Abstract
Rapid screening for enteric bacterial pathogens in clinical environments is essential for biosecurity. Salmonella found in veterinary hospitals, particularly Salmonella enterica serovar Dublin, can pose unique challenges for culture and testing because of its poor growth. Multiple Salmonella serovars including Dublin are emerging threats to public health given increasing prevalence and antimicrobial resistance. We adapted an automated food testing method to veterinary samples and evaluated the performance of the method in a variety of matrices including environmental samples (n = 81), tissues (n = 52), feces (n = 148), and feed (n = 29). A commercial kit was chosen as the basis for this approach in view of extensive performance characterizations published by multiple independent organizations. A workflow was established for efficiently and accurately testing veterinary matrices and environmental samples by use of real-time PCR after selective enrichment in Rappaport–Vassiliadis soya (RVS) medium. Using this method, the detection limit for S. Dublin improved by 100-fold over subculture on selective agars (eosin–methylene blue, brilliant green, and xylose–lysine–deoxycholate). Overall, the procedure was effective in detecting Salmonella spp. and provided next-day results.
Introduction
Salmonella spp. represent one of the leading causes of foodborne illness across all regions of the world. 13 A wide range of animals can be affected, including poultry and other birds, horses, cattle, pigs, sheep, goats, dogs, cats, and reptiles.8,24 Screening for Salmonella in clinical settings is critical given the high prevalence of multidrug resistance. 5 Animals fed raw meat diets can shed Salmonella intermittently without showing clinical signs. 2 In veterinary hospitals, animals that are shedding present significant biosecurity challenges, possibly because of the increased risk of infection associated with transportation, immunosuppressive therapies or other infections, as well as oral antimicrobial treatments. 24 Hospital-acquired cases may begin to shed subclinically or develop clinical disease that may be costly to treat and sometimes fatal, and may transfer to the home premises of the animal.3,4,9
Salmonellosis induces clinical patterns including gastroenteritis, bacteremia, enteric fever, and abortion depending on the host and serovar. 24 The cattle-adapted serovar Dublin is an emerging concern given the association of this serovar with invasive disease, multi-drug resistance, and opportunities for infection following survival in the environment.17,27 The organism also shows a high predilection for causing severe bacteremia in humans, with a higher case fatality rate than other serovars. 12 This serovar can be difficult to detect in view of intermittent shedding and poor growth in commonly used culture media.11,18,27 The sensitivity for detection of S. Dublin by culture has been estimated as 20% at best in feces from subclinically infected cattle. 18
One of the standard means of direct detection of Salmonella in food, animal, and environmental samples is through real-time (rt)PCR, which permits the rapid determination of the presence or absence of the bacterial genome. This procedure is typically performed on pre-enriched samples either directly from lysates 15 or on extracted DNA.7,22 Buffered peptone water (BPW) is a common pre-enrichment medium that is compatible with both of these approaches. Tetrathionate and selenite have been used for further selective enrichment of veterinary matrices. Given the presence of PCR inhibitors in the common sample types tested, extraction of DNA is preferred over lysates for maximizing sensitivity and accommodating a wider variety of broths and other matrices.14,16,21 Few commercial performance-tested rtPCR assays that include an extraction step are available. Routinely in veterinary laboratories, a single consolidated extraction procedure is sought for all pathogen detection assays and matrices in order to optimize workflow efficiency. Retaining both DNA and RNA allows for flexibility in use of the procedure across a wide range of assays.
We describe a protocol modification and matrix extension to the MicroSEQ Salmonella spp. detection kit (Applied Biosystems, Foster City, CA) that has been performance certified by 2 external organizations for Salmonella spp. detection in: (1) human foods and animal feed (certification 031001, AOAC Research Institute, Rockville, MD), 1 and (2) human food products, animal feed stuffs, animal feces, and environmental samples from the primary production stage (certification ABI 29/02-09/10, French Association for Standardization [AFNOR], La Plaine St. Denis, France). We modified several aspects of the published workflow. First, the recommended pre-enrichment broth of BPW was changed to Rappaport-Vassiliadis soya (RVS) selective enrichment broth for veterinary matrices in order to maximize recovery of serovars frequently encountered in animals (including S. Dublin). Second, the extraction method was changed from an automated enzymatic lysis-based extraction method to an automated total nucleic acid isolation procedure that can accommodate raw clinical samples such as feces and tissues and preserve RNA for detection of other potential targets in the same workflow. Applications of these modifications were assessed using matrices commonly encountered in veterinary testing and environmental surveillance of clinical settings. We focused our evaluation on those aspects of test performance likely to be affected by the proposed modifications, namely the limit of detection. To evaluate the modifications, the agreement between the combined selective enrichment–rtPCR method and culture alone was tested in both spiked samples and clinical specimens. Correlation between viability and threshold cycle (Ct) values was also assessed for different sample types.
