Abstract
The pestiviruses bovine viral diarrhea virus 1 and 2 (BVDV-1 and -2, respectively) and HoBi-like pestivirus (HoBiPeV) are important pathogens of cattle, and a number of reverse-transcription PCR (RT-PCR)–based assays have been developed for their detection in clinical specimens. We evaluated a newly designed set of pan–bovine pestivirus primers (BP189-389) in a gel-based RT-PCR screening test for pestiviruses in the sera of beef calves destined for export from southern Brazil. Serum samples positive for BVDV antigens by an antigen ELISA (n = 135) were submitted to RT-PCR assays using different sets of primers, followed by nucleotide sequencing of the amplicons. RT-PCR with pestivirus primers 324-326 detected 110 positive samples: BVDV-1 (n = 62), BVDV-2 (n = 38), and HoBiPeV (n = 10). A PCR using primers HCV90-368 detected 97 positive samples (64 BVDV-1; 33 BVDV-2). An additional RT-PCR round using BVDV-2–specific primers (2F-2R) detected 45 positive samples (including 38 detected by primers 324-326 and 33 by HCV90-368); whereas a RT-PCR using HoBiPeV-specific primers (N2-R5) detected 26 positive samples (including 10 detected by primers 324-326).The assay using the primers BP189-389 detected all 135 ELISA-positive samples, including the 26 HoBiPeV detected by primers N2-R5. Our results demonstrated that primers BP189-389 compare favorably against other primer sets in the detection of bovine pestiviruses, especially HoBiPeV. This conventional PCR may be useful for efficient detection of pestiviruses in bovine sera and other specimens as well, especially in laboratories without real-time PCR equipment.
Bovine pestiviruses comprise 2 well-recognized viral species: bovine viral diarrhea virus 1 (BVDV-1; species Pestivirus A) and 2 (BVDV-2; species Pestivirus B), and HoBi-like pestivirus (HoBiPeV; species Pestivirus H). 17 Pestiviruses comprise a genus within the family Flaviviridae and are small (45 nm), enveloped, single-stranded, positive-sense RNA viruses. The pestivirus genome is 12.3 kb in length and contains a long open-reading frame (ORF) flanked by 2 untranslated regions (5’- and 3’-UTRs). The ORF encodes a long polyprotein that is cleaved by viral and host proteases in 11–12 mature viral polypeptides. 18 Pestivirus species are determined by criteria that include comparisons of the complete coding nucleotide (nt) sequences and cross-neutralization titers, in addition to the host of origin. Nucleotide sequencing and comparison of the highly conserved 5’-UTR, Npro, and E2 have served as indicators for pestivirus phylogeny and classification into subtypes.3,9,14,19
BVDV-1 and -2 are among the most important pathogens of cattle, have a global distribution, and are associated with a variety of clinical manifestations in cattle. 8 HoBiPeV were initially identified as contaminants of fetal bovine serum (FBS) of Brazilian origin 16 and subsequently isolated in South America, 20 Italy, 4 and Asia 10 from FBS and from clinical conditions similar to those classically attributed to BVDV-1 and BVDV-2. HoBiPeV isolates are genetically and antigenically related to BVDV-1 and BVDV-2, yet existing genetic and antigenic differences between these groups of viruses may influence diagnostic and control strategies. 1
Reliable tests for pestivirus detection are pivotal for BVDV control, in addition to biosecurity measures, monitoring for BVDV exposure, and vaccination in some cases. Molecular detection through reverse-transcription PCR (RT-PCR) and its variations has been widely used for pestivirus detection. Most RT-PCR assays for pestivirus detection target the highly conserved 5’-UTR.5,7,11–13,19
We employed a newly designed set of primers (BP189-389) to screen the sera of beef calves for pestiviruses in a gel-based RT-PCR. Serum samples positive for BVDV antigens in an antigen ELISA (n = 135) were submitted to RT-PCR using different sets of primers targeting the 5’-UTR of the viral genome (Table 1).
Primer data and sensitivity of the 5’-UTR–based RT-PCR assays used for detection of pestiviruses in sera of calves.
