Abstract
We evaluated duration of PCR-positive results following administration of modified-live viral (MLV) vaccines to beef calves. Twenty beef calves were randomly assigned to either group 1 and vaccinated intranasally with a MLV vaccine containing bovine alphaherpesvirus 1 (BoHV-1), bovine respiratory syncytial virus (BRSV), and bovine parainfluenza virus 3 (BPIV-3), or to group 2 and vaccinated subcutaneously with a MLV vaccine containing bovine viral diarrhea virus 1 and 2 (BVDV-1, -2), BoHV-1, BRSV, and BPIV-3. Deep nasopharyngeal swabs (NPS) and transtracheal washes (TTW) were collected from all calves, and whole blood was collected from group 2 calves and tested by PCR. In group 1, the proportions of calves that tested PCR-positive to BVDV, BoHV-1, BRSV, and BPIV-3 on any sample at any time were 0%, 100%, 100%, and 10%, respectively. In group 1 calves, 100% of calves became PCR-positive for BoHV-1 by day 3 post-vaccination and 100% of calves became PCR-positive for BRSV by day 7 post-vaccination. In group 2, the proportions of calves that tested positive to BVDV, BoHV-1, BRSV, and BPIV-3 on any sample at any time were 50%, 40%, 10%, and 0%, respectively. All threshold cycle (Ct) values were >30 in group 2 calves, irrespective of virus; however, Ct values <25 were observed in group 1 calves from PCR-positive results for BoHV-1 and BRSV. All calves were PCR-negative for all viruses after day 28. Following intranasal MLV viral vaccination, PCR results and Ct values for BRSV and BoHV-1 suggest that attempts to differentiate vaccine virus from natural infection is unreliable.
Keywords
Introduction
Bovine respiratory disease (BRD) complex is a multifactorial disease problem in the cattle industry. In addition to stress, environmental conditions, and management-related factors, infectious disease agents are important in the development of disease, and include viruses, bacteria, and mycoplasmas. 6 The primary viral respiratory pathogens implicated in BRD are bovine viral diarrhea virus (BVDV), bovine alphaherpesvirus 1 (BoHV-1), bovine parainfluenza virus 3 (BPIV-3; bovine respirovirus 3), and bovine respiratory syncytial virus (BRSV; bovine orthopneumovirus). These viruses primarily infect the upper respiratory tract, resulting in rhinitis, pharyngitis, tracheitis, and bronchitis, which subsequently predisposes the lower respiratory tract to bacterial infection. 4 Timely detection of viral respiratory pathogens is critically important in order to implement interventions strategies to control BRD.
Many different tests are available for detection of viral infections of the respiratory tract in cattle. 7 These include indirect testing, usually paired serology, to detect an immune response following infection by a viral agent, and direct determination of the presence of virus. Tests used to detect whole virus, viral antigen, or nucleotides include virus isolation (VI), fluorescent antibody and immunoperoxidase staining, antigen ELISA, and PCR.7,11 The introduction of PCR has transformed the effectiveness of viral detection by the diagnostic laboratory. Compared to traditional virus detection methods, PCR offers rapid results with high sensitivity and specificity. The advent of real-time (rt)PCR technologies has further improved the already significant benefits of traditional PCR, and rtPCR has become the tool often used by diagnostic laboratories for the detection of viral RNA and DNA.12,17,19 This technique enables simultaneous amplification and quantification of specific nucleic acid sequences, allowing multiplex tests to detect multiple agents in the same sample.10,13,20 The rtPCR platform is also adaptable for testing of large sample numbers rapidly. The ability to detect multiple infectious agents in a high-volume platform is leading to large-scale adoption of rtPCR in diagnostic laboratories for evaluation of the presence of agents contributing to BRD.
