Abstract
Salmonella enterica resistance to extended-spectrum cephalosporins (ESC) conferred by cefotaximases (blaCTX-M) is a growing concern in the United States. Among food-producing animals, poultry are a major reservoir of ESC-resistant Salmonella. A retrospective study was carried out to further characterize 38 ceftiofur-resistant clinical Salmonella enterica isolates obtained from poultry during 2007–2018. Of the isolates tested, 31 displayed resistance to ceftriaxone and harbored blaCMY-2, whereas 7 isolates demonstrated resistance or reduced susceptibility to cefepime in addition to ceftriaxone resistance. These 7 isolates displayed extended-spectrum β-lactamase activity, harbored blaCTX-M-1, and were recovered only from recent poultry diagnostic submissions made in 2011–2018 as opposed to the 31 isolates that were recovered in 2007–2018. Further characterization of the blaCTX-M-1 gene determined that it was located on conjugative IncN/ST1 and IncI1/ST87 plasmids in the isolates from commercial turkeys and broilers, respectively. These plasmids have been responsible for extensive spread of blaCTX-M-1 in livestock, poultry, and humans in Europe. Potential transfer of IncN and IncI1 plasmids and/or nontyphoidal Salmonella carrying these plasmids through the food chain, or by other means to humans, may result in treatment failures. Our study demonstrates the importance of further characterization of ceftiofur-resistant S. enterica isolates detected by veterinary diagnostic laboratories to identify the sources of blaCTX-M-1 and to mitigate the spread of ESC-resistant Salmonella in the poultry production pyramid.
Introduction
The emergence of resistance to extended-spectrum cephalosporins (ESCs) in Salmonella enterica is a growing public health concern in the United States (NARMS integrated report, 2015. Available from: https://www.fda.gov/downloads/AnimalVeterinary/SafetyHealth/AntimicrobialResistance/NationalAntimicrobialResistanceMonitoringSystem/UCM581468.pdf). ESC resistance is conferred predominantly by Ambler class C cephamycinases (CMY), carbapenamases, and Ambler class A extended-spectrum β-lactamases (ESBLs). 2 The ESBLs consist of many different families of β-lactamases, including TEM, SHV, and CTX-M enzymes. 2 Among the ESBL families, the newest family (CTX-M), represents the highest number of variants, which have been classified into 5 major clusters. 1 In Europe, the incidence of ESBL- and/or AmpC-producing, ESC-resistant Salmonella in humans, food-producing animals, and foods of animal origin has been extensively reported since ~2000.3,11 Although ESBL and/or AmpC-producing bacteria have been recovered from all major food-producing animals, poultry and poultry products are the most frequently reported to carry these strains.13,35 In contrast to reports from other countries, blaCTX-M ESBL-producing Salmonella of livestock and poultry origin have been rarely reported in the United States (NARMS integrated report, 2015). ESC resistance in human, animal, and retail meat nontyphoidal Salmonella of U.S. origin was predominantly associated with plasmid-mediated blaCMY-2 (NARMS integrated report, 2015). Food-producing animals are considered to be the primary reservoir of ESC-resistant Salmonella and, therefore, these Salmonella and their resistance gene repertoire are potentially transferred to humans via the food chain, contaminated environment, or animal handling.4,28
Ceftiofur, a third-generation ESC, had been approved for the control of early mortality associated with Escherichia coli infections in day-old chicks and poults in U.S. hatcheries (U.S. FDA, Extra label use and antimicrobials, 2018.08.24. Available from: https://www.fda.gov/animalveterinary/safetyhealth/antimicrobialresistance/ucm421527.htm). Ceftriaxone, another third-generation ESC, is the drug of choice for the treatment of invasive Salmonella infections in children (U.S. FDA, Drug approvals and databases, 2019.01.04. Available from: https://www.fda.gov/Drugs/InformationOnDrugs/default.htm). Resistance in Salmonella isolates to ceftiofur and ceftriaxone is highly correlated with the presence of the blaCMY-2 gene in the United States. 24 Studies have rarely focused on the clinical isolates of Salmonella from food-producing animals, although Salmonella isolated from veterinary diagnostic submissions display a high level of resistance to ceftiofur.20,41 Such resistance may be the result of selection pressure imposed by the prophylactic use of ceftiofur or acquisition of resistant Salmonella or their genes from other birds or a contaminated poultry house environment. Our objective was to characterize ceftiofur-resistant nontyphoidal Salmonella, previously isolated from poultry clinical specimens in 2007–2018, to better understand the molecular mechanisms responsible for their ESC resistance.
