Abstract
Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) has emerged as a reliable method to identify fungal isolates. The success of this approach relies on the availability of exhaustive databases, but the latter were built with a focus on human pathogens. We assessed a large in-house database of reference spectra and a dedicated web application for their suitability for use in veterinary laboratories. A panel of 290 mold and yeast isolates representing 69 different fungal species was isolated from various animals (including pets, cattle, and zoo animals) and identified using both MALDI-TOF MS and conventional techniques. The performance of the 2 methods was compared, and identifications were confirmed by DNA sequencing. MALDI-TOF MS allowed distinction between some closely related species and achieved 89% correct identification at the species level. In comparison, only 60% of the isolates were correctly identified with conventional approaches. Using this online application, MALDI-TOF MS thus appears to be a relevant alternative for the identification of fungal isolates encountered by animal health professionals.
The first attempts to identify fungi by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) date back to the early 2000s and were performed on a limited range of species.7,15,19 Following successful MALDI-TOF MS identification of bacteria in clinical laboratories, interest increased in applying the method for fungal pathogens. In this context, numerous studies have demonstrated the reliability of MALDI-TOF MS, which is now routinely used in medical settings for yeasts and increasingly implemented for molds. 10
The performance of this tool, however, depends on a complete database of reference spectra that represents the broad diversity of potential fungi that can be retrieved from clinical samples. However, although yeast species are well covered by commercial databases, molds are relatively poorly covered. 10 Using mainly strains from the Belgian Coordinated Collections of Microorganisms (BCCM/IHEM), an extended database was therefore built for filamentous fungi and made freely available through a website. 18 This database was further complemented with yeast reference spectra and, as of early 2019, has data for ~1,900 strains and 900 different species. For better management of animal fungal infections by veterinary professionals, a fast, reliable, and cost-effective identification method such as MALDI-TOF MS is an advantage. Given that the database was initially created for a human clinical purpose, our goal was to validate the use of the database for isolates sourced from veterinary laboratories.
Isolates were collected in March–September 2017 from the routine activities of 3 Belgian veterinary centers: 1) the Royal Zoological Society of Antwerp that provided isolates from diseased turtles and penguins; 2) Zoolyx, a private veterinary laboratory analyzing samples generally from cats, dogs, and horses, and mostly from superficial skin lesions; and 3) Animal Health Care Flanders, a public laboratory responsible for farm animal health and performing notably systematic mycologic screening on aborted bovine fetuses.
Each center identified isolates using classical methods, namely microscopy for molds and physiologic testing for yeasts. These results were compared to MALDI-TOF MS identification performed in parallel at the BCCM/IHEM. MALDI-TOF MS spectra were obtained in quadruplicate (4 spots per isolate) using standardized methodology described previously. 16 Identification of the spectra was performed using our in-house database and dedicated web application publicly available online at https://biological-mass-spectrometry-identification.com/msi/welcome. Only the MALDI-TOF MS identification with the highest score of the 4 spots was taken into account and considered as a reliable result provided that this score was at least 20.17,18
Multilocus sequencing was also applied to the isolates, as it is considered the gold standard method for fungal identification. 12 It was first used on a random set of 75 isolates representing 25 species for which both MALDI-TOF MS and classical methods resulted in the same identification. DNA sequencing confirmed the obtained identifications in all cases and therefore sequencing was then limited to validating discrepant results between MALDI-TOF MS and traditional identifications (122 isolates). Isolates were cultivated in Sabouraud dextrose broth (molds) or agar (yeasts) at room temperature, and genomic DNA was extracted (Invisorb spin plant mini kit; Stratec, Berlin, Germany) following the manufacturer’s instructions, after prior treatment including freeze-drying and bead-beating. PCR was then performed using primers targeting one or several of the following genes depending on the taxon considered: internal transcribed spacer, 26S rDNA, beta-tubulin, translation elongation factor 1 alpha, calmodulin, RNA polymerase II, glyceraldehyde 3-phosphate dehydrogenase, and actin (Supplementary Table 1). Amplicons were purified (ExoSAP-IT Express PCR product cleanup kit; Affymetrix, High Wycombe, UK) and sequenced (BigDye terminator v.3.1 cycle sequencing kit, 3130xl Genetic Analyzer; Applied Biosystems, Foster City, CA). Sequences were analyzed (Lasergene software v.10; DNASTAR, Madison, WI) and then compared with the MycoBank database (www.mycobank.org) and the GenBank database using the BLAST tool. 3 The best-matched taxon was selected and a minimal 99% sequence similarity were chosen as the criterion for identification at the species level.