The enhanced sensitivity of rtPCR has led to some confusion regarding the significance of the presence of viral sequences and the correlation to disease. 7 Two potential circumstances that complicate interpretation of PCR results include samples from live cattle where no tissue is available for morphologic evaluation of lesion presence to the PCR result, and circumstances in which samples are procured from cattle recently vaccinated with modified-live viral (MLV) vaccines. In these situations, a PCR-positive result could indicate vaccine virus shedding, shedding of wild-type viruses from subclinical carrier animals, or the presence of viral respiratory pathogens from diseased animals. Multiple commercial multivalent MLV and killed viral vaccines are available and widely used for the prevention of BRD. In the recent U.S. Department of Agriculture (USDA) National Animal Health Monitoring System feedlot survey, nearly all individual cattle entering feedlots were vaccinated against viral respiratory pathogens, 23 with the majority of feedlots vaccinating within 72 h of arrival. 22 Because BRD is often observed in cattle within the first several weeks following arrival at a feedlot or stocker operation, vaccination soon after arrival could potentially complicate interpretation of PCR results for viral respiratory pathogens from diseased cattle. Therefore, we evaluated sequential post-vaccination PCR results from calves given parenteral or intranasal commercial multivalent MLV vaccines. Samples included deep nasopharyngeal swabs (NPS), transtracheal washes (TTW), and whole blood, which are commonly obtained from live calves for detection of viral pathogens associated with BRD complex. 2
Materials and methods
Animals
All study procedures were reviewed and approved by the Institutional Animal Care and Use Committee of Auburn University (PRN 2015-2748). Twenty clinically normal, crossbred beef steer calves were used in this study. All calves were born between January 6, 2015 and February 16, 2015 and raised at the Upper Coastal Plain Agricultural Research Center (Winfield, Alabama). Calves were identified by ear tattoo and ear tag at birth and allowed to remain with their dams with consumption of colostrum occurring under natural conditions in the pasture.
Experimental design
Our study was designed as a randomized control trial. Calves were weaned at 154–197 d of age. At weaning, all calves were weighed and assigned to either treatment group according to randomized block design. Calves were sorted by weight from highest to lowest. Random numbers were generated for each calf using commercial software (Excel 2010, Microsoft, Redmond, WA). For each pair of calves, starting with the heaviest pair, the higher number was assigned to group 1 and the lower number was assigned to group 2 for a total of 10 replicates. At 15 d prior to study initiation, all calves were transported from Winfield, AL to the North Auburn Beef Unit in Auburn, AL. Calves were isolated from all other cattle until study initiation.
On day 0 (vaccination), blood, TTW, and nasopharyngeal swabs were obtained from all calves. Following sample collection, calves were transported to the North Auburn BVDV Unit where they were vaccinated in different working chutes assigned to each group, housed in separate pastures, and subsequently handled through the 2 chute systems at the unit during sampling days. Calves were kept in separate pens to eliminate contact between groups, but each vaccine group was housed together. To prevent inadvertent transfer of vaccine viruses, different personnel cared for and sampled the calves during the study. At 3, 7, and 14 d post-vaccination (dpv), TTW and NPS were collected from all calves, with blood being an additional sample collected on group 2 calves. Sample collections were continued at 21, 28, 35, and 42 dpv only on calves that met a predetermined criterion for sampling, which was that a calf was no longer sample-eligible once that individual calf became negative for all viruses in all samples on 2 consecutive samples.
Vaccination
Vaccines used were USDA-licensed products approved for commercial use. Calves in group 1 were administered via intranasal cannula a multivalent MLV vaccine (Inforce 3, Zoetis, Florham Park, NJ) containing BRSV and temperature-sensitive variants of BoHV-1 and BPIV-3. Group 2 calves received a multivalent MLV vaccine (BoviShield Gold 5, Zoetis) containing BVDV-1, BVDV-2, BoHV-1, BRSV, and BPIV-3 subcutaneously. All vaccines were used within 1 h of reconstitution. All vaccines were administered according to label directions.
Sample collection
Whole blood was collected from all calves at weaning and at vaccination for VI testing for BVDV. Serum was collected from all calves at weaning, at vaccination (day 0), and at 28 dpv for determination of virus neutralizing antibody titers for each virus. On the day of vaccination, serum, whole blood, TTW, and NPS were collected from all calves. For group 1 calves that received the intranasal vaccine, TTW and NPS were collected on subsequent sample days and multiplex PCR assays were performed for the 4 viruses. For group 2 calves, whole blood, TTW, and NPS were collected on subsequent sample days, and multiplex PCR assays were performed on TTW and NPS samples; individual PCR assays were performed for BVDV and BoHV-1 on whole blood. For TTW collections,
2
calves were restrained in a chute with head pulled dorsally. The ventral neck area was surgically prepared and anesthetized using local analgesia. A small stab incision was made through the skin, and a 12-gauge, 5-cm sterile needle was inserted into the tracheal lumen between 2 tracheal rings. A sterile polyethylene tube (#5 French, 60-cm length; Argyle suction catheter, Covidien, Mansfield, MA) was inserted through the needle and advanced ~50 cm. Sterile 0.9% saline (30 mL) was administered through the tube into the trachea, and 5–10 mL of the wash was collected and placed in sterile transport tubes. The deep NPS were obtained using a double-guarded culture swab (Equine uterine culture swab, Agtech, Manhattan, KS) immediately following the collection of the TTW samples.