Materials and methods
Bacterial isolates
Clinical isolates of S. enterica (n = 38) obtained from poultry and known to be resistant to ceftiofur (minimum inhibitory concentration [MIC] >4 µg/mL) using the Sensititre avian MIC panel AVIAN1F (Thermo Fisher Scientific, Cleveland, OH), tested in 2007–2018 at the Animal Diagnostic Laboratory of The Pennsylvania State University (PSU; University Park, PA), were identified and further characterized.
Antimicrobial susceptibility testing for ESBL
Salmonella isolates were tested with a broth microdilution test using a panel designed for ESBL confirmatory testing (Sensititre extended spectrum beta-lactamase plate; Thermo Fisher Scientific). The panel contained cefazolin (8–16 μg/mL), cefepime (1–16 μg/mL), cefoxitin (4–64 μg/mL), meropenem (1–8 μg/mL), cephalothin (8–16 μg/mL), cefpodoxime (0.5–64 μg/mL), ceftriaxone (1–128 μg/mL), ciprofloxacin (1–2 μg/mL), gentamicin (4–16 μg/mL), ampicillin (8–16 μg/mL), imipenem (0.5–16 μg/mL), piperacillin–tazobactam (4/4–64/4 μg/mL), ceftazidime (0.25–128 μg/mL), ceftazidime–clavulanic acid (0.25/4–128/4 μg/mL), cefotaxime (0.25–64 μg/mL), and cefotaxime–clavulanic acid (0.25/4–64/4 μg/mL). Antimicrobial MICs were determined using the Sensititre complete automated system (Thermo Fisher Scientific) in accordance with the manufacturer’s instructions, and results were interpreted in accordance with the Clinical and Laboratory Standards Institute (CLSI) criteria breakpoints. 7 E. coli ATCC 25922, Enterococcus faecalis ATCC 29212, Staphylococcus aureus ATCC 29213, and Pseudomonas aeruginosa ATCC 27853 were used as quality control organisms in the antimicrobial MIC determinations.
Characterization of β-lactamase genes
To detect the presence of β-lactamase genes, PCR amplifications and sequencing were performed. Genomic DNA was prepared by boiling the bacteria at 95°C for 10 min and collecting the supernatant after centrifugation for 10 min at 5,000 × g. Initially, the presence of 5 different β-lactamase genes (blaCMY, blaCTX-M, blaOXA, blaSHV, and blaTEM) was determined by PCR and sequencing as described previously.9,21,30 The primers used and the amplicon sizes detected by the PCR were as follows: CMY-2 (F, 5’-GCACTTAGCCACCTATACGGCAG-3’; R, 5’-GCTTTTCAAGAATGCGCCAGG-3’; 758 bp), CTX-M (F, 5’-ATGTGCAGYACCAGTAARGTKATGGC-3’; R, 5’-TGGGTRAARTARGTSACCAGAAYCAGCGG-3’; 593 bp), SHV (F, 5’-TTATCTCCCTGTTAGCCACC-3’; R, 5’-GATTTGCTGATTTCGCTCGG-3’; 795 bp), TEM (F, 5’-GCGGAACCCCTATTTG-3’; R, 5’-ACCAATGCTTAATCAGTGAG-3’; 964 bp), OXA-1 (F, 5’-ATGAAAAACACAATACATATCAACTTCGC-3’; R, 5’-GTGTGTTTAGAATGGTGATCGCATT-3’; 820 bp), and OXA-2 (F, 5’-ACGATAGTTGTGGCAGACGAAC-3’; R, 5’-ATYCTGTTTGGCGTATCRATATTC-3’; 602 bp). The CTX-M–positive samples were subsequently amplified by PCR with primers specific to CTX-M-1 (F, 5’-GACGATGTCACTGGCTGAGC-3’; R, 5-AGCCGCCGACGCTAATACA-3’; 499 bp). 31 The amplicons were gel purified and sequenced on both strands at the Genomics Core Facility (PSU). The obtained sequences were queried against the National Center of Biotechnology Information (NCBI) nucleotide database using the BLASTn algorithm.
Plasmid characterization
Plasmids were extracted from 7 isolates of blaCTX-M-1–positive Salmonella for further analysis. Briefly, bacteria were grown overnight at 37°C in 5 mL of Luria Bertani (LB) broth. Plasmids were extracted by removing chromosomal DNA with alkaline sodium dodecyl sulfate followed by a phenol-chloroform extraction method, 27 as well as by using a commercial kit (Mini plasmid kit; Qiagen, Valencia, CA). Extracted plasmids were separated on 1.2% agarose gel along with plasmids of known sizes for molecular weight determination.