A total of 290 isolates, representing 69 different species and including 176 yeasts and 114 molds, was collected in the 3 centers over the 6-mo period (Supplementary Table 2). Yeasts belonged mainly to Candida spp., Diutina spp., and Pichia spp.; molds were represented primarily by Aspergillus spp., Paecilomyces spp., as well as Mucorales (7 species) and dermatophytes (5 species). These isolates corresponded either to true pathogens or to colonizers and contaminants. Pathogens included notable zoophilic dermatophytes that were found on their typical hosts. Dermatophytosis cases were caused by Microsporum canis in cats, Trichophyton equinum in horses, and T. benhamiae from a guinea pig. Noteworthy, a geophilic species, Nannizzia gypsea, was also obtained from a horse skin lesion. Other infectious agents were also found in the animals from the zoo. All 5 Humbolt penguins (Spheniscus humboldti) were affected by aspergillosis caused by Aspergillus fumigatus, a common fungal infection in birds including penguins.4,20 A Branderhorst snapping turtle (Elseya branderhorsti) was found with hyalohyphomycosis caused by Purpureocillium lilacinum, as reported in other turtle species.9,13
Of the 290 isolates, 258 (89% of the total isolates) representing 47 different species were correctly identified at the species level using MALDI-TOF MS (Table 1). This result indicates a parity in correct identification by MALDI-TOF MS between human and veterinary clinical isolates given that it is in the range of the rates obtained in studies using human clinical isolates.6,14,18 Indeed, using the same database as used in our study, an identification rate of 85.6% was attained on a total of 390 clinical molds isolated during the routine activities of 2 Belgian hospitals. 6 In an evaluation of the same web application used in our study, 87.3% correct identification was obtained on a panel of 501 molds and dermatophytes derived from 5 medical laboratories. 18 Regarding clinical yeasts, identification rates for MALDI-TOF systems generally exceed 90% with, for example, up to 95.1% on a large collection of >1,100 human clinical yeast isolates. 5
Results obtained using conventional methods on the 258 isolates correctly identified at the species level by MALDI-TOF MS.

Number of isolates in parentheses.
Morphologic identification at the species complex level.
In comparison, identifications based on morphologic observations and physiologic tests (Supplementary Table 2) were correct at the species (or species complex) level for 174 isolates (60%). For an additional 22 isolates, identification was only correct at (or limited to) the genus level; 63 isolates could not be identified. For the 31 remaining isolates, identification of the genus was incorrect. Consequently, MALDI-TOF MS resulted in fewer limitations as compared to classical methods and appears to be a reliable option for the identification of fungal isolates in animal health settings.
For 30 isolates representing 21 different species, a corresponding reference spectrum was not available in the MALDI-TOF MS database (Table 2). This could indicate that isolates encountered in veterinary practice differ significantly from the isolates obtained in human clinical settings. Such differences suggest that the database requires further extension to deal with important fungal diversity and to identify species that are rarely isolated. Noteworthy, of these 30 isolates, 23 were not identified, showing that species not represented in the database generally do not result in false identifications and that the user should choose another method of identification. Finally, 9 isolates (3% of the total isolates) were incorrectly identified by MALDI-TOF MS (Table 2). These misidentifications resulted generally from confusion between some closely related species, especially if the correct reference was lacking in the database.