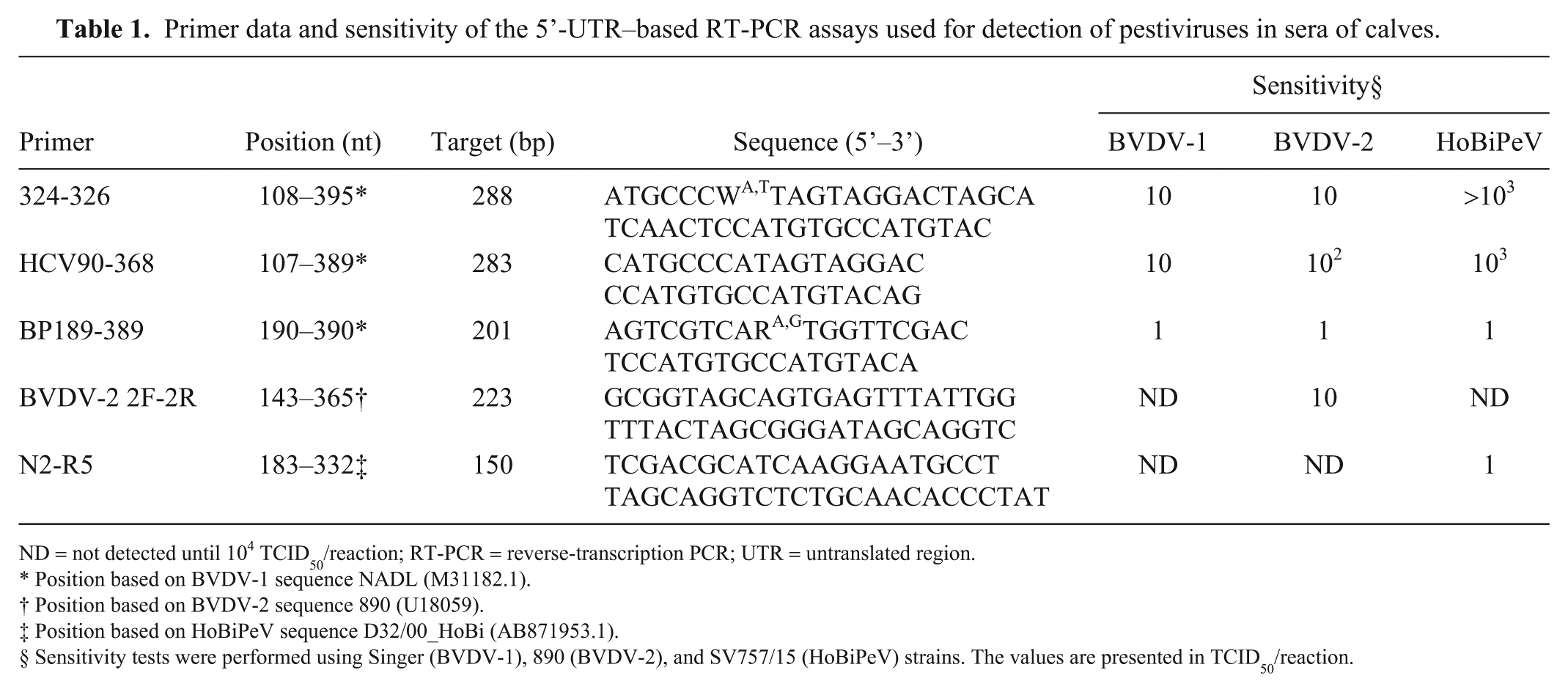
ND = not detected until 104 TCID50/reaction; RT-PCR = reverse-transcription PCR; UTR = untranslated region.
Position based on BVDV-1 sequence NADL (M31182.1).
Position based on BVDV-2 sequence 890 (U18059).
Position based on HoBiPeV sequence D32/00_HoBi (AB871953.1).
Sensitivity tests were performed using Singer (BVDV-1), 890 (BVDV-2), and SV757/15 (HoBiPeV) strains. The values are presented in TCID50/reaction.
BVDV-1 strain Singer, BVDV-2 strain 890, and Brazilian HoBiPeV strain SV757/15 were used as controls in RT-PCR and sensitivity tests. Brazilian HoBiPeV isolates have been described elsewhere. 6 Viruses were amplified and quantified in pestivirus-free Madin–Darby bovine kidney cells (MDBK, ATCC-CCL22) cultured in MEM (Gibco-BRL minimum essential medium, Thermo Fisher Scientific, Waltham, MA), supplemented with 10% equine serum and penicillin, streptomycin, and amphotericin B. Virus stocks were titrated by limiting dilution, and titers were expressed as log10 median tissue culture infective dose (TCID50).