Conjugation
To assess the transferability of resistance, conjugation experiments were carried out following both broth-mating and filter-mating techniques as described previously. 6 Each of the blaCTX-M-1–positive Salmonella isolates was used as the donor strain; nalidixic acid–resistant E. coli DH5α served as the recipient strain. For broth mating, fresh overnight cultures of donor and recipient cells were mixed at a donor-to-recipient ratio of 1:10 and were incubated at 37°C with gentle agitation for 4 h. The bacterial mixture was vortexed and plated on MacConkey agar containing 20 μg of nalidixic acid/mL and 16 µg of cefepime/mL. Colonies were counted after incubation at 37°C for 48 h. Filter mating was performed by collecting the fresh 1:10 donor–recipient mixture on a membrane filter (Nalgene 0.2 µm polyethersulfone membrane; Thermo Fisher Scientific) and placing the filter on MacConkey agar followed by incubation at 37°C overnight. Then, the filter containing the bacteria was suspended in 1 mL of LB, and serial dilutions were made prior to plating on MacConkey agar containing 20 µg of nalidixic acid/mL and 16 µg of cefepime/mL to select for the transconjugants. Colonies were counted after incubating the plates for 48 h. Corresponding donor bacteria alone were plated to determine the efficiency of conjugation (transfer efficiency per donor).
Replicon typing
DNA was isolated from the transconjugants by a rapid boiling method as described elsewhere. 25 Replicon types were determined by a previously described PCR-based method that detects 18 replicon types in 3 multiplex PCR assays. 25
Plasmid multi-locus sequence typing
Plasmids were extracted from the transconjugants (Mini plasmid kit; Qiagen), and incompatibility groups (IncI1, IncHI2, IncF, and IncN) were determined according to the scheme available from the plasmid multi-locus sequence typing (pMLST) website (http://pubmlst.org/plasmid/). Briefly, the corresponding genes for each incompatibility group were amplified with PCR, and the amplicons were sequenced on both strands (Genomics Core Facility, PSU) as described previously.15-17,38 Sequences were queried against the database of the pMLST website to determine the ST types.
Pulsed-field gel electrophoresis
CTX-M-1–positive Salmonella isolates were subjected to pulsed-field gel electrophoresis (PFGE), according to the Centers for Disease Control (CDC) PulseNet protocol, using XbaI as the restriction enzyme (Document PNL05, 2017.12. Available from: https://www.cdc.gov/pulsenet/pdf/ecoli-shigella-salmonella-pfge-protocol-508c.pdf). PFGE profile designations were assigned by the CDC according to PulseNet USA, the national database for Salmonella PFGE profiles, which contains >500,000 entries. 32 PFGE patterns were compared to understand the epidemiologic and genetic relatedness of isolates belonging to the same serovar using previously published guidelines.19,37
Results
Ceftiofur-resistant Salmonella collection
The 38 ceftiofur-resistant poultry clinical Salmonella isolates were from commercial turkey (n = 19), commercial broiler (n = 17), pheasant (n = 1), and guinea fowl (n = 1). These isolates belonged to the following 10 serovars: S. Kentucky (n = 13) followed by S. Heidelberg (n = 8), S. Bredeney (n = 7), S. Typhimurium (n = 2), S. Worthington (n = 2), S. Senftenberg (n = 2), S. Derby (n = 1), S. Infantis (n = 1), S. Montevideo (n = 1), and S. Newport (n = 1; Table 1).
Distribution of β-lactamases among 38 ceftiofur-resistant clinical Salmonella isolates of poultry, and minimum inhibitory concentration (MIC) ranges of cefoxitin, cefotaxime, ceftriaxone and cefepime.
