Abstract
In September 2015, a large outbreak caused by epizootic hemorrhagic disease virus (EHDV) was identified in Israeli dairy and beef farms. The main clinical signs were reduced milk production, weakness, drooling, lameness and recumbency, fever, slight erythema of nasal and oral mucosae, weight loss, and abortion. Dyspnea, cachexia, and death were observed less frequently. The clinical diagnosis was confirmed by ELISAs and EHDV-specific real-time reverse transcription PCR (RT-rtPCR), followed by conventional RT-PCR of the VP2 gene and sequence analysis. According to the sequence and phylogenetic analysis of theVP2 gene, the 2015 Israeli EHD outbreak was caused by EHDV-6, which was found not only in clinically ill cattle, but also in aborted fetuses.
Epizootic hemorrhagic disease (EHD) is an infectious, non-contagious viral disease transmitted by insects of the genus Culicoides; it affects wild and domestic ruminants, primarily white-tailed deer (Odocoileus virginianus) and cattle 6 (https://goo.gl/U1JHsf). The disease is caused by species Epizootic hemorrhagic disease virus (EHDV; genus Orbivirus, family Reoviridae). It is a double-stranded RNA virus with a linear genome of 10 segments, coding 7 structural (VP1–VP7) and 3 or 4 nonstructural (NS1–NS4) proteins. It shares many morphologic and structural characteristics with the other members of the genus, such as bluetongue virus (BTV), African horse sickness virus, and equine encephalosis virus 4 (https://goo.gl/U1JHsf). Like BTV, the primary determinant of serotype specificity is the outer capsid VP2 protein; at least 7 distinct serotypes have been identified (https://goo.gl/q3QzbL).
EHDV has been isolated from cattle worldwide, including Africa, North America, Australia, Japan, and, during the last decade, the Mediterranean basin. 6 Severe clinical signs also have been observed in cattle infected by the Ibaraki strain (EHDV-2) in Japan 5 and EHDV-6, viruses similar to EHDV-6/EHDV-318 in the countries of the Mediterranean basin.6,9 Many cases of EHD were observed in the United States in cattle in the summer and fall of 2012 caused by the North American serotypes EHDV-1, EHDV-2, and EHDV-6. 7
In Israel, EHD was suspected in 1951, 3 but was first confirmed as an EHDV-7 outbreak in 2006. 11 The disease was reported in 83 dairy herds, each comprising 380 cows on average, and 22 beef herds. 2 Observed clinical signs—not all in any individual animal—included reduced milk production, anorexia, reduced rumination, fever, weakness, clear-to-purulent nasal discharge, drooling, nasal and labial redness with scaling, tongue swelling, cyanosis and erosions, conjunctival hyperemia, petechiae on tips of the lingual and buccal papillae, hemorrhages in the conjunctiva, dyspnea, emphysema, stiff gait, lameness, and muscle tremors. 12 Mortality was very low. Duration of clinical disease was at least 1 wk, but has still to be defined. No cases have been seen in adjacent sheep and goat farms (ProMed, archive 20060913.2593 and 20060917.2651).
The recent outbreak of EHD was recognized clinically in late September 2015, simultaneously in dairy farms in 2 different geographic loci in northern Israel and in the area around the Dead Sea. The infection quickly spread through the country, from Galilee in the north to the Arava Desert in the south (Fig. 1). The most prevalent clinical signs observed during this outbreak were reduced milk production, weakness, drooling, lameness and recumbency, fever, light erythema of nasal and oral mucosae, weight loss, and abortion. Dyspnea, cachexia, and death were observed less frequently.