Aliquots of 150 µL of ELISA-positive sera were submitted to RNA extraction (TRIzol reagent, Thermo Fisher Scientific), according to the manufacturer’s instructions. After RNA extraction, complementary DNA (cDNA) was synthesized (SuperScript III reverse transcriptase, Thermo Fisher Scientific). Briefly, total RNA (~500 ng) was mixed with 50 ng of random primers and annealing buffer followed by incubation at 65°C for 5 min, and immediately placed on ice for 1 min. Then, 10 μL of 2× First-strand reaction mix and 2 μL of SuperScript III were added and incubated for 5 min at 25°C, followed by incubation at 50°C for 50 min and 85°C for 5 min. The synthesized cDNA were submitted sequentially to 5 PCR assays, using different sets of primers targeting the 5’-UTR (Table 1), with Taq DNA polymerase, recombinant (Thermo Fisher Scientific). PCR conditions for primers 324-326 19 and HCV90-368 15 were: 94°C for 5 min, 35 cycles of 94°C, 48°C (HCV90-368) or 50°C (324-326), and 72°C, 30 s each, with a final extension at 72°C for 10 min. PCR conditions for BVDV-2–specific primer (2F-2R) 13 were: 94°C for 5 min, 35 cycles of 94°C, 58°C, and 72°C, 45 s each, with a final extension at 72°C for 7 min. PCR conditions for primers specific for HoBiPeV N2-R52 were: 94°C for 3 min, 35 cycles of 94°C, 55°C, and 72°C, 30 s each, with a final extension at 72°C for 5 min. Virus identification was determined by nucleotide sequencing of the amplicons and phylogenetic analysis as described previously. 13
The new primer set (BP189-389) was constructed based on the sequences of BVDV-1 (n = 90), BVDV-2 (n = 60), and HoBiPeV (n = 30) deposited in GenBank (https://www.ncbi.nlm.nih.gov/genbank). Alignment was performed on BioEdit Sequence Alignment Editor v.7.2.6.1 (http://www.mbio.ncsu.edu/BioEdit/bioedit.html). The obtained primers set were analyzed using the primer-BLAST tool to specificity parameters (https://www.ncbi.nlm.nih.gov/tools/primer-blast/). PCR conditions were: 94°C for 5 min, 35 cycles of 94°C, 47°C, and 72°C extension for 30 s each, with a final extension at 72°C for 7 min.
All PCR reactions were performed in a 25-μL volume, using 2 μL of cDNA template (100–200 ng), 0.5 μM of each primer, 3 mM MgCl2, 0.8 mM of dNTPs, 1× reaction buffer, and 1 U of Taq DNA polymerase. PCR products were resolved in 1.5% agarose gel, stained (Gel Red, Biotium, Fremont, CA), and visualized under ultraviolet light after electrophoresis (60 V, 40 min). Total RNA extracted from MDBK cells infected with BVDV-1 Singer, BVDV-2 890, and HoBiPeV SV757/15 were used as controls.
The sensitivity of the assays was investigated by performing PCR in cDNA synthesized from RNA extracted from 150 μL of 10-fold dilutions of supernatants of MDBK cells infected with control viruses.
The range of HoBiPeV detection by primers BP189-389 was further investigated by submitting 3 FBS samples contaminated with HoBiPeV 13 and the supernatant of MDBK cells infected with each of 8 Brazilian isolates 6 to RT-PCR
The design and optimization of primers BP189-389 for annealing to the highest number of available sequences required the introduction of a degenerate nucleotide in the forward primer (nucleotide 200) given the presence of a guanine in BVDV-1 and an adenine in BVDV-2 and HoBiPeV genomes. Primer BP389 (reverse) is very close to primer HCV-368, being shifted by a single nucleotide. Differentiation of the pestivirus species by the amplicon size was not possible using these primers. Unfortunately, these oligonucleotides would not anneal to the genome of pestiviruses of other species (e.g., Bungowannah virus, pronghorn virus, giraffe pestivirus, etc.), thus precluding their use as pan-pestivirus reagents. On the other hand, they would anneal to genomes of the 3 bovine pestivirus species and, thus, may be designated pan–bovine pestivirus primers.
Primers BP189-389 had a broader range of detection—including 26 HoBiPeV detected by primers N2-R5—and higher sensitivity (even for BVDV-1 and BVDV-2) than the PCR assays based on 324-326 and HCV90-368 primers (Table 2). These results confirm previous observations that pestivirus primers (HCV90-368 and 324-326) may fail to detect HoBiPeV, 2 mainly when their genomes are present in small amounts. The specificity of primers N2-R5 for HoBiPeV 2 and primers 2F-2R for BVDV-2 13 was also confirmed.