Materials and methods
Sample collection and enrichment
Veterinary clinical specimens were submitted for routine testing to the Cornell Animal Health Diagnostic Center (Ithaca, NY). Clinical specimens were from a wide variety of animals including cattle, horses, dogs, birds, goats, deer, and reptiles. All samples tested were coded for blinding purposes, either internally by supervisory staff or by shipment to the Food and Drug Administration’s Veterinary Laboratory Investigation and Response Network (FDA Vet-LIRN) for external blinding.
Environmental samples were collected by wiping surfaces (recently cleaned, disinfected, and dried) with a sterile 4 × 4 gauze pad or commercial cloth cleaning pad (Swiffer, Procter & Gamble Professional, Cincinnati, OH) and placing the pad into sealed bags with ~5 mL of sterile evaporated skim milk to preserve viability. On the same day (or next day for samples shipped with cold packs), 45 mL of RVS broth was added to the bags and incubated for 20–24 h at 40–44°C in ambient air. Feed and water samples were pre-enriched as a ratio of 1:10 in BPW followed by separate enrichments in both RVS and Muller–Kauffmann tetrathionate–novobiocin (MKTTN) broth with each of those broths plated. Fecal and cloacal swabs submitted in transport media (Amies with or without charcoal) were added to 9 mL of RVS broth and incubated 20–24 h at 40–44°C in ambient air. For fecal material, intestinal scrapings, intestinal contents, or other enteric tissue, ~1 g of material was added to 9 mL of RVS broth and incubated as for the clinical swabs.
Confirmatory subcultures were performed on all samples by subculturing the incubated RVS broths onto eosin–methylene blue (EMB), brilliant green (BG), and/or xylose–lysine–deoxycholate (XLD) agar plates and incubating overnight at 33–37°C. As well, a 1-mL aliquot of the incubated RVS broth (and the incubated MKTTN broth for feed samples) was transferred to a 2-mL tube for screening by rtPCR.
Characterization of isolates
Colonies grown on selective agars (EMB, BG, or XLD) that showed the appropriate colony characteristics were confirmed as belonging to the Salmonella genus by matrix-assisted laser-desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry (Microflex Biotyper, Bruker Daltonics, Billerica, MA). Scores ≥2.0 were considered confirmatory (out of a maximum of 3). Any isolate with a score of <2.0 was subcultured to a nonselective agar and repeated. Colonies with subsequent scores of 1.7–1.999 were subjected to further biochemical tests according to standard procedures to confirm or rule out Salmonella. Any colonies scoring <1.7 were considered negative by culture. Following confirmation by MALDI-TOF, serogrouping was performed by slide agglutination using commercial antisera (Statens Serum Institut, Copenhagen, Denmark; Difco, BD Diagnostics, Sparks, MD). MALDI-TOF–confirmed clinical Salmonella isolates were sent to a national reference laboratory (National Veterinary Services Laboratories, Veterinary Services, Animal Plant Health Inspection Service, U.S. Department of Agriculture Services, Ames, IA) for complete serotyping by standard techniques. 6 Serovar confirmation of isolates obtained from spiked samples was performed by rtPCR according to a published method for S. Dublin 20 and a commercial kit for S. Typhimurium (TaqMan Salmonella Typhimurium Assay, Applied Biosystems).
Spiking studies
Salmonella spiking experiments were performed on the matrices listed below using fresh cultures prepared on blood agar plates that had been incubated for 18–24 h prior to spiking. Based on nephelometer readings calibrated with a 0.5 McFarland certified standard (verified by routine calibration against plate counts), the cultures were serially diluted in phosphate-buffered saline (PBS) prior to adding to the homogenized matrix. For environmental matrix spiking studies, poultry environmental samples were collected from a farm known to be free of Salmonella spp. as described above for environmental samples. Cultures from 8 recent clinical isolates were added to these environmental samples at final dilution targets of 10, 100, and 10,000 cfu/mL.
Bovine intestinal scrapings were obtained from an ~0.6 m section of a non-diarrheic adult dairy cow (that was euthanized because of a uterine laceration with peritonitis), combined with contents, and homogenized with enough PBS to create a 50-mL slurry to be used as the matrix. An aliquot of the matrix was determined to be free of Salmonella spp. using the enrichment–rtPCR method outlined herein including plating of the enrichment broth. The remaining matrix was frozen at −80°C until spiking. Cultures of serovars Dublin and Typhimurium were added to the matrix to achieve target concentrations of 1, 5, 10, 100, or 1,000 cfu/mL. Each of the samples was split into aliquots for either sending to FDA Vet-LIRN and return shipment of blinded samples or testing in-house after being held at 4°C. The outside temperature at the time of shipping ranged from −20°C to 3°C over the course of 3 d.
We additionally tested 12 Salmonella-spiked canine fecal samples from a network proficiency test administered by FDA Vet-LIRN in collaboration with the FDA Moffett Campus Proficiency Testing Laboratory. After testing for stability and homogeneity, samples were spiked at the FDA Moffett Center with either S. enterica serovar Typhimurium or Heidelberg at 0, 1, 5, or 10 cfu/g, coded, and then sent to our laboratory.