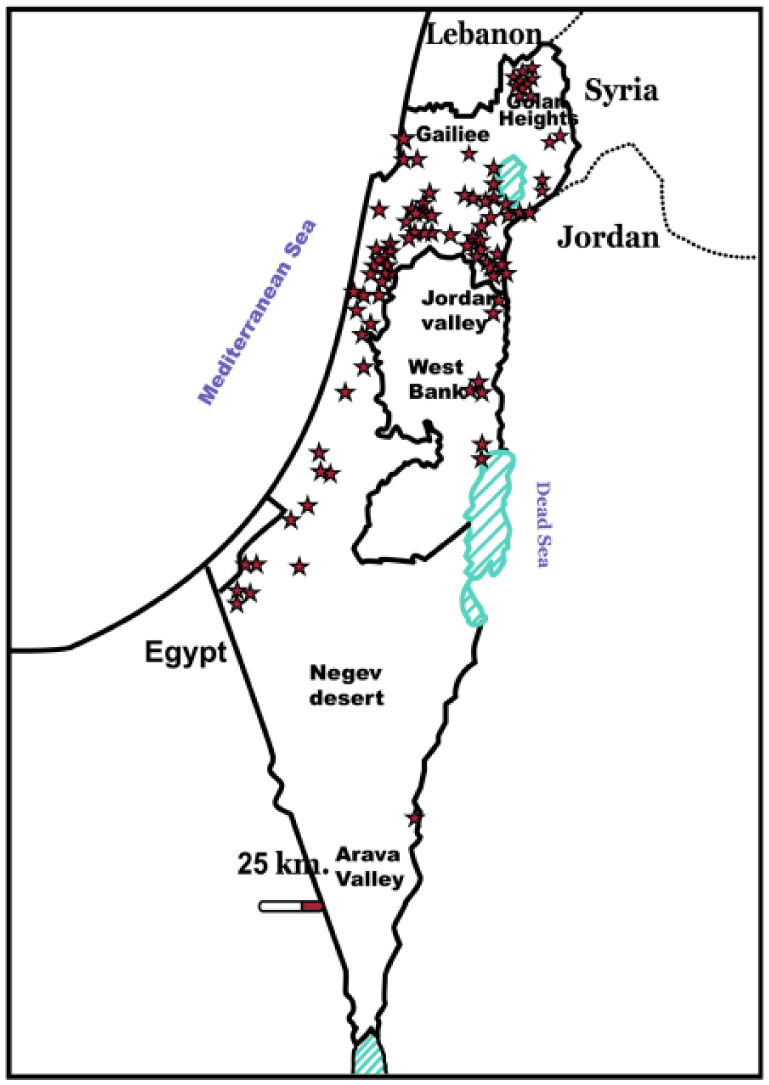
Geographic distribution of epizootic hemorrhagic disease virus–positive cattle farms confirmed by RT-rtPCR and/or antibody ELISAs.
Viral RNA was extracted (Invisorb Spin Virus RNA Mini kit, STRATEC Molecular, Berlin, Germany) from EDTA–anticoagulated whole blood, spleen, brain, placenta, pooled organs (lung, kidney, liver) from aborted fetuses, and homogenized embryonated chicken eggs. All samples were tested by real-time (rt) reverse transcription (RT) PCR (RT-rtPCR) systems (Epizootic Hemorrhagic Disease Virus Real-Time PCR kit, LSI VetMAX, Lissieu, France) and RT-rtPCR 11 using a commercial kit (SuperScript III Platinum One -Step qRT-PCR System, Invitrogen, Carlsbad, CA) that specifically identify EHDV, but do not provide information about serotype of the viruses. Conventional RT-PCR for virus confirmation, followed by sequence analysis, was performed with a RT-PCR kit (One-Step RT-PCR kit, Qiagen, Hilden, Germany), together with several in-house primer systems based on the VP2 gene for identification of EHDV-2, -6, -7, and -8 (Table 2). Sequence analyses revealed that the outbreak was caused by EHDV-6. Additionally, 5 sets of overlapping primers for full-length sequencing of the EHDV-6 VP2 gene also were developed and used for sequencing of virus isolates (Table 2). Nucleotide and amino acid sequences were aligned and phylogenetically analyzed (https://goo.gl/cGEvL7; https://goo.gl/XBny1).