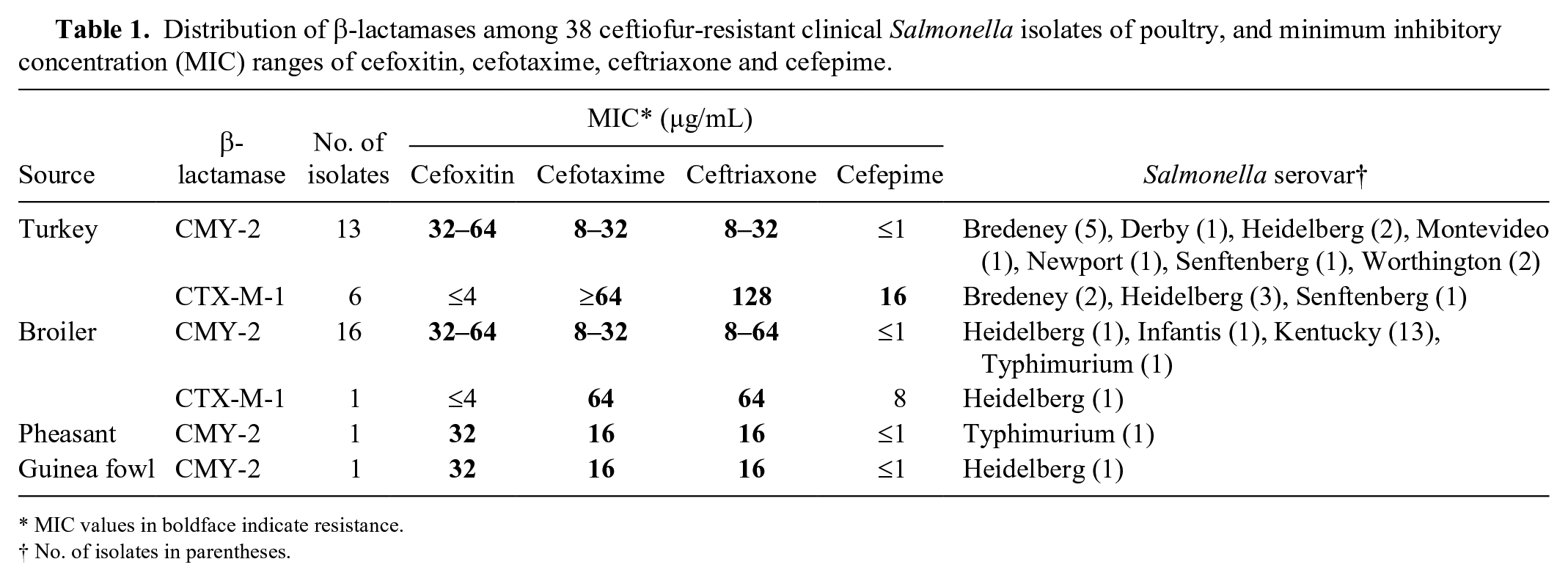
MIC values in boldface indicate resistance.
No. of isolates in parentheses.
Antimicrobial susceptibility testing for ESBL
All 38 ceftiofur-resistant S. enterica isolates were also resistant to ceftriaxone. Additionally, 6 isolates were resistant to cefepime (MIC = 16 µg/mL); another isolate had reduced susceptibility to cefepime, with an MIC of 8 µg/mL. These 7 isolates demonstrated a ≥8 times concentration decrease in MIC for both cefotaxime and ceftazidime in combination with clavulanic acid than the MIC of cefotaxime and ceftazidime alone, and showed sensitivity to cefoxitin consistent with ESBL production. The other 31 isolates were resistant to cefoxitin. None of the isolates were resistant to ciprofloxacin or the 2 carbapenems tested (imipenem and meropenem). Ten of the 38 isolates were also resistant to gentamicin (Table 1).
Characterization of β-lactamase genes
Most isolates assayed (31 of 38) were PCR-positive for blaCMY-2, and 7 isolates were positive for blaCTX-M. PCR amplicon sequencing confirmed the blaCTX-M gene type as blaCTX-M-1 (GenBank accessions MH195155, MH195156, MH195157, MH195158, MH195159, MH195160, MK341546). None of the isolates possessed both blaCMY-2 and blaCTX-M−1. All of these isolates were negative for blaOXA, blaSHV, and blaTEM by PCR (Table 1).
Plasmid characterization
Plasmid profiling of wild-type and corresponding transconjugant strains revealed that ESBL resistance was conferred by transferable plasmids. Six of 7 resistance plasmids were transferable to E. coli DH5α by both broth and filter mating at a frequency of ~4.4 × 10−6 to 3.8 × 10−5, whereas one isolate was not transferable under the experimental conditions used in the study. The ESBL resistance plasmid in turkey isolates of S. Heidelberg and S. Bredeney was ~40 kb in size, whereas the resistance plasmid of the broiler isolate of S. Heidelberg was ~100 kb (Fig. 1). Plasmid replicon typing and pMLST performed on the transconjugants typed the ~40-kb plasmids as IncN/ST1 and the ~100-kb plasmid as IncI1/ST87 (Table 2).