9
Nostrils were first cleaned with disposable gauze. The swab was inserted into the lowest portion of the nostril in order to enter the ventral meatus of the nasal passage. The nasal swab was advanced ~30 cm until making contact with the nasopharynx. The swab was retracted slightly and the inner guard pushed through the end of the outer guard. The cotton-tipped swab was then pushed through the inner guard and vigorously rotated against the pharyngeal mucosa for 15–30 s. The cotton-tipped swab was retracted within the inner guard and the entire double-guarded swab removed from the calf’s nose. The swab was immediately placed into sterile transport medium (Gibco Eagle minimum essential medium [Life Technologies, Grand Island, NY] supplemented with 10% equine serum [Hyclone, GE Healthcare Life Sciences, Logan, UT],
Molecular testing for BVDV, BoHV-1, BRSV, and BPIV-3
The TTW and NPS specimens were collected and shipped by overnight courier to the Animal Health Diagnostic Center (AHDC) at Cornell University (Ithaca, New York) as standard test submissions. A multiplex PCR for BVDV, BoHV-1, BRSV, and BPIV-3 was performed. Personnel that performed the assays at the AHDC were blinded to the group designation, and results were recorded as negative or positive with threshold cycle (Ct) values provided. The Ct value is defined as the number of cycles required for the fluorescence signal to cross the threshold that corresponds to a fluorescence signal significantly above background fluorescence. Ct values are inversely proportional to the amount of target nucleic acid in the sample, with a lower Ct value indicating a greater amount of target nucleic acid in the sample.
Whole blood was subjected to rtPCR testing for BVDV and BoHV-1 on all calves on day 0 and from group 2 calves on days 3, 7, 14, 21, and weekly thereafter from each individual calf until PCR negative on 2 consecutive weeks. A membrane kit (High Pure PCR Template preparation kit, Roche, Branchburg, NJ) was used to extract viral RNA or DNA from whole blood specimens according to the manufacturer’s instructions. SYBR Green–based rtPCR protocols were developed to detect BVDV and BoHV-1 as described previously. 8 The optimal PCR conditions were determined to be the lowest concentration of the standard template required to amplify the correct gene fragment corresponding to each virus. Forward and reverse primers were used for both viruses (BVDV-F: 5’-TAGCCATGCCCTTAGTAGGAC-3’, BVDV-R: 5’-GACGACTACCCTGTACTCAGG-3’; BoHV-1-F: 5’-GTAAGGGTATATTATTGATTGC-3’, BoHV-1-R: 5’-GAGAGTGAGTATGAGGAC-3’), respectively. For BVDV, the reaction mixture in each tube contained 5 µL of RNA template, 10 µL of iTaq SYBR Green mix (Bio-Rad, Hercules, CA), 0.25 µL of reverse transcriptase, 0.3 µM (0.3 µL) each of forward and reverse primers, and sterile PCR water to make up the total reaction volume of 15 µL. The PCR protocol for detection of BVDV was 50°C for 10 min for 1 cycle, 95°C for 1 min for 1 cycle, 95°C for 10 s for 1 cycle, 56.5°C for 30 s for 1 cycle, and 95°C for 10 s for 39 cycles. A positive control was maintained using the BVDV standard template. No-template control was maintained using PCR-grade water instead of the standard template. An additional control without reverse transcriptase was maintained for the BVDV PCR assay. For BoHV-1, the reaction mixture contained 5 µL of DNA template, 10 µL of SYBR Green supermix (ssoAdvanced, Bio-Rad), 0.5 µM (0.5 µL) each of forward and reverse primers for BoHV-1, and sterile PCR-grade water for a total volume of 15 µL. The PCR protocol for detection of BoHV-1 was 1 cycle at 98°C for 3 min, 95°C for 15 s, 60°C for 30 s, followed by 35 cycles at 95°C for 15 s. The standard DNA template served as positive control. The non-template control served as negative control, which contained PCR-grade water instead of DNA template. All of the reactions were subjected to rtPCR using a rtPCR detection system (CFX96 Touch, Bio-Rad). The rtPCR yielded the expected products for BoHV-1 (128 bp) and BVDV (96 bp).