Nucleic acid extraction
An undiluted aliquot was taken from the incubated RVS broth and added directly to bead tubes containing lysis/binding solution for the alternative automated total nucleic acid isolation kit (MagMAX total nucleic acid isolation kit [AM1840], Applied Biosystems) or proteinase K for the recommended kit used in previous performance evaluations for food testing (PrepSEQ Nucleic Acid Extraction Kit for Food and Environmental Testing [4480466], Applied Biosystems). Using either 250 µL or 175 µL of the suspension for MagMAX or PrepSEQ extraction, respectively, the nucleic acid was then purified using the manufacturer’s recommendation for automated preparation in the 96-well format, including bead beating for the MagMAX kit (Mini-Beadbeater-96, BioSpec, Bartlesville, OK). A 96-well robot (MagMAX/Kingfisher model 710, Thermo Fisher Scientific, Waltham, MA) was used for both methods with the recommended program for each kit. The yield of eluted nucleic acid from both kits was ~80 µL. Positive extraction controls consisted of saturated overnight cultures of S. enterica serovar Cerro grown in RVS broth and frozen at −80°C in aliquots at dilutions of 10−4, 10−5, and 10−6 in the same broth. Acceptable ranges were established for these controls based on 2 standard deviations above and below the mean Ct for each dilution. Negative extraction controls consisting of uninoculated RVS broth were also prepared at the time of inoculation; one such control was set up for every 20 samples.
Real-time PCR
An aliquot of 30 µL of nucleic acid from each sample was placed into a tube provided in the Salmonella spp. detection kit and tested using the recommended platform (7500-FAST with SDS 1.4.1 software, Applied Biosystems) and cycling conditions. Internal positive amplification controls are included as part of the lyophilized reaction mix (run by multiplex). A negative amplification control (provided with the kit in a separate tube) was included for each run. If both of the targets failed to amplify, elutions were diluted at 1:5 and repeated as recommended by the manufacturer. The fractional positive level for spiked samples was defined as 50% ± 25% of tests positive (Guidelines for the Validation of Analytical Methods for the Detection of Microbial Pathogens in Foods and Feeds, https://goo.gl/Q33Sjs).
Data analysis
Data were analyzed using statistical analysis software (Prism, GraphPad Software, La Jolla, CA). Standard curves were analyzed by linear regression, and overall differences between methods were tested by the Wilcoxon signed rank test. Percent agreement was calculated using the Cohen kappa coefficient.