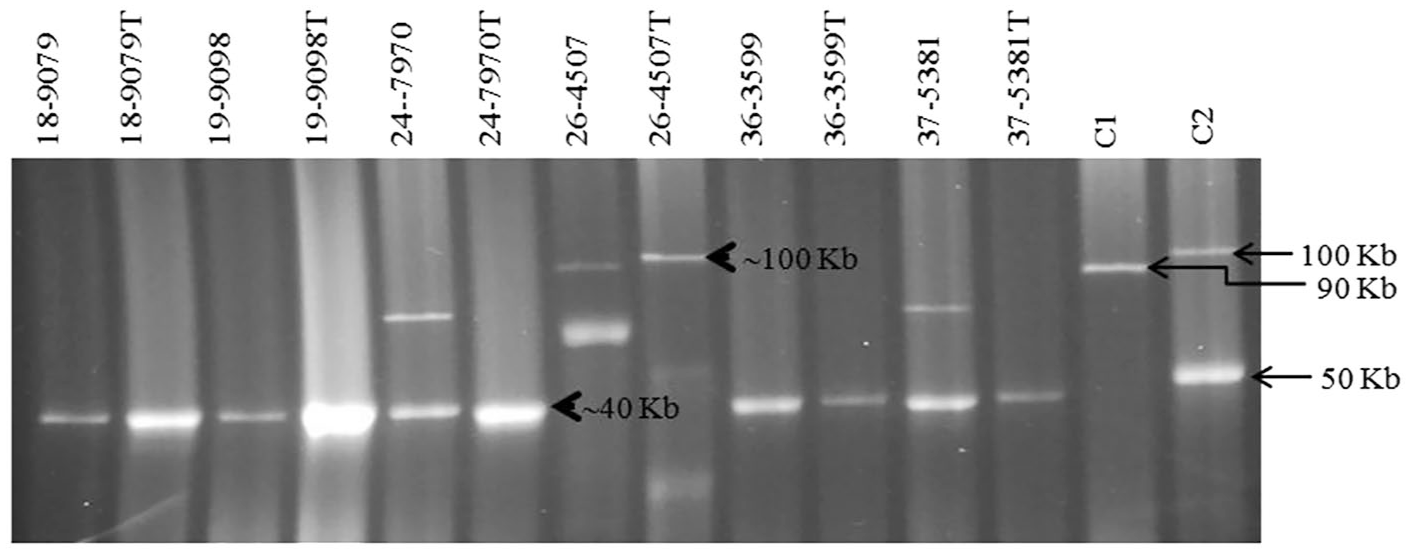
Plasmid profiles of 6 Salmonella clinical isolates of poultry harboring blaCTX-M-1 wild type and their transconjugants. Lanes C1 and C2 are Escherichia coli plasmids with known sizes. Isolate designations ending with letter “T” indicate plasmid profiles of corresponding transconjugants.
Characteristics of Salmonella clinical isolates of poultry harboring blaCTX-M-1 and their corresponding Escherichia coli DH5α transconjugants (T).
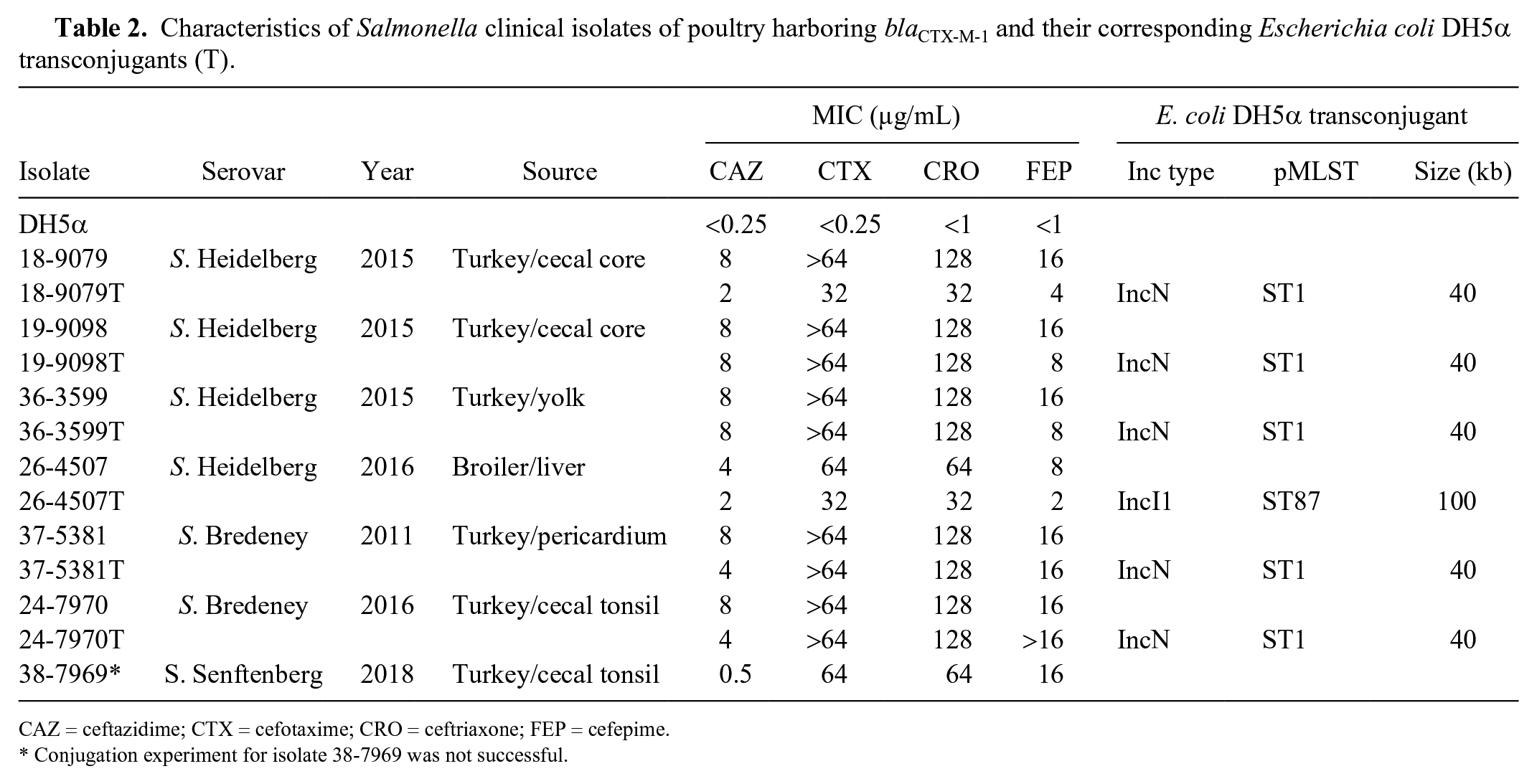
CAZ = ceftazidime; CTX = cefotaxime; CRO = ceftriaxone; FEP = cefepime.
Conjugation experiment for isolate 38-7969 was not successful.
PFGE
Each of 7 blaCTX-M-1–positive isolates produced a different PFGE profile. Based on the PFGE band patterns, pulsotype designations were assigned by CDC PulseNet for blaCTX-M-1–positive S. Heidelberg and S. Bredeney isolates, but not for the S. Senftenberg isolate. Although the PFGE patterns of the isolates belonging to the same serovar were not identical, 3 turkey S. Heidelberg isolates differed from each other by only 1–3 bands, suggesting that these isolates are similar, represent subtypes, and are epidemiologically related (Fig. 2). The broiler isolate of S. Heidelberg was different from turkey S. Heidelberg by 4–6 bands, suggesting a possible genetic relatedness and that this isolate may have the same genetic lineage as that of the other isolates, but is less likely to be epidemiologically related. The PFGE patterns of the 2 S. Bredeney isolates differed by only 1 band, indicating these 2 isolates represent subtypes and are epidemiologically related.