VI
Whole blood and serum samples were collected for BVDV detection by VI at weaning and on the day of vaccination (day 0). Briefly, isolated WBC samples (buffy coat) were subjected to co-cultivation in 24-well plates containing monolayers of Madin–Darby bovine kidney (MDBK) cells. Serum samples were inoculated into 24-well plates containing monolayers of MDBK cells. The 24-well plates were incubated for 4 d at 37°C and 5% CO2. Following cultivation, all plates were frozen at −80°C and thawed, and then 10 μL of the sample was inoculated in triplicate into wells on 96-well microtiter plates followed by MDBK cells in culture medium. After 72 h of incubation at 37°C and 5% CO2, all wells were stained to detect BVDV using the immunoperoxidase monolayer assay as described previously. 25
Virus neutralization
The standard virus neutralization (VN) microtiter assay was used to detect antibodies to BVDV, BoHV-1, BRSV, and BPIV-3 in serum collected at weaning, day 0, and day 28. 25 Sera were tested for neutralizing antibodies against BVDV-1a (BVDV strain NADL), BVDV-2 (BVDV strain 125c), BoHV-1 (Colorado strain), BRSV (A51908 strain), and BPIV-3 (TVMD60). Following heat inactivation of serum samples at 56°C for 30 min, serial 2-fold dilutions (1:2 to 1:4,096) were made in 50 μL of culture medium. For each dilution, 3 wells of a 96-well plate were inoculated with an equal volume (50 μL) of culture medium containing 100–500 CCID50 (50% cell culture infective dose) of the respective virus strain. After inoculation, the plate was incubated at 38.5°C in a humidified atmosphere of 5% CO2 and room air for 1 h. Then, 2.5 × 103 MDBK cells (BVDV, BoHV-1, BPIV-3 assays) or bovine turbinate cells (BRSV assay) in 50 μL of culture medium were added to each well. Plates were incubated for 72 h and evaluated visually for cytopathic effect. 25 For the BoHV-1 VN assay, after each well was inoculated with the respective serum dilution, plates were incubated for 120 h at 38.5°C in a humidified atmosphere of 5% CO2 and air, and then each well was examined for the characteristic cytopathic effect of BoHV-1. The geometric mean of antibody titers was calculated from the endpoint log2 titers of the animals in each group.
Results
Following randomization, average weight for group 1 calves and group 2 calves was 211 kg and 212 kg, respectively, at weaning and 226 kg and 228 kg on day 0. Group 1 calves averaged 219 d in age (211–229 d) and group 2 calves averaged 214 d in age (189–232 d) on day 0. Testing for BVDV by VI at weaning and on day 0 was negative. With respect to general results, all group 1 calves were positive for at least 1 virus on at least 1 sample collection day; however, 3 group 2 calves were negative for all viruses at any sample collection day.
Following vaccination, all calves in both groups were sampled at 3, 7, and 14 dpv. By 21 dpv, all group 1 calves and 7 group 2 calves were sampled, as the 3 aforementioned group 2 calves were ineligible for further sampling because of 2 consecutive negative results from weekly testing. By 28 dpv, 9 group 1 calves and 6 group 2 calves were sampled; by 35 dpv, 5 group 1 calves and 1 group 2 calf were sampled; and, by day 42, 2 group 1 calves and no group 2 calves were sampled. No calves had a positive PCR result for any virus beyond 28 dpv, and all calves had met the eligibility criterion of no further sampling after 2 consecutive negative weekly tests for all viruses by 42 dpv.
In group 1, the proportions of calves that tested positive to BVDV, BoHV-1, BRSV, and BPIV-3 on any sample at any time were 0%, 100%, 100%, and 10%, respectively. In group 2, the proportions of calves that tested positive to BVDV, BoHV-1, BRSV, and BPIV-3 on any sample at any time were 50%, 40%, 10%, and 0%, respectively. The overall detection by sample type varied between groups (Table 1), with NPS specimens more likely to result in a PCR-positive outcome than TTW in group 1, and TTW specimens more likely to result in a PCR-positive outcome in group 2. With respect to buffy coat samples, BVDV was the only virus to result in a PCR-positive outcome and was only observed in group 2. One calf in group 2 was positive for BVDV in buffy coat but was negative for BVDV in all testing performed on NPS and TTW. For BPIV-3, all calves were negative on all sample days with the exception of a single group 1 calf on day 7 after intranasal vaccination.