Results
Comparison of nucleic acid extraction methods
In order to evaluate the effect of modification of the nucleic extraction method on rtPCR results, 6 replicates of S. enterica serovar Enteritidis spiked at 1 cfu/mL into poultry farm environmental samples, serially diluted following enrichment, and extracted by both methods, were tested (Fig. 1A). The overall median difference in Ct value was 1.513 (p = 0.03). The slopes for each method were not statistically different, indicating comparable efficiencies. The fractional positive level was the 10−6 dilution (4 of 6 replicates positive by MagMAX and 2 of 6 positive by PrepSEQ). Dilutions beyond the fractional positive level were not detected for either method. Based on these results, the MagMAX extraction method was considered to be an equivalent alternative to PrepSEQ.
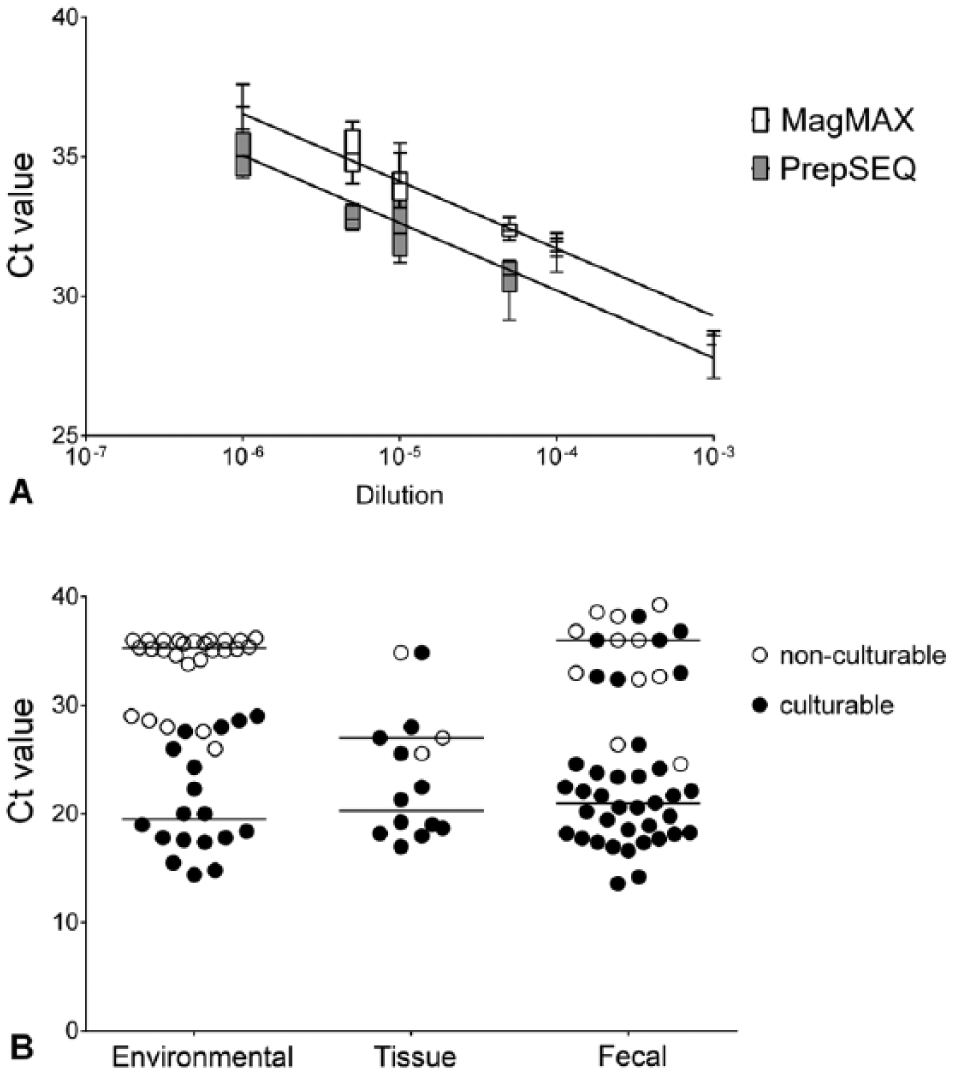
Threshold cycle (Ct) values from endpoint analysis of 2 different nucleic acid extraction methods
Viability analysis
Ct values from enrichment–rtPCR were compared with confirmatory culture in aggregate for each matrix in order to determine if Ct values could be used to predict viability (Fig. 1B). No environmental samples with Ct values >29 were viable. No such comparison could be made for other sample types. Viability of 3 nucleic acid elutions from extracted positive cultures was assessed by further culture in order to determine if the extraction process inactivated the bacteria; no growth was seen.
Environmental samples
Isolates from 8 different origins were spiked into environmental matrix from a poultry farm at 3 dilutions and tested by selective enrichment–rtPCR and confirmatory subculture (Table 1). Unspiked samples were negative by both methods. For all serovars tested, except for Dublin, the 10 cfu/mL spike was detected by both methods, corresponding to an average Ct value of 15 (range: 14–17). For S. Dublin, only the 10,000 cfu/mL spike was detectable by confirmatory subculture. This spike level had a Ct value of 29, and the non-culturable 100 cfu/mL spike level had a Ct of 35.1.
Results of a selective enrichment broth with real-time PCR assay for Salmonella, expressed as threshold cycle (Ct) values, for recent clinical isolates spiked into an environmental matrix from a poultry farm.
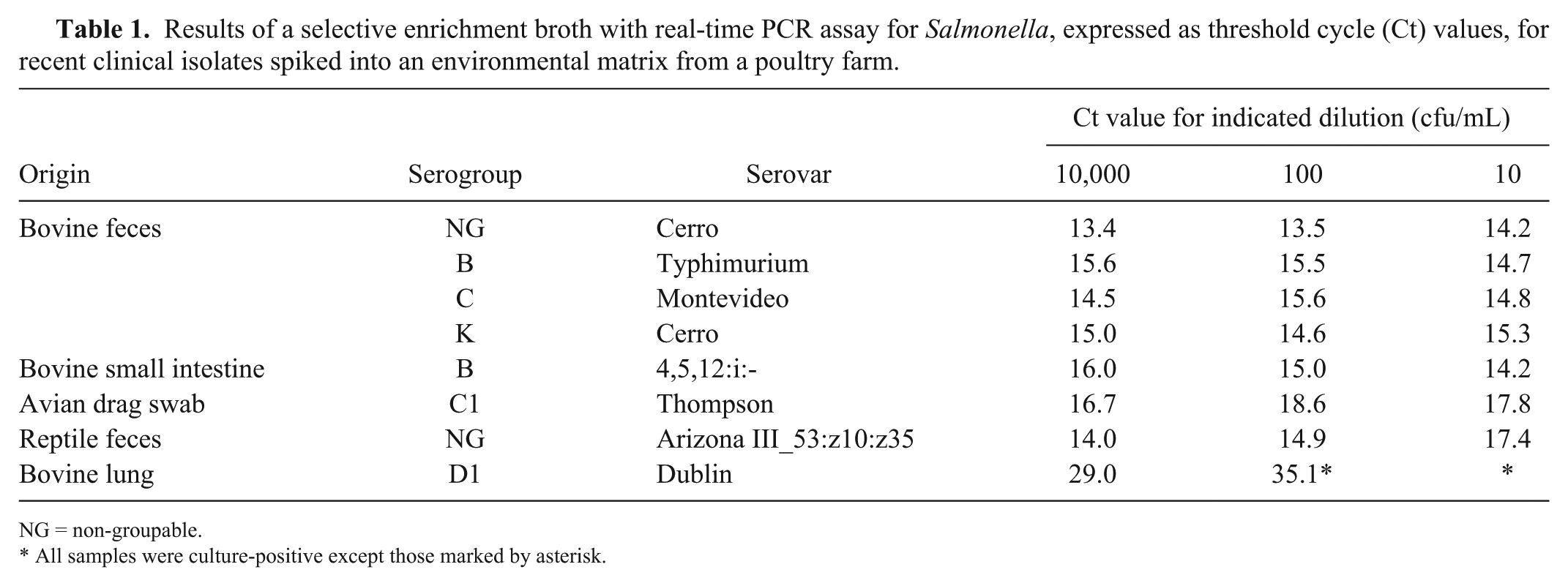
NG = non-groupable.