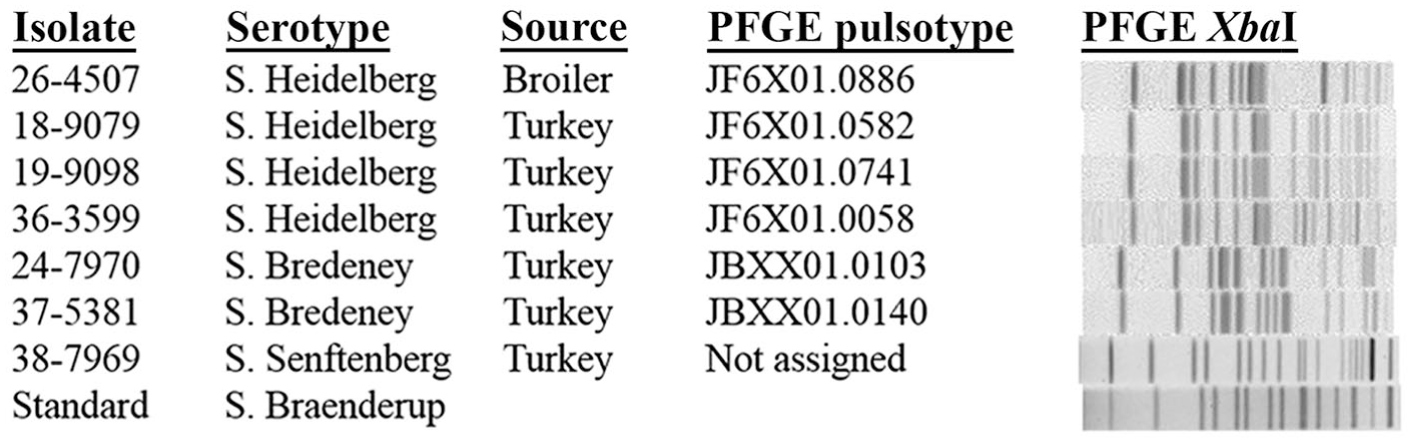

Pulsed-field gel electrophoresis (PFGE) profiles of Salmonella enterica clinical isolates positive for CTX-M–1.
Discussion
Ceftriaxone, a third-generation ESC, is the antibiotic recommended to treat invasive Salmonella infections in children (U.S. FDA, Drug approvals and databases, 2019.01.04). Accordingly, increasing resistance to ESCs in nontyphoidal Salmonella has become a serious public health issue in the United States. Resistance of S. enterica of livestock and poultry origin to ceftiofur, the veterinary counterpart of ceftriaxone, has increased during the last 2 decades (NARMS integrated report, 2015). This may pose a public health risk given that ceftiofur resistance in Salmonella is known to be correlated with resistance to ceftriaxone.10,24 As well, ceftriaxone resistance in S. Heidelberg in humans is strongly correlated with ceftriaxone resistance in S. Heidelberg isolated from poultry at slaughter and at the retail level. 24 This is of public health concern because poultry serves as a reservoir for nontyphoidal Salmonella, and S. Heidelberg, in particular, is the most common cause of invasive Salmonella infection in humans. 8 In the United States, ceftiofur and ceftriaxone resistance in clinical isolates of human nontyphoidal Salmonella and nonclinical isolates of livestock and poultry Salmonella was predominantly associated with plasmid-mediated blaCMY-2, whereas blaCTX-M–mediated ESBL resistance is rarely reported.29,40
We describe the detection of blaCTX-M-1 ESBL-producing S. Heidelberg, S. Bredeney, and S. Senftenberg in poultry clinical isolates. Although infrequently reported in the United States at present, the high prevalence of plasmid-mediated blaCTX-M ESBL-producing Salmonella serovars in humans and food-producing animals in other countries, in particular in Europe, suggests a potential risk of their dissemination in the United States (JIACRA report, 2017.06.28. Available from: https://www.ema.europa.eu/documents/report/ecdc/efsa/ema-second-joint-report-integrated-analysis-consumption-antimicrobial-agents-occurrence_en.pdf). In our study, all 38 isolates of ceftiofur-resistant poultry clinical Salmonella were resistant to ceftriaxone. However, only 7 (18%) isolates that were also resistant, or demonstrated reduced susceptibility, to cefepime were positive for blaCTX-M–1. The ESC resistance in a majority (n = 31, 81%) of Salmonella was conferred by blaCMY-2, and the most common serovar among blaCMY-2–positive isolates was S. Kentucky (n = 13, 42%), a serovar that is not generally implicated in foodborne infection (NARMS integrated report, 2015). However, the contribution of this serovar to the blaCMY-2 gene pool cannot be underestimated. Interestingly, except for one S. Heidelberg isolate from the liver of a commercial broiler chicken, all other blaCTX-M-1–positive Salmonella (S. Heidelberg = 4, S. Bredeney = 2, and S. Senftenberg = 1) were isolated from turkey specimens. The MICs of turkey blaCTX-M-1–positive isolates for ceftriaxone and cefepime were higher than the MIC of the broiler isolate. Although all Salmonella were isolated at autopsy, presence of blaCTX-M-1–positive Salmonella indicates a potential zoonotic transmission of resistant Salmonella, blaCTX-M-1, or the plasmid harboring blaCTX-M-1 through the food chain or by direct contact as a result of environmental contamination. The ESBL-encoding plasmids reported herein were transmissible to E. coli DH5α at a rate of 4.4 × 10−6 to 3.8 × 10−5, providing evidence for potential widespread dissemination of these resistance plasmids to other bacteria inside or outside of the bird intestines. However, transferability of plasmids is dependent on a number of factors, including the presence or absence of multiple plasmids in the donor and recipient strains as well as plasmid incompatibility groups. 14 Further, in our study, one of the turkey cecal specimens (18-9079) was coinfected with both S. Heidelberg and E. coli harboring identical ESBL-plasmid IncN/ST1 (data not shown). It is widely known that blaCTX-M-1 has been frequently associated with IncN and IncI1 plasmids in avian enteric E. coli. 26