Proportion of calves with a PCR-positive result by sample type and median days to first PCR-positive and second PCR-negative result following vaccination with a multivalent modified-live viral (MLV) intranasal vaccine (group 1) or a multivalent MLV parenteral vaccine (group 2).*
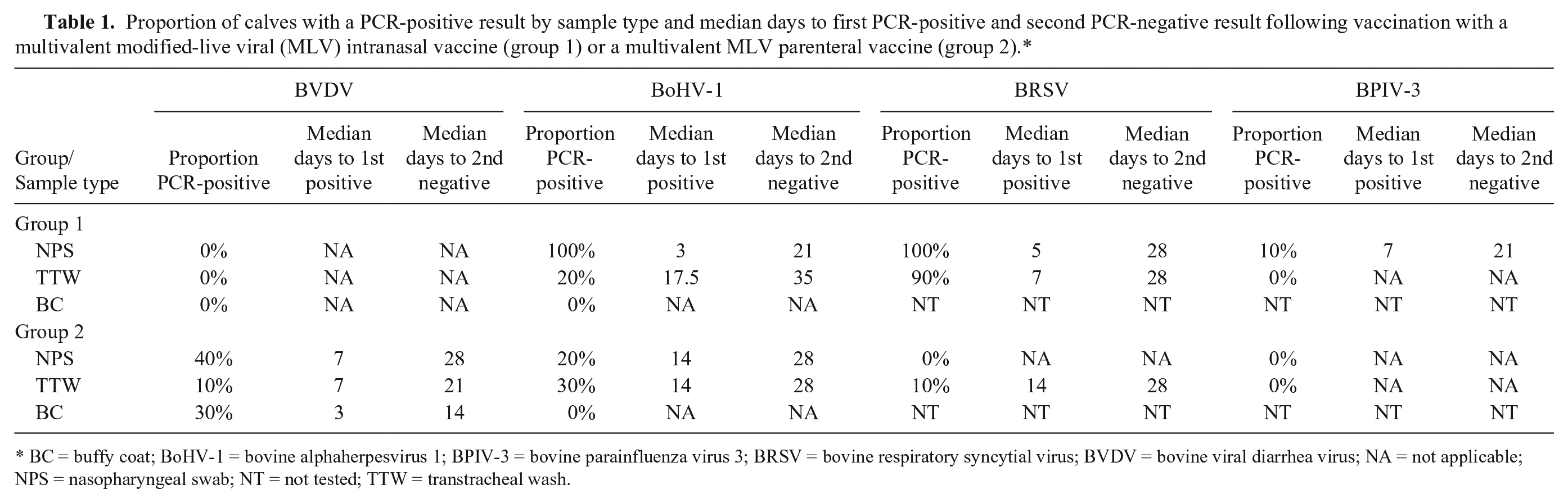
BC = buffy coat; BoHV-1 = bovine alphaherpesvirus 1; BPIV-3 = bovine parainfluenza virus 3; BRSV = bovine respiratory syncytial virus; BVDV = bovine viral diarrhea virus; NA = not applicable; NPS = nasopharyngeal swab; NT = not tested; TTW = transtracheal wash.
PCR testing results from deep NPS
In group 2 calves, 3 and 4 calves were positive for BVDV from NPS on day 7 and 14, respectively (Table 2). Two calves were PCR-positive for BVDV only in NPS, and were positive 7 and 14 dpv. All Ct values were >35 on all calves positive for BVDV in NPS. As expected, BVDV was not recovered from any NPS at any sample day because BVDV was not a component of the vaccine administered to group 1 calves.
Proportion of calves with a PCR-positive result for bovine viral diarrhea virus (BVDV), bovine alphaherpesvirus 1 (BoHV-1), and bovine respiratory syncytial virus (BRSV) by day post-vaccination (dpv) with corresponding threshold cycle (Ct) values (mean and range) following vaccination with a multivalent modified-live viral (MLV) intranasal vaccine (group 1) or a multivalent MLV parenteral vaccine (group 2).*
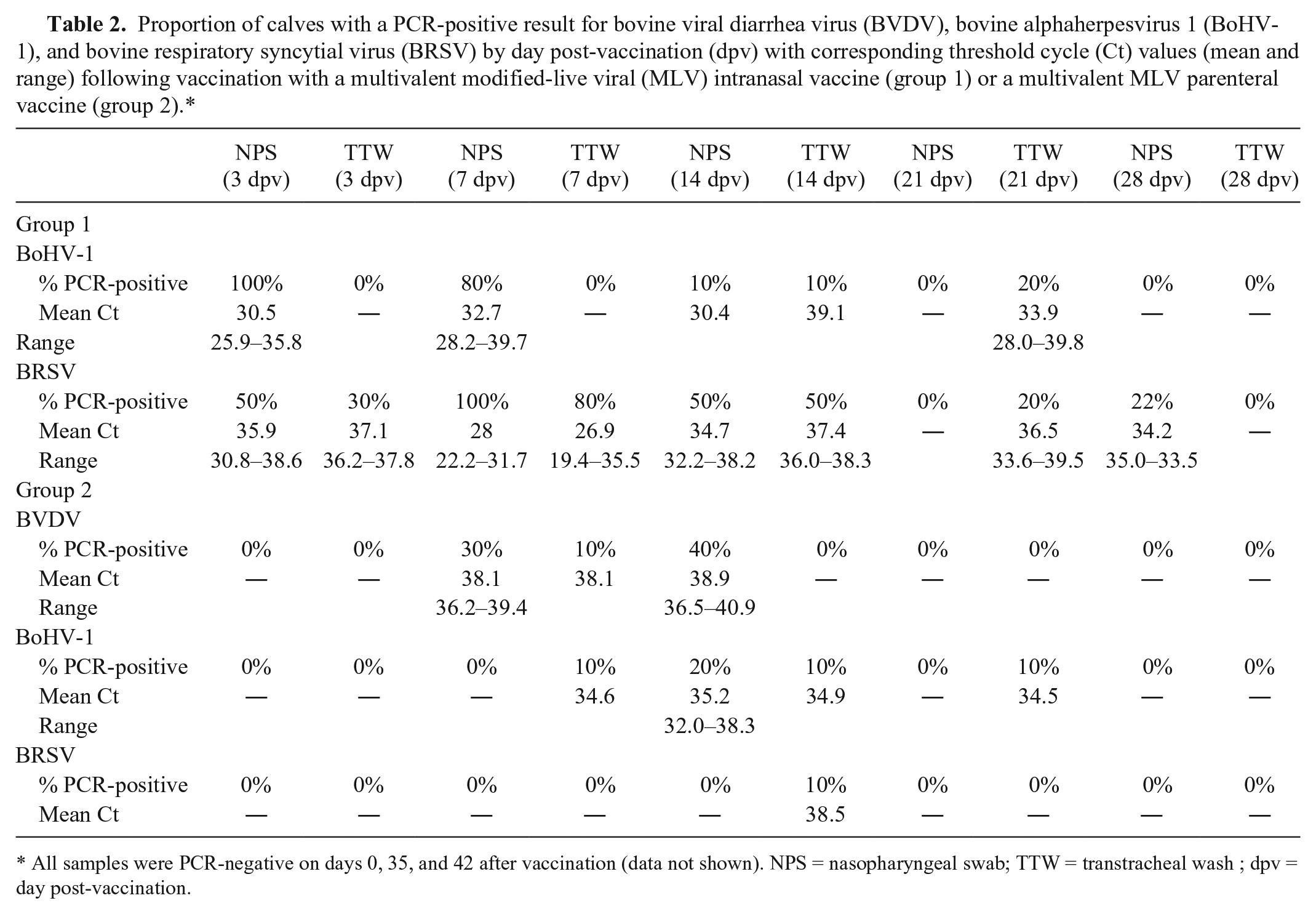
All samples were PCR-negative on days 0, 35, and 42 after vaccination (data not shown). NPS = nasopharyngeal swab; TTW = transtracheal wash ; dpv = day post-vaccination.
