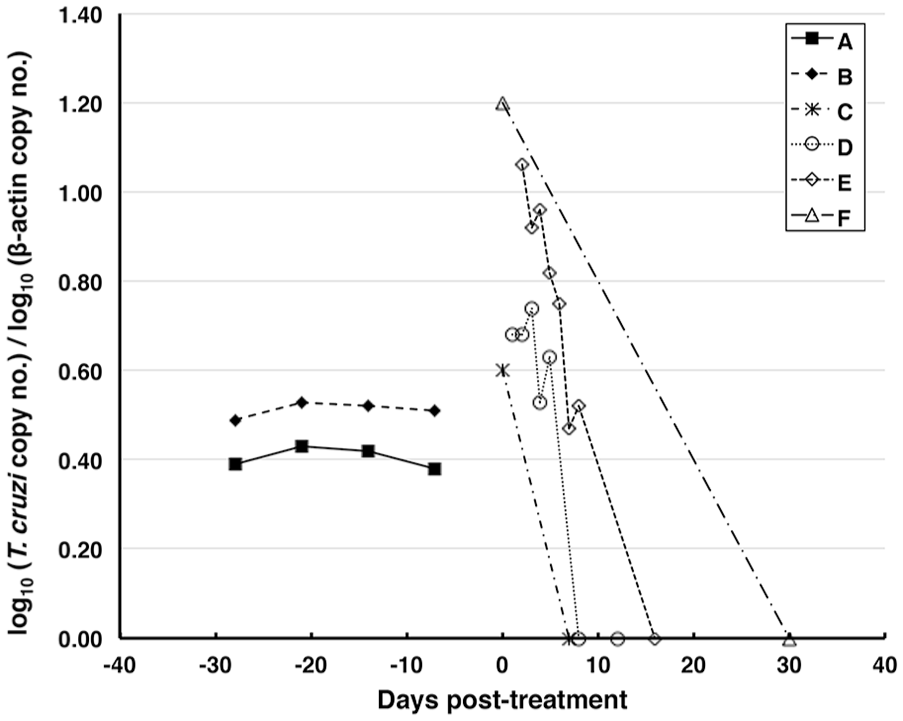
Parasite circulating blood load kinetics using kDNA-based qPCR. Letters A and B are Chagas-positive control dogs with no treatment. Letters C–F are Chagas-positive dogs with drug treatment.
To measure T. cruzi parasite loads in canine whole blood during drug treatment, quantitative kDNA rtPCR was performed because the kDNA-based PCR method was slightly more sensitive than the nDNA-based PCR method. Parasite loads were dramatically reduced within 7–30 d in the 4 representative drug-treated, Chagas-positive dogs (Fig. 2). Furthermore, 7 dogs that were outliers in their DNA copy baselines also showed reduced parasite loads within 30 d (data not shown).
In the highly T. cruzi–prevalent state of Texas, the seropositive rate was 44% (57 of 131) and PCR positive rate was 21–24% (27–31 of 131) from testing the randomly collected untreated dogs (Tables 1, 2). There is no U.S. Food & Drug Administration–approved treatment of canine Chagas disease. To evaluate the antiparasitic drug choice to treat canine Chagas disease, our qualitative rtPCR can be useful to rapidly screen blood samples for the presence of parasite DNA during drug treatment (Table 4), and is more cost-effective than qPCR because no standards with various copy numbers are included in each run. Based on the slightly higher DSe observed in kDNA- than in nDNA-based qualitative rtPCR, the former assay may be used for routine screening and the latter, with a slightly higher DSp, may be used to confirm results. Quantitative rtPCR, however, can be applied to assess the kinetic impact of drug therapy by measuring changes in circulating parasite loads. Positive results obtained from our qualitative and qPCR after 2 wk post itraconazole and amiodarone treatment may indicate treatment failure. Negative rtPCR results indicate undetectable parasitemia, but such results cannot be used as the sole criterion for Chagas disease cure because their correlation with tissue parasitism and clinical outcome is unknown. Using seroreversion of anti–T. cruzi IgG antibodies after treatment as an indicator of canine Chagas disease cure is also controversial because an anti–T. cruzi IgG titer may not persist after treatment.3,5,10 In our drug-treated dogs, no seroreversion of anti–T. cruzi IgG was observed up to 3 y post-treatment (unpublished data). Additional negative results from serologic tests, such as testing anti-live amastigote, anti-live trypomastigote, and fixed epimastigote antibodies simultaneously with flow cytometry, 1 are used to assess human Chagas disease cure after drug treatment, and they have the potential to be adapted for testing dog samples.
Another limitation of our PCR methods is that genetic variability of T. cruzi strains may affect our PCR test performance and cause false-negative PCR results. For example, copy numbers of nuclear satellite repeats might be different in T. cruzi strains with distinct discrete taxonomic units (DTUs), causing the sensitivity of nDNA PCR to be T. cruzi strain–dependent.6,7 At least 6 DTUs were reported, but only TcI and/or TcIV were found in dogs in Texas, and the majority belong to TcI.4,18 The performance of our PCR methods should be tested using T. cruzi strains with different DTUs, and cross-reactivity with other Trypanosoma species needs to be determined.
Following trypanocidal treatment, low or undetectable levels of parasitemia are expected. Multiple blood collections and PCR re-testing, similar to the procedure for detecting low levels of parasitemia in infected dogs during the chronic phase, 2 are therefore recommended to prevent false-negative results from being interpreted as treatment success. Furthermore, using nDNA- and kDNA-based rtPCR methods in parallel may be complementary to detect parasitic DNA for the evaluation of treatment outcome.
Footnotes
Acknowledgements
Declaration of conflicting interests
The authors declare no potential conflicts of interest with respect to the research, authorship, and publication of this article.
Funding
The authors received no financial support for the research, authorship, and publication of this article.