From September 2015 through March 2016, field samples from cattle were positive for EHDV by RT-rtPCR. According to the clinical and laboratory findings, the peak of the outbreak occurred in October–November 2015. During the outbreak, 152 of 321 field samples (mostly whole blood samples) were positive. Five samples of spleen were positive from 6 dead animals suspected to be infected with EHDV. Four of the 5 positive spleen samples were from animals 3–10 mo of age, and 3 of 4 calves had epicardial hemorrhages. The first aborted fetus positive for EHDV was sampled and tested at the beginning of October, and the last one in mid-December 2015. Additionally, from September 2015 through January 2016, 11 of 30 aborted bovine fetuses were positive for EHDV by RT-rtPCR tests; 7 of 16 brain tissue samples and 6 of 13 placenta samples were positive, whereas no positive results were obtained from internal organs of aborted cattle fetuses. In 2 cases, both placenta and brain samples were positive. Four fragments of the VP2 gene, 2 of 272 bp and 2 of 266 bp, were amplified from aborted material (Table 1). The geographic distribution of positive bovine fetuses is shown in Figure 2.
Epizootic hemorrhagic disease virus–positive samples and/or isolates used in our study.
Sequenced using RNA from virus isolated in embryonated chicken eggs.
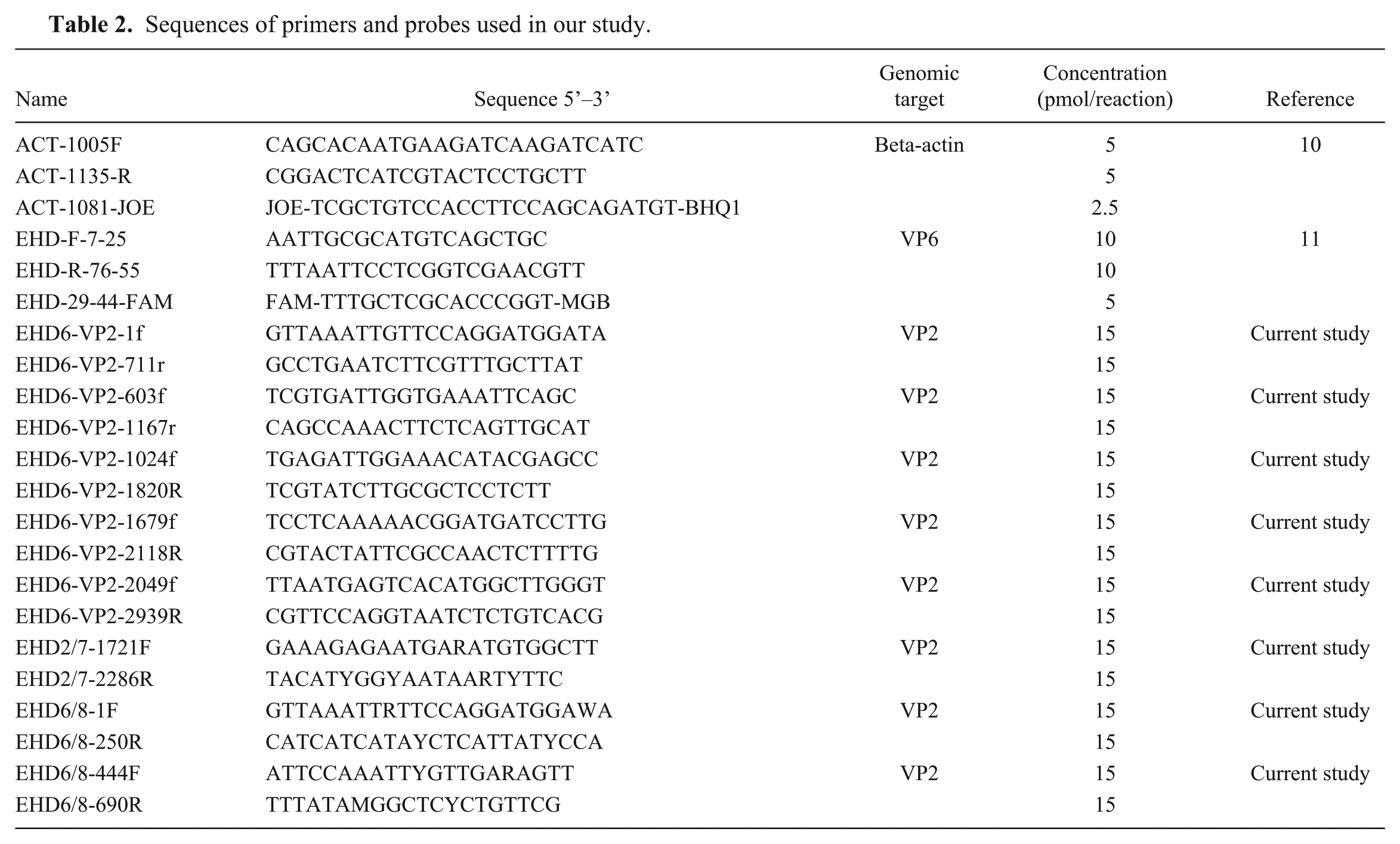
Sequences of primers and probes used in our study.
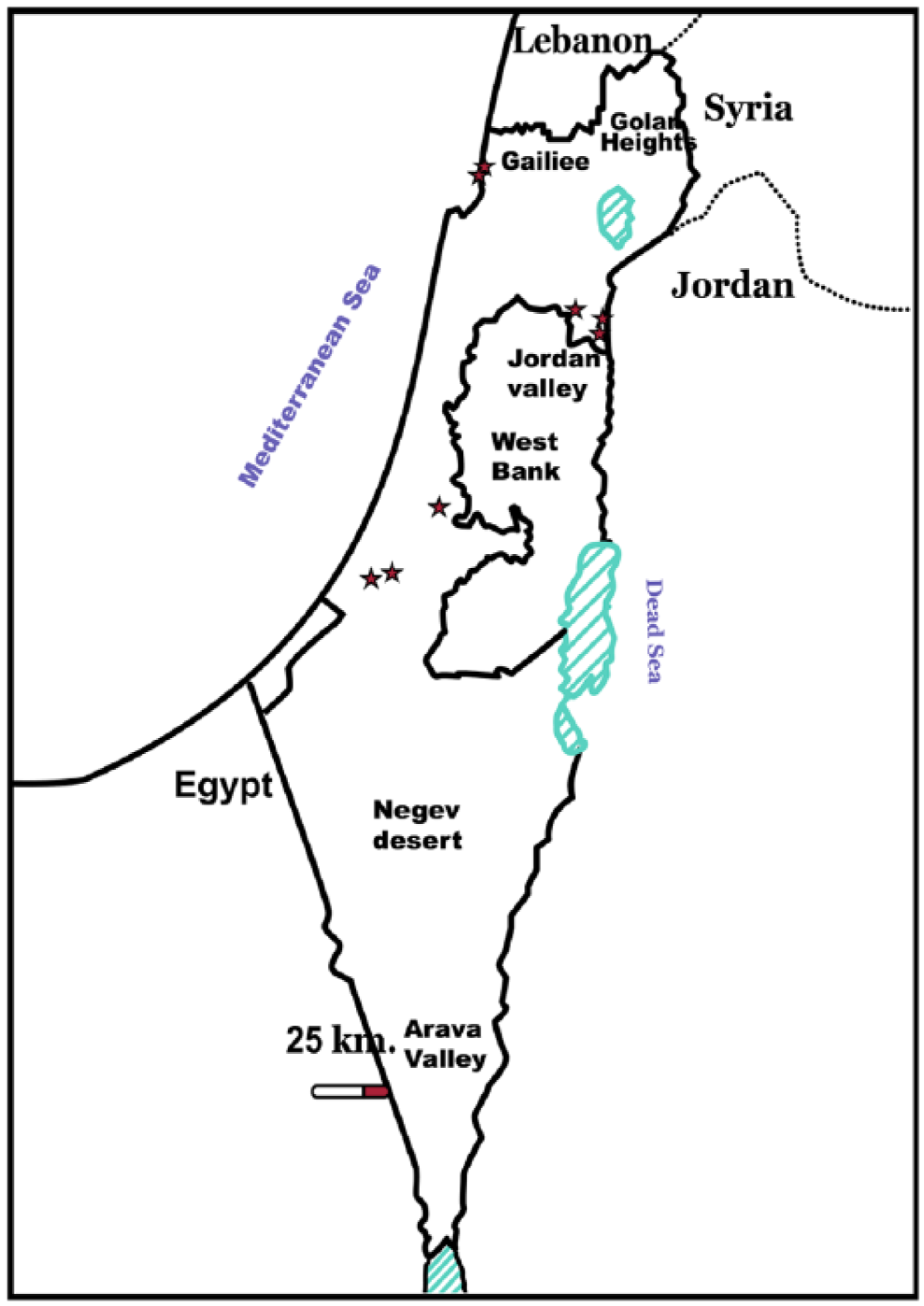
Locations of epizootic hemorrhagic disease virus–positive abortions in cattle.
The outbreak of 2015 resulted in milder clinical signs than observed during the 2006 outbreak, but the spread of EHDV was significantly wider than in 2006. 2 In the 2015 outbreak, EHDV was identified in 86 loci by means of RT-rtPCR (Epizootic Hemorrhagic Disease Virus Real-Time PCR kit, LSI VetMAX) 11 and/or antibody ELISA (LSIVet Ruminant EHDV Serum ELISA kit, LSI VetMAX; Fig. 1). Antibody ELISA was used because the cattle population was free of EHDV antibody >3 y previously.
Twelve viruses were isolated in embryonated chicken eggs and cell cultures,1,3 but incubation of the eggs required up to 9 d post-inoculation.
Two full-length sequences from 2 virus isolates (ISR-4487/15 and ISR-1975/2/15; Table 1), partial VP2 sequences from all other EHDV-6 isolates, and several selected field samples were analyzed but not uploaded to GenBank. Analogous regions of theVP2 gene (nucleotide positions 1–2,956) from 2 virus isolates (ISR-4487/15 and ISR-1975/2/15) shared 100% identity (Fig. 2). Two brain and 2 placenta samples from aborted cattle fetuses were positive in conventional PCR for VP2 genes (Table 1).
According to the phylogenetic analyses, all viruses identified in this Israeli EHD outbreak clustered with EHDV-6 of Asian/African origin; the prototype strain is strain 318 from Bahrain (isolate BAR1983/01). Pairwise analysis showed that the ALG2006/02(R10/06-1) strain from Algeria was the EHDV-6 strain closest to Israeli strains, with 96.3% identity; the next closest was the MOR2006/05(A113/06 1) strain from Morocco, with 96.2% identity (Fig. 3).