For BoHV-1, all group 1 calves were positive in NPS by 3 dpv, with 8 calves and 1 calf positive for BoHV-1 in NPS by 7 and 14 dpv, respectively (Table 2). All group 1 calves were negative for BoHV-1 after day 14. In contrast, only 2 group 2 calves were positive for BoHV-1 and only at 14 dpv. The average Ct values from group 1 calves were lower on days 3, 7, and 14 as compared to the 2 group 2 calves on day 14 (Table 2). All individual Ct values were >30 except for 4 group 1 calves on day 3 and 2 group 2 calves on day 7. The lowest Ct value from group 2 calves was 32.
For BRSV, 5 group 1 calves were positive in NPS by 3 dpv, with all 10 calves and 5 calves positive for BRSV in NPS by 7 and 14 dpv, respectively. Interestingly, all group 1 calves were negative for BRSV on day 21, but then 2 calves were positive for BRSV in NPS on day 28. These 2 calves had previously been positive for BRSV in NPS specimens at 3 and 7 dpv. The Ct values were generally high (>30 cycles) with the exception of the 14 dpv sample date, when 8 of the 10 calves had Ct values <30, with the lowest reported Ct value being 22. In contrast, no group 2 calves were positive for BRSV in NPS specimens on any sample day. For BPIV-3, only a single group 1 calf was PCR-positive in a NPS specimen, with a Ct value of 39. No group 2 calves were PCR-positive for BPIV-3 in NPS specimens.
PCR testing results from TTW
For BVDV, only a single group 2 calf was PCR-positive for BVDV in TTW specimen, with the positive result 7 dpv (Ct = 38; Table 2). As expected, no group 1 calves were BVDV PCR-positive on any day post-vaccination.
For BoHV-1, only 2 group 1 calves were PCR-positive, with 1 calf positive at 14 and 21 dpv and the other calf only positive at 21 dpv. In contrast, 3 group 2 calves became PCR-positive for BoHV-1, with each calf having a single positive result on 3 separate sampling days (7, 14, and 21 dpv). All calves, irrespective of group, had Ct values >34 in TTW samples.