S. Heidelberg is more commonly implicated in invasive infection in humans than the other nontyphoidal serovars. 8 This serovar is also capable of causing extraintestinal infections and severe septicemia in broiler chickens. 39 Studies in the United States have found that ceftriaxone-resistant S. Heidelberg in humans is strongly correlated with ceftriaxone resistance in isolates from poultry at slaughter.12,23,24 A previous study also found a strong correlation between isolation of ceftiofur-resistant S. Heidelberg from retail chicken and incidence of ceftiofur-resistant S. Heidelberg infections in humans in Canada. 10 The major source of S. Heidelberg associated with human outbreaks in the United States is considered to be the consumption of poultry.5,24 As well, multidrug- and pan-resistant strains of S. Heidelberg have been isolated from chicken meat and ground turkey.18,33,36 It is noteworthy to mention that 4 of 7 blaCTX-M-1–positive Salmonella found in our study were also S. Heidelberg, highlighting the importance of implementing mitigation strategies to control the spread of these organisms. Most importantly, blaCMY-2–positive Salmonella belonging to 10 different serovars, including S. Heidelberg and S. Bredeney, were present throughout the study period (2007–2018), whereas blaCTX-M-1–positive S. Heidelberg, S. Bredeney, and S. Senftenberg were found during more recent years. The only blaCTX-M-1–positive serovars of Salmonella of poultry origin reported by previous studies were S. Bredeney and S. Ouakam, 40 whereas we report the presence of blaCTX-M-1–positive S. Heidelberg and S. Senftenberg in poultry in the United States.
In our study, ESBL resistance in turkey isolates was linked to an IncN/ST1 type 40-kb plasmid, whereas the ESBL resistance of the commercial broiler isolate of S. Heidelberg was conferred by a 100-kb plasmid that was typed as IncI1/ST87. Another study also reported blaCTX-M-1–positive, ESBL-producing poultry Salmonella harboring blaCTX-M-1 on an IncN type plasmid. 40 However, the IncN plasmid detected in our study was further characterized as IncN/ST1, which has been found to be the pandemic ESBL plasmid widely distributed in livestock and humans in Europe. 34 The presence of a pandemic IncN/ST1 ESBL plasmid in S. Heidelberg is important considering the potential for this serovar to cause systemic infections through zoonotic and foodborne spread in the United States. The pandemic plasmid IncN/ST1 was responsible for the extensive spread of the blaCTX-M-1 gene in livestock and humans in Europe. 34 To date, no human infections linked to the IncN/ST1 pandemic plasmid have been reported in the United States, to our knowledge. However, the risk of potential zoonotic and foodborne transmission of ESBL-resistant S. Heidelberg highlights the need for active targeted surveillance. PFGE profiling revealed that S. Heidelberg isolates are genetically related and might have descended from a common lineage. However, the broiler S. Heidelberg strain isolated in 2016 is less likely to be epidemiologically related to the turkey S. Heidelberg strains isolated in 2015, suggesting a recent genetic event that made this isolate different from turkey isolates. However, the 2 S. Bredeney strains isolated in 2011 and 2016 were closely related, demonstrating epidemiologic relatedness. Interestingly, the S. Senftenberg isolate from 2018 did not match any existing PFGE profiles and may indicate a novel epidemiologic clone harboring blaCTX-M−1.
Because ceftiofur-resistant S. enterica may harbor blaCTX-M-1, it is important to further characterize ceftiofur-resistant Salmonella isolates detected by veterinary diagnostic laboratories. Without this extra step, it will not be possible to identify the sources of blaCTX-M-1, a step that is required to prevent the spread of ESC-resistant Salmonella in the poultry production pyramid. 22
Footnotes
Acknowledgements
We thank National Veterinary Services Laboratories (Ames, IA) for providing technical assistance for PFGE profile designations.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was partly supported by a grant from the Pennsylvania Department of Agriculture (grant PEN04588).