Details on the 32 isolates either not identified or incorrectly identified by MALDI-TOF MS.
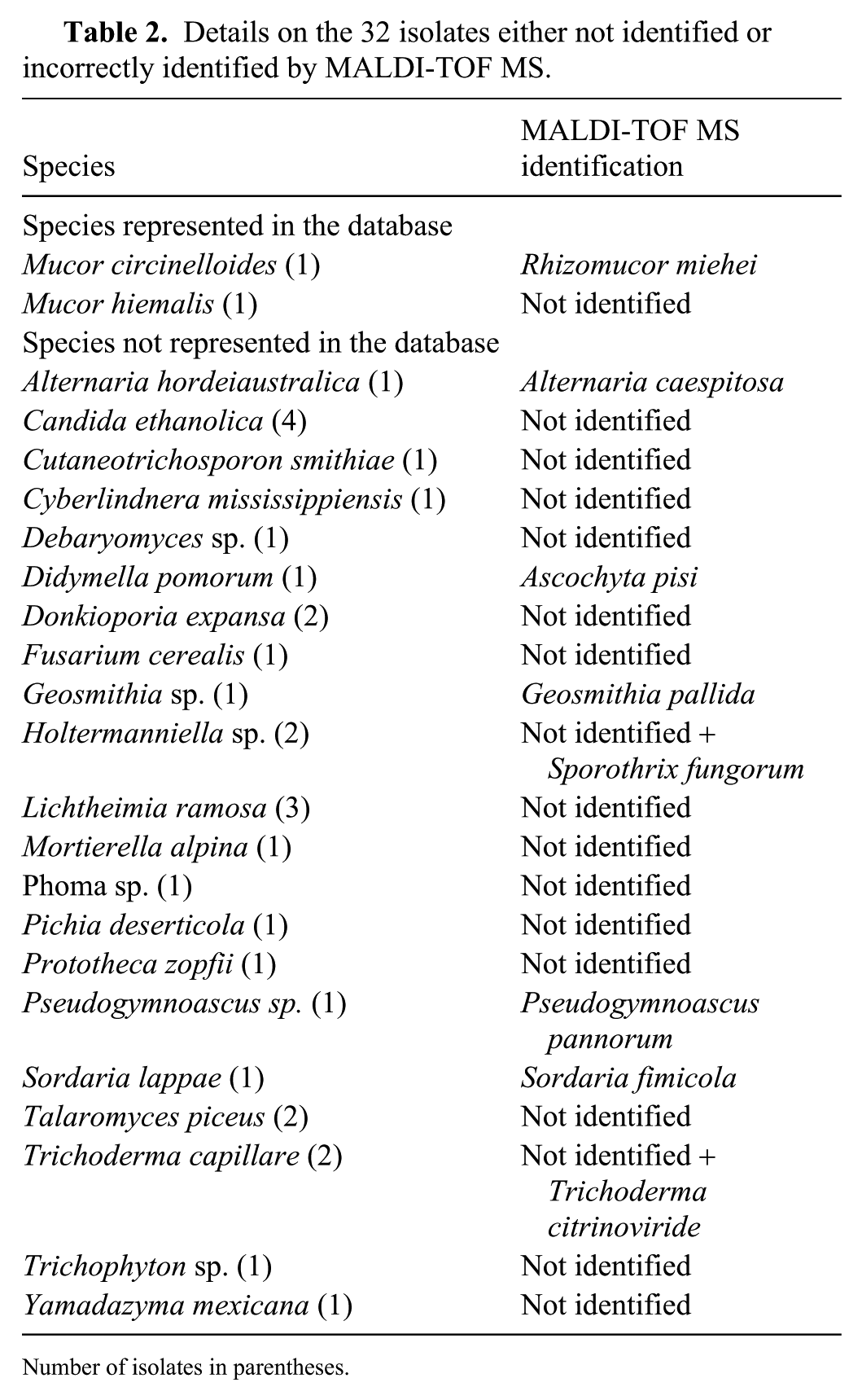
Number of isolates in parentheses.
Interestingly, MALDI-TOF MS was able to differentiate species of the Aspergillus Nigri complex given that it correctly identified A. tubingensis and A. welwitschiae, both representatives of this group of fungi. Likewise, an isolate of Scedosporium boydii, a member of the S. apiospermum complex, was correctly identified. In contrast, microscopy was not able to distinguish these species sharing the same morphology (the 2 isolates of the A. Nigri section were identified as A. niger, whereas S. boydii was recognized as S. apiospermum). Similar discriminatory levels for MALDI-TOF MS were reported previously1,8 and have interesting implications, such as the improvement of epidemiologic data. Moreover, it allows adaptations of treatment given that species of the same complex can display different susceptibilities to antifungal drugs. This is, for example, the case of A. tubingensis identified in our panel, which is more resistant to azoles than other species of the A. Nigri complex.2,11
Our study illustrates that the limitation for successful identification by MALDI-TOF MS lies mainly in the database, which is particularly important regarding the huge diversity in mycology. Limitations of our study include the fact that the sampling period did not cover a whole year. Collecting isolates during the winter season could result in broader species diversity. Moreover, although an effort was made in the study design to recruit laboratories with different scopes, animals and anatomic isolation sites represented only a fraction of the possible situations encountered in veterinary practice.
Overall, our study suggests that our online database, although developed in the first instance for clinical practice in humans, can also be used by animal health professionals for the identification of veterinary isolates by MALDI-TOF MS. It is available for the medical and veterinary community after free registration on the application webpage. It is also regularly supplemented by new reference spectra and curated to fit with the taxonomic developments in mycology.
Supplemental Material
DS1_JVDI_10.1177_1040638719835577 – Supplemental material for Identification of fungal isolates by MALDI-TOF mass spectrometry in veterinary practice: validation of a web application
Supplemental material, DS1_JVDI_10.1177_1040638719835577 for Identification of fungal isolates by MALDI-TOF mass spectrometry in veterinary practice: validation of a web application by Pierre Becker, Anne-Cécile Normand, Gerty Vanantwerpen, Mia Vanrobaeys, Roel Haesendonck, Francis Vercammen, Dirk Stubbe, Renaud Piarroux and Marijke Hendrickx in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Jessie Claessens, Sam Roesems, Kevin Charlier, Sophie Ligot, Tania Bus, Sylvie Goessens, and Ivy Meert for technical assistance.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the Belgian Science Policy (BELSPO).
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