For BRSV, PCR-positive results were obtained on TTW specimens in 90% of group 1 calves. By 3 dpv, BRSV PCR-positive results were obtained from 3 group 1 calves, but Ct values were >36 (Table 2). By 7 dpv, BRSV PCR-positive results were obtained from 8 group 1 calves, with Ct values <30 in 6 calves. The lowest Ct value for BRSV was 19 and occurred at 7 dpv. By 14 dpv, 5 calves were PCR-positive, but Ct values were all >35. Only 2 group 1 calves were PCR-positive for BRSV on day 21, and all calves were negative for BRSV in TTW samples by day 28. In group 2 calves, a single calf was PCR-positive for BRSV in TTW samples at 14 dpv with a Ct value >38.
VN
On the day of vaccination, all calves were negative (titer < 1:2) for antibodies against all viruses with the exception of 3 calves. A single group 1 calf possessed an antibody titer of 2 against BPIV-3 and BRSV, a single group 2 calf possessed an antibody titer of 2 against BRSV, and a single group 2 calf possessed an antibody titer of 32 against BPIV-3. By 28 dpv, all group 2 calves had seroconverted to BVDV, but all group 1 calves remained negative by VN (Table 3). Comparison of antibody responses directed against BVDV-1 and BVDV-2 revealed that group 2 calves possessed a higher concentration of antibodies directed against BVDV-1 by 28 dpv. For BoHV-1, only 1 calf from group 1 had any measurable antibody response by 28 dpv, with an antibody titer of 4. In contrast, all group 2 calves had a measurable antibody response by day 28; however, antibody titers were generally low (2–8). All calves in both groups developed a measurable antibody response to BRSV, with group 2 calves possessing higher antibody concentrations than group 1. For BPIV-3, 80% of group 1 and 80% of group 2 calves demonstrated a rise in antibody titer, with group1 calves possessing higher antibody concentrations than group 2.
Geometric mean virus neutralizing antibody titers against bovine viral diarrhea virus 1 and 2 (BVDV-1, -2), bovine alphaherpesvirus 1 (BoHV-1), bovine respiratory syncytial virus (BRSV), and bovine parainfluenza virus 3 (BPIV-3) following vaccination with a multivalent modified-live viral (MLV) intranasal vaccine (group 1) or a multivalent MLV parenteral vaccine (group 2).*
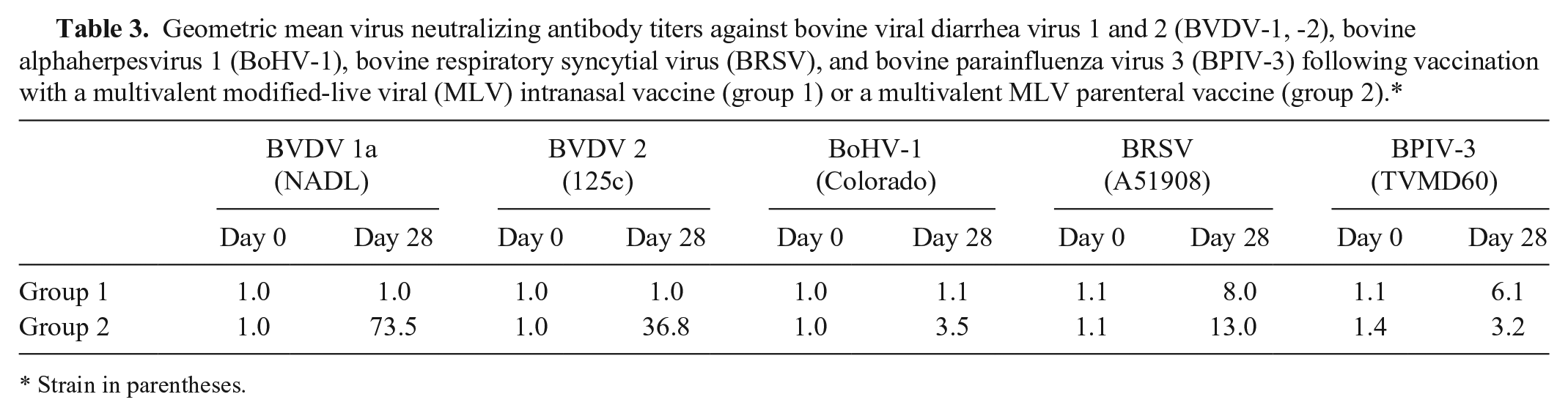
Strain in parentheses.