All samples were culture-positive except those marked by asterisk.
A total of 81 environmental samples were tested (Table 2). These included samples collected from clinical treatment areas of a large animal teaching hospital and the autopsy room following environmental contamination from a Holstein bull with diarrhea confirmed to be shedding Salmonella enterica serovar 4,5,12:i:- on direct culture 4 d after hospitalization in October 2013. Samples were also received from other sites for routine environmental surveillance. The serovars detected from environmental samples were 4,5,12:i:-, Berta, Cerro, Newport, Oranienburg, and Typhimurium var. 5-. In addition to environmental samples, we tested 8 water (pond and sink) samples and 1 bedding sample from a dairy farm with an ongoing outbreak of salmonellosis. The bedding sample was positive by both methods (typed as serovar Typhimurium var. 5-). One of the 8 water samples was detected by rtPCR only (Ct 39.71 with enrichment and 35.97 without enrichment); all water samples were negative by culture. The overall kappa correlation between testing methods for environmental samples was 0.335 (95% confidence interval [CI]: 0.186–0.483).
Results for environmental and clinical specimens tested with selective enrichment with real-time (rt)PCR compared to culture only.
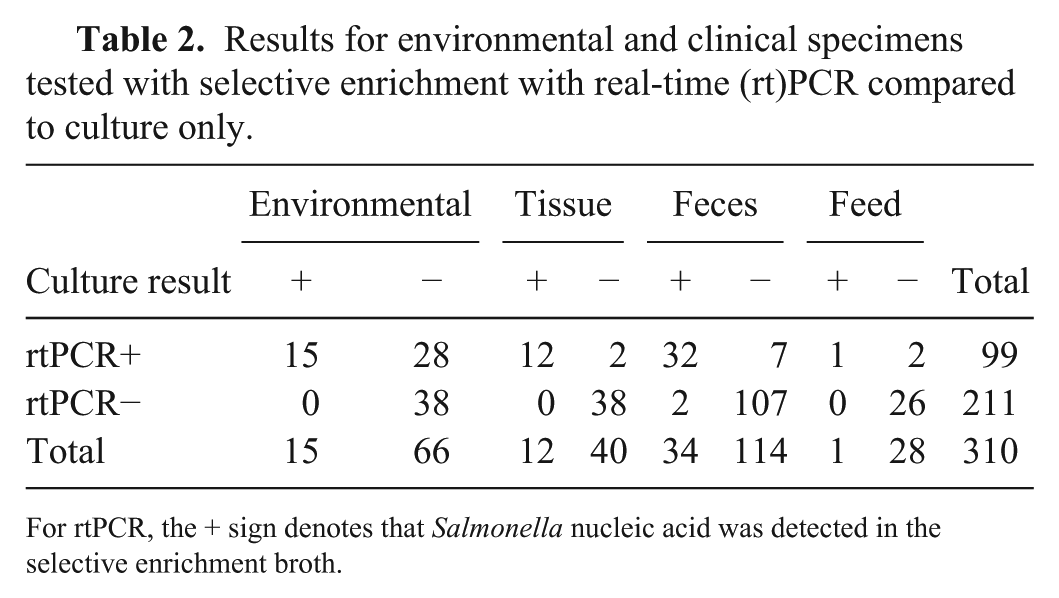

For rtPCR, the + sign denotes that Salmonella nucleic acid was detected in the selective enrichment broth.
Tissue samples
For the spiking study on bovine intestine, all results for testing of shipped versus held samples ultimately agreed except for the 100 cfu spike of serovar Typhimurium, which was culture positive when held at 4°C but not detected by culture after being shipped to and from FDA Vet-LIRN. Enriched clinical tissue samples (n = 52) submitted for routine testing were also tested by rtPCR (Table 2). These were primarily bovine intestinal scrapings, but also included enteric or cloacal samples from alpaca, chicken, clouded leopard, deer, duck, gazelle, goat, goose, hooded crane, horse, rabbit, raccoon, and turtle. The 2 samples producing discrepant results were jejunum samples from a cow and a New World alpaca. The serovars detected were Albany, Dublin, Havana, Heidelberg, Mbandaka, Meleagridis, and Typhimurium, and the correlation coefficient for tissues was 0.898 (95% CI: 0.759–1.000).