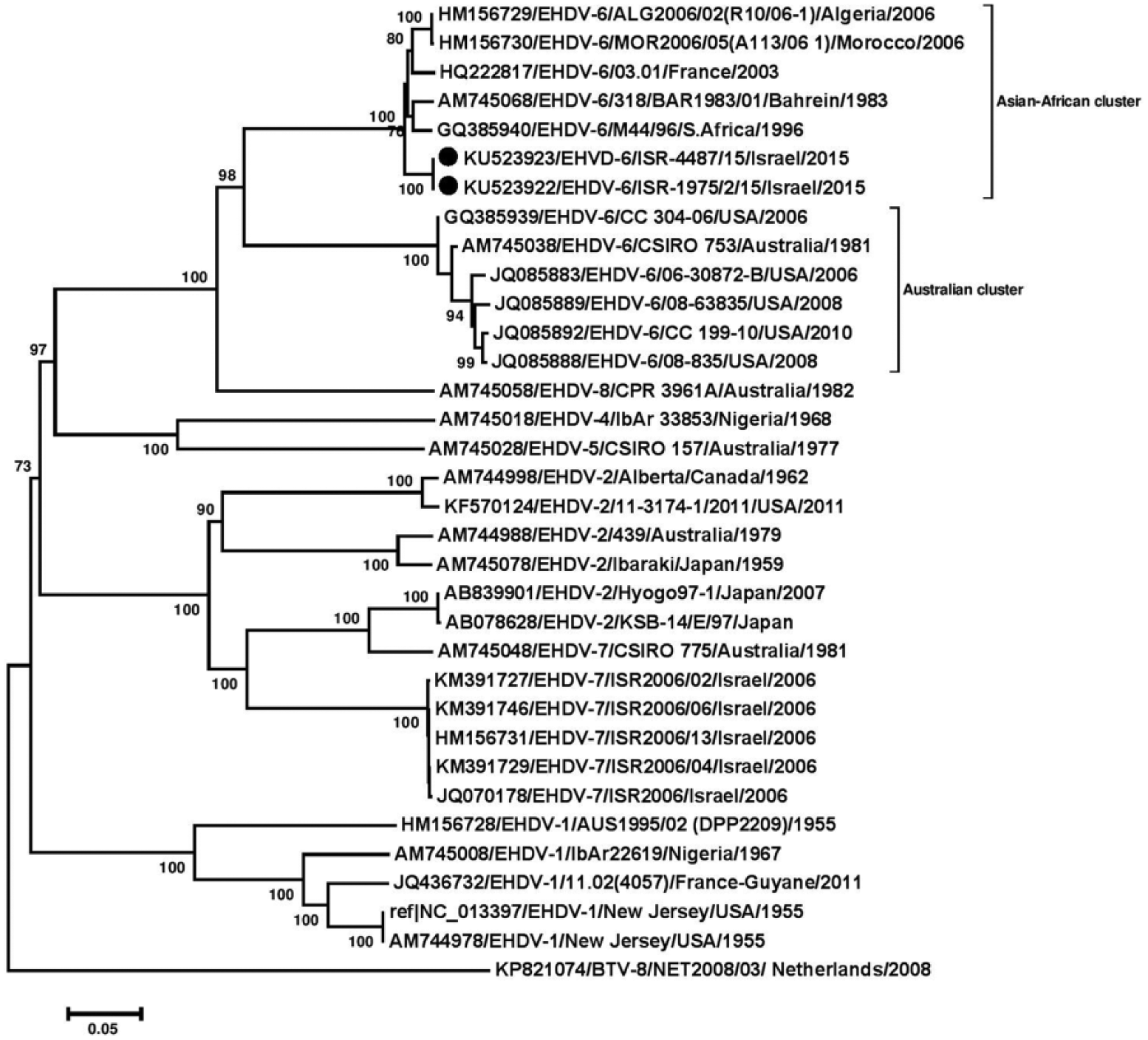
Phylogenetic analysis based on full-length sequences of segment 2 of 2 Israeli epizootic hemorrhagic disease virus (EHDV)-6 isolates with global EHDVs and bluetongue virus (BTV)-8 from GenBank. Nucleotide sequences (n = 34) were analyzed, and their phylogenetic relationship was inferred by using the neighbor-joining method. 8 Numbers below branches indicate bootstrap values. Israeli isolates from 2015 are marked with black dots. Viruses are identified by accession/serotype/isolate/location/year.
Footnotes
Acknowledgements
We thank Drs. Alexander Panshin, Yehuda Stram, Dan David, and Anita Kovtunenko, and Mrs. Marisol Rubinstein-Giuni.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