Pestivirus detection in sera of beef calves by RT-PCR assays using different sets of pestivirus primers.
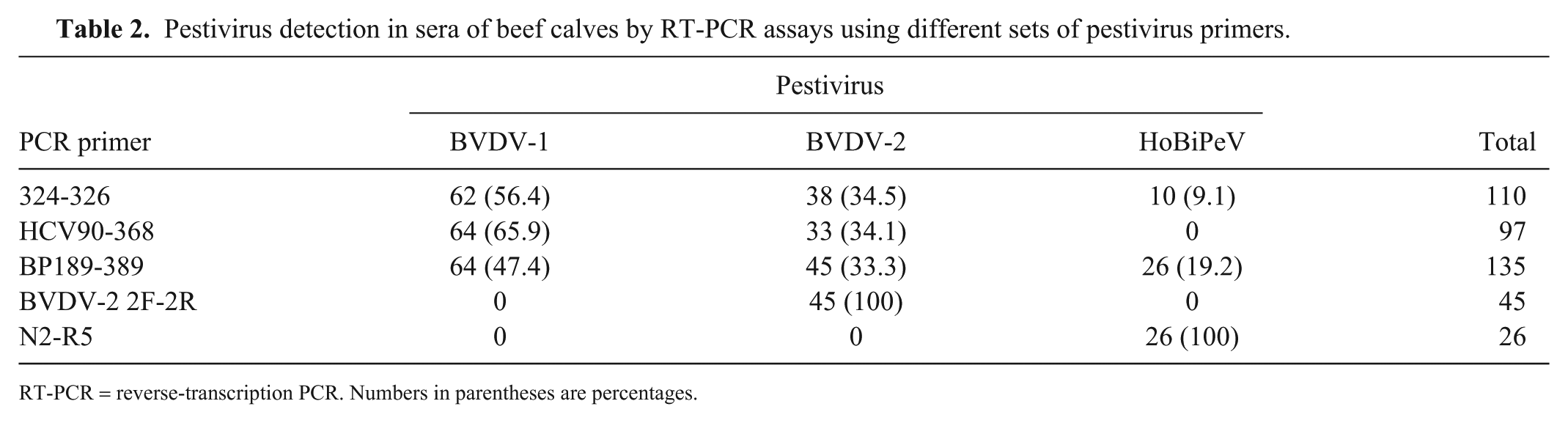
RT-PCR = reverse-transcription PCR. Numbers in parentheses are percentages.
PCR based on primers BP189-389 was also able to detect HoBiPeV in batches of FBS and in culture supernatants of MDBK cells inoculated with each of 8 Brazilian HoBiPeV (Table 3), confirming the spectrum of detection of these primers and their ability to detect HoBiPeV in different clinical specimens.
Detection of HoBiPeV in various clinical specimens by RT-PCR assays using different sets of primers.
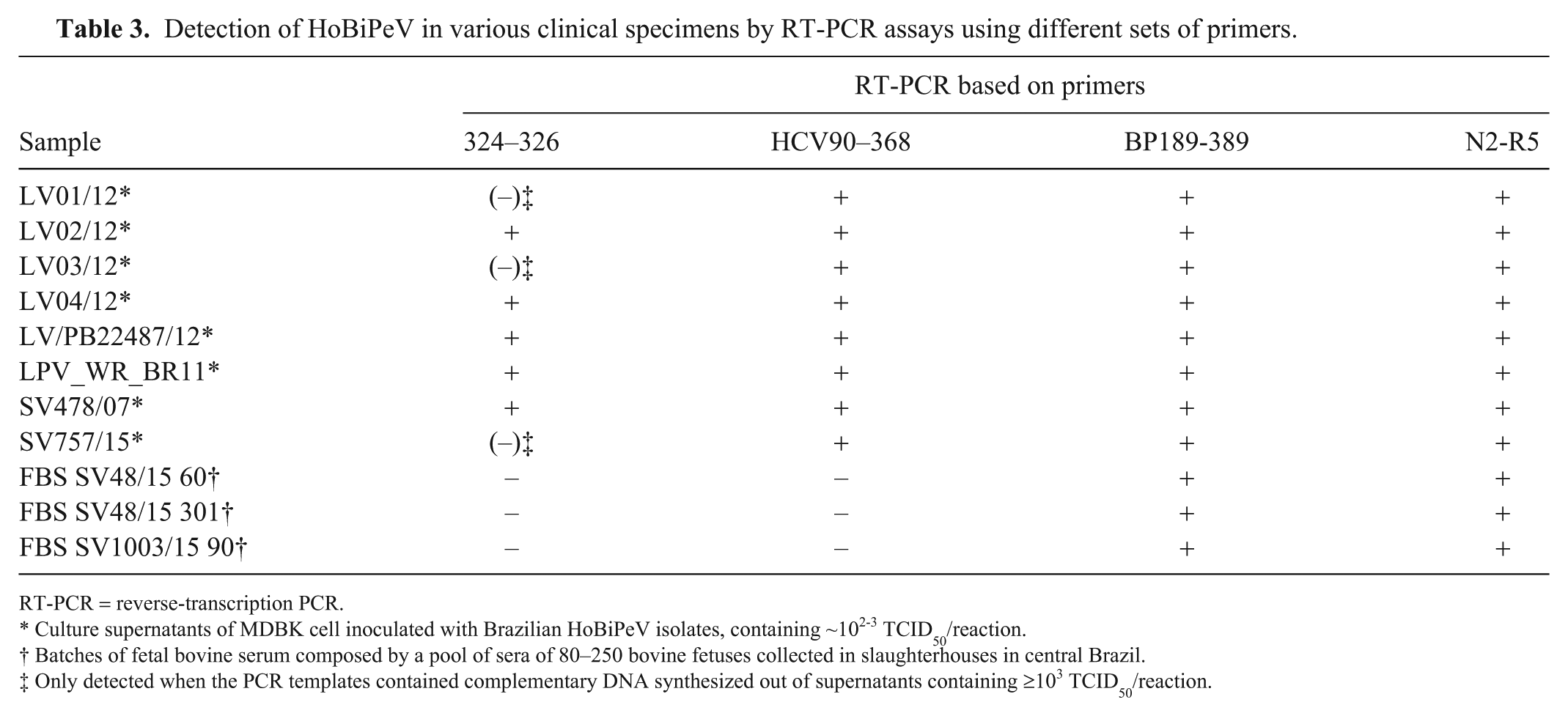
RT-PCR = reverse-transcription PCR.
Culture supernatants of MDBK cell inoculated with Brazilian HoBiPeV isolates, containing ~102-3 TCID50/reaction.
Batches of fetal bovine serum composed by a pool of sera of 80–250 bovine fetuses collected in slaughterhouses in central Brazil.
Only detected when the PCR templates contained complementary DNA synthesized out of supernatants containing ≥103 TCID50/reaction.
Primers HCV90-368 did not amplify any of the 26 HoBiPeV detected with pan–bovine pestivirus primers (BP189-389) and by primers N2-R5, whereas primers 324-326 detected 10 HoBiPeV. In a previous study, primers 324-326 were able to detect some HoBiPeV genomes in a gel-based PCR, 12 similar to the findings of our study. On the other hand, in a screening of North American lots of FBS for pestiviruses, only BVDV sequences were amplified by primers 324-326 and HCV90-368, even in the presence of large amounts of HoBiPeV genomes. 2 In that study, HoBiPeV sequences were detected only using the specific primers N2-R5. 