Fecal samples
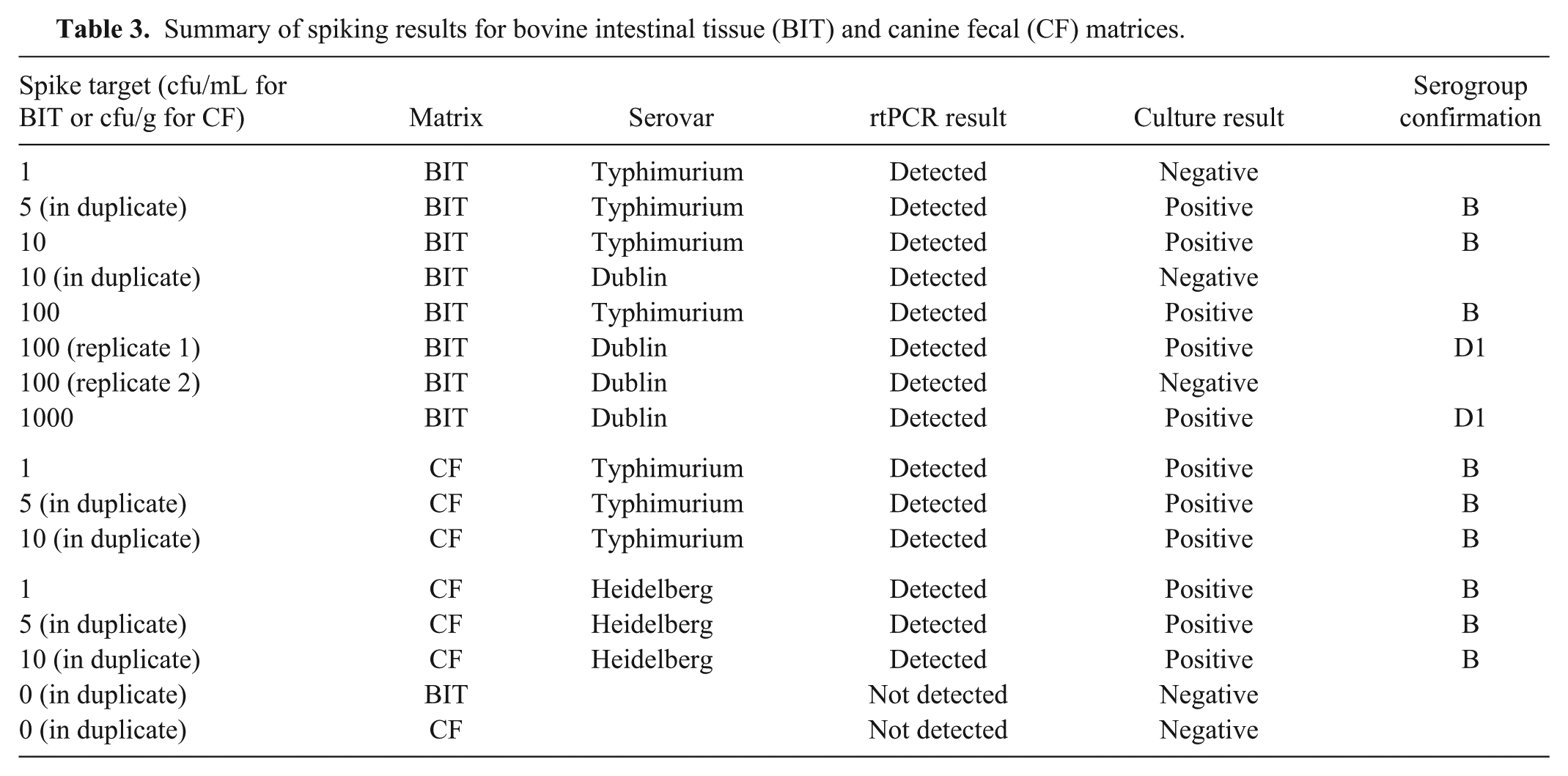
A total of 148 clinical fecal samples were tested (Table 2). These were primarily from cows, dogs, and horses. The serovars detected in fecal samples were Anatum, Cerro, Dublin, Holcomb, Montevideo, Muenster, Thompson, and Typhimurium, and the correlation coefficient for fecal samples was 0.837 (95% CI: 0.734–0.939). The 2 false-negatives on enrichment–rtPCR were from a dairy cow (Cerro) and a giraffe (Muenster). Internal control Ct values for these samples were 30.5 and 31.2. The Cerro sample could be detected at Ct 36 after testing a diluted nucleic acid extract. The 7 samples detected only by enrichment–rtPCR were from 5 dogs, 1 goat, and 1 cow, with Ct values of 24.6–39.3. Six of 36 (16.7%) horse fecal samples and 3 of 6 goat fecal samples tested had a failed internal control, although this was effectively managed by dilution. For spiked fecal samples, all results were correct by both methods (Table 3).
Summary of spiking results for bovine intestinal tissue (BIT) and canine fecal (CF) matrices.
Feed samples
A total of 29 feed samples were tested by both methods. These included raw meat, frozen feeder rodents, milk replacer, colostrum, hay, and various types of chow. One sample (canola meal) was detected by both methods (serotyped as Havana and Meleagridis). Two samples (1 chow and 1 milk) were detected by enrichment–rtPCR only (Ct: 39.02 and 39.46). Of the 2 enrichment procedures performed on the chow, only the BPW/RVS-enriched sample produced a Ct result. All other samples were negative by both methods. The kappa correlation for feed was 0.473 (95% CI: 0.126–1.000).
Discussion
There are many different procedures for isolation of Salmonella from different matrices. Variables included in the choice of Salmonella culture protocols include the expected level of Salmonella, the amount of moisture in the sample, the presence of competitive microbial flora (feces, intestinal samples, autopsy tissues), and whether the sample matrix endured any heating or drying conditions during processing (grain, dog treats, dry dog food/chow) that likely injure or damage the Salmonella. Depending on the matrix to be cultured, a pre-enrichment step may be needed before the final selective enrichment step in the culture protocol. The rationale for this pre-enrichment step lies in the fact that during manufacture, and perhaps as a result of storage conditions, Salmonella may be “injured” or damaged and may need to be revived so as to increase their growth rate and better detect them by culture. 26 There are a number of pre-enrichment broths that have been used by microbiologists to engender repair of injured Salmonella. BPW is commonly used as a pre-enrichment broth in food microbiology. 26 For our study, pre-enrichment steps with BPW were limited to food matrices and water. Veterinary samples with competitive microbial flora were selectively enriched only in order to inhibit competitive flora while minimizing turnaround time and expense.
We determined that RVS enrichment is compatible with automated nucleic acid extraction for Salmonella spp. testing of veterinary matrices using either extraction kits. Although PrepSEQ gave significantly lower Ct results, all aspects of test performance with MagMAX were acceptable. The lower Ct values from PrepSEQ are likely the result of the larger volume of enrichment culture used in the automated extraction procedure. PrepSEQ is designed for purification of DNA from enrichment cultures, whereas MagMAX isolates both DNA and RNA from most types of samples encountered in veterinary clinical submissions including blood, feces, fluids, tissues, and cultures. For an efficient extraction workflow in clinical laboratories, the versatility of being able to extract samples for multiple tests and matrices together is essential. Future work can be directed at evaluating this extraction workflow on other combinations of matrices and pathogen targets.
The method described herein improved the limit of detection (LOD) for S. Dublin. All serovars were reliably detected down to the 1 cfu/mL spike level for enriched environmental samples, with the exception of Dublin, which could only be detected at 10 cfu/mL by enrichment–rtPCR (1,000 cfu/mL by culture). In a suspension of spiked bovine intestinal scrapings, 1 or 5 cfu of S. Typhimurium were detected by rtPCR or culture, and 10 or 1,000 cfu were detected for S. Dublin. For fecal samples (spiked with Typhimurium or Heidelberg), 1 cfu/g was detected by both methods. The LOD established for the unmodified method for the MicroSEQ rtPCR system has been reported as 1,000 cfu/mL in food matrices. 1
Although we only investigated a limited number of S. Dublin samples, these results merit further examination, as the comparison between the results for S. Dublin and the other serovars tested suggests that conventional methods and enrichment–rtPCR both have difficulties in detecting this serovar, though selective enrichment–rtPCR did have greater detection sensitivity. This finding is in contrast to a previous study comparing bacteriologic culture reference methods to rtPCR performed on enrichment media, which did not show an improvement by rtPCR in detection of S. Dublin. 11 The epidemiology of S. Dublin infection includes the development of persistent infections without clinical illness, or carriers. 25 The site of persistence can be either intestinal or in the mammary gland. 18 Some of the limitation in methods for detection of S. Dublin infection has been assumed to be related to intermittent shedding and/or bacteremia in these infected carrier animals because of colonization of the S. Dublin organisms in lymphoid or other non-enteric tissues, which could impact antigen-based testing applied to fecal or tissue specimens. 18 However, if the bacteria are actually present in some matrices such as feces at very low numbers, a detection methodology able to improve the detection sensitivity by a factor of 102 could significantly improve infection management. Previous studies using fecal culture alone for the detection of carrier animals demonstrated very low detection sensitivity, ranging from ~3 to 17%.10,19 Serologic methods have been developed and used to enhance detection of carrier animals. However, the persistent infection status of an individual animal cannot be determined by solely using serology without repeated testing over many months, given that animals that are infected and clear the infection will routinely have detectable antibodies for ~5–7 mo. 18 Highly sensitive molecular testing combined with serologic techniques that would allow veterinarians and cattle owners to accurately classify seropositive animals as carrier or uninfected would assist with the cost-effective control of this infection within herds.