2 Molecular screening of Brazilian lots of FBS for pestiviruses using RT-PCR with different primers reinforced these findings 13 (e.g., HoBiPeV detection was only achieved by a RT-PCR using primers N2-R5). Primers (2F-2R) have also been shown to specifically amplify BVDV-2 with higher sensitivity than other previously described primers, even in mixtures containing a large excess of BVDV-1 genomes. 13
Concerns about the efficacy of available BVDV tests to detect HoBiPeV viruses became especially relevant considering the risk of introduction of these novel pestiviruses into free areas.1,2 Given that the detection of HoBiPeV in different locations seems to be an ongoing process and their degree of genetic diversity is poorly known, it is expected that continuous surveillance and reformulation of molecular tools may be required for updated and efficient detection of these viruses. In this sense, our results are promising toward the use of primers BP189-389 in RT-PCR for pestivirus detection—including HoBiPeV in bovine samples—because they were able to detect an equivalent number of samples detected by a combination of 3 PCR assays, including sensitive detection of HoBiPeV. A conventional RT-PCR based on these primers may be especially useful for laboratories in which real-time PCR equipment is not available.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
FL Monteiro, R Weiblen, and EF Flores are recipients of CNPq fellowships (Brazilian Council for Research). JF Cargnelutti was the recipient of a PNPD/CAPES fellowship (Programa Nacional de Pós-Doutorado/Coordenação de Aperfeiçoamento de Pessoal de Nível Superior).