The internal control for the rtPCR assay effectively indicated the presence of rtPCR inhibitors, highlighting the importance of using an internal control. Because the internal control for this method is introduced at the amplification stage of the workflow, we also used a series of positive extraction controls on each plate to ensure that extraction was successful. As expected, feces was the matrix containing the most inhibitors. Fecal PCR inhibition is likely caused by a combination of bile salts, lipids, complex polysaccharides, urate, and/or hemoglobin. 23 Possibly because of the lack of a gallbladder, equine fecal samples are particularly prone to inhibition. Low levels of inhibitors that may not affect the internal control are a concern for equine fecal samples, which suggests the use of repeated sampling schemes in large animal hospital settings.
Detection sensitivity as assessed by correlation between enrichment–rtPCR and culture alone was quite good for clinical specimens (tissue and feces). For environmental and feed samples, the correlation was lower, presumably because of the fact that surfaces had been cleaned and disinfected, and some feed may have been irradiated prior to sampling. Some of the surfaces that were culture-positive included office spaces, highlighting the importance of biosecurity for non-clinical areas. Assessment of viability based on Ct values was informative for environmental samples but not successful for clinical specimens. For this reason, enrichment–rtPCR should be considered a screening tool for detection of Salmonella spp. in animal tissues and feces. Our results using the modified workflow are consistent with interpretation guidelines for food testing, namely that rtPCR-negative enrichments may be considered final and do not need to be confirmed by culture. Overall, the data do not support the use of Ct cutoffs for confirmatory culture. Any enrichment producing an amplification curve with a Ct value should be assessed for viability and further characterized by culture and subsequent testing; our recommended guidelines for interpretation are outlined in Table 4.
Selective enrichment with real-time PCR interpretation guidelines recommended based on threshold cycle (Ct) results.
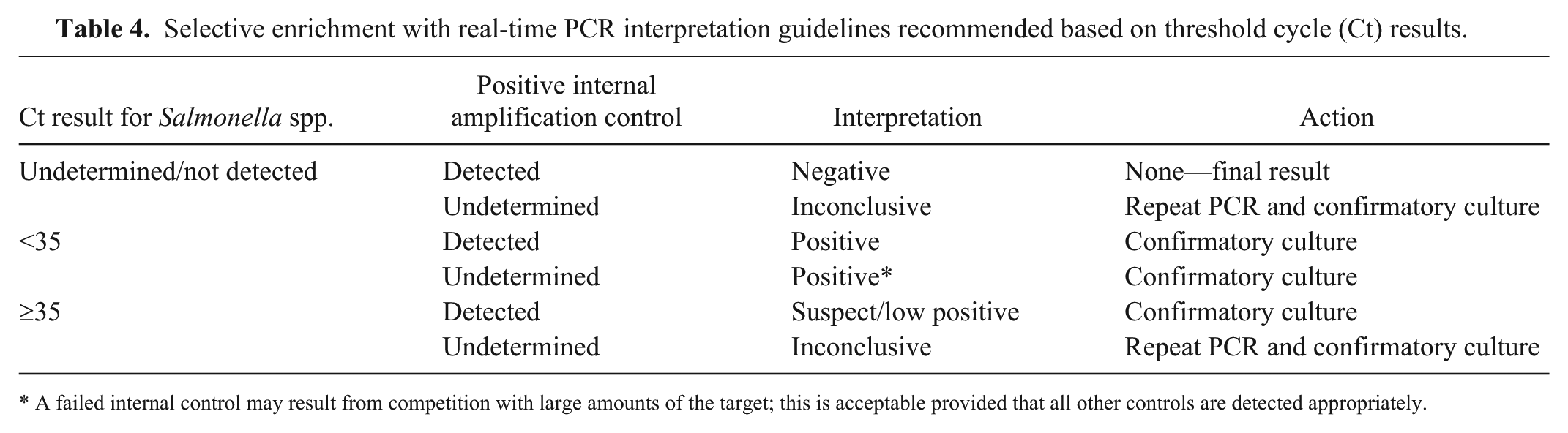
A failed internal control may result from competition with large amounts of the target; this is acceptable provided that all other controls are detected appropriately.
The workflow described herein allows the use of retained enrichment broths for other testing such as culture and evaluation of isolates. Phenotypic antimicrobial susceptibility testing generally has the ability to detect emerging antimicrobial resistance before the molecular or genetic basis of that resistance may be fully understood. Therefore, there may be lag time in the ability to use molecular methods for comprehensive antimicrobial resistance evaluation for clinical treatment or surveillance purposes. Maintaining access to culture isolates may provide the materials necessary for molecular investigations into changes in antimicrobial resistance or pathogenicity.
The detection of Salmonella spp. using a selective enrichment prior to rtPCR can be used on a variety of veterinary samples from feed, water, environmental samples, feces, and intestinal scrapings. The method can be incorporated into a high-throughput workflow using an extraction method commonly used for multiple matrices and targets, allowing for next-day results. Enrichments with any amount of rtPCR amplification should be cultured for assessment of viability and further characterization.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was funded (FOA PA-13-244) and performed in collaboration with the Food and Drug Administration’s Veterinary Laboratory Investigation and Response Network (FDA Vet-LIRN) under grant 1U18FD005144-01.