Discussion
In our study, vaccination of cattle with multivalent MLV vaccines resulted in high proportions of calves with PCR-positive results for viral respiratory pathogens in clinically relevant samples routinely collected from cattle undergoing respiratory disease.1,2,9 Use of multivalent MLV vaccines containing BVDV, BoHV-1, and BRSV in cattle soon after arrival at stocker or feedlot operations could potentially impact the utility of molecular testing for these viral respiratory pathogens in sick or diseased cattle when testing is performed within several weeks after arrival. Interpretation of PCR-positive results then becomes difficult for veterinarians and diagnosticians in the absence of lesions or recovery of infectious virus in these cattle. 7 To address this problem, diagnostic laboratories may try to use Ct values obtained from PCR-positive NPS or TTW samples as a method for differentiating infection with wild-type virus from vaccination with MLV vaccines. In general, the Ct values were high following the parenterally administered MLV vaccine. The variability and lower Ct values for BRSV and BoHV-1 following administration of the intranasal MLV vaccine to calves makes differentiation of infection with wild-type virus from vaccine virus difficult.
The intranasally administered multivalent MLV vaccine containing BoHV-1, BRSV, and BPIV-3 resulted in PCR-positive results in NPS specimens for BoHV-1 and BRSV from 100% of calves vaccinated. The TTW samples yielded far fewer PCR-positive calves for BoHV-1 following intranasal MLV vaccination than NPS samples. This finding is interesting because differences could potentially be observed between NPS and TTW PCR-positive results during infection with unattenuated wild-type BoHV-1 strains as compared to intranasal vaccination with the temperature-sensitive variant of BoHV-1, although further research is necessary. As expected, BVDV was not detected in any calves administered the trivalent intranasal vaccine because that vaccine did not contain BVDV. Although no attempt was made to culture viruses, given that our purpose was to evaluate PCR testing results, previous studies evaluating intranasal MLV vaccines containing a temperature-sensitive variant of BoHV-1 have demonstrated high viral replication in the upper respiratory tract with potential for shedding.5,16,26 Therefore, PCR-positive results for BoHV-1 could be expected following intranasal vaccination with BoHV-1. For BRSV, a previous study demonstrated that reverse transcription PCR testing performed on nasal swab samples detected vaccinal BRSV in 15 of 17 calves following intranasal administration of a MLV vaccine containing BRSV and BPIV-3. 21 The Ct values in some of the calves with PCR-positive nasal swabs were in the low to mid-twenties, 21 which was a finding also observed in our study. Interestingly, 2 calves in our study had a PCR-positive result for BRSV beyond 14 dpv and occurring after 1–2 wk of PCR-negative results. This finding was also observed in previous research. 21 Although the cause for this reoccurrence is unknown, possibilities include reinfection by circulating BRSV vaccine virus or inconsistent quantity of nasal epithelium containing BRSV thus impacting PCR sensitivity on a weekly basis. A previous study demonstrated that live vaccinal BRSV was not shed, as in-contact susceptible calves remained seronegative to BRSV. 21 The quantity of nasal epithelium obtained during deep NPS collections does not seem highly plausible because the finding of intermittent PCR-positive results following PCR-negative results was not observed for BoHV-1 or BVDV, and because once a calf became PCR-negative for BoHV-1 and BVDV, they remained negative. BPIV-3 was only recovered from 1 calf on a single day following intranasal vaccination. The BPIV-3 strain in the intranasal vaccine used in our study is a temperature-sensitive variant that only replicates in the nasal epithelium. Although studies are limited on shedding of BPIV-3 vaccine strains, one study reported no shedding of BPIV-3 at 10 dpv in cattle receiving the same temperature-sensitive mutant of BPIV-3 used in our study. 24
The parenterally administered multivalent MLV containing BVDV, BoHV-1, BRSV, and BPIV-3 caused far fewer PCR-positive results in NPS and TTW specimens compared to the intranasal MLV vaccine. Interestingly, the TTW samples were more likely to result in PCR-positive results for BoHV-1 and BRSV than NPS following parenteral administration of MLV vaccine, which is in contrast to the intranasal MLV vaccine. However, NPS were more likely to be PCR-positive for BVDV than TTW. Buffy coat samples have long been identified as ideal samples for detecting BVDV in acutely and persistently infected animals.11,18 Our study indicates that a nasal swab is a more sensitive sample for detecting BVDV following MLV vaccination than other samples. In spite of the paucity of PCR-positive results for all 4 respiratory viruses, all group 2 calves developed neutralizing antibody responses, thus indicating systemic immune recognition, likely resulting from vaccine virus replication. The BVDV and BoHV-1 vaccine components used in our study have been shown to not be transmitted to in-contact animals, 3 so the results of less than half of vaccinated calves having PCR-positive results for these individual viruses is expected. The high Ct values and short duration of detection after vaccination would support that this systemic MLV vaccine would be unlikely to be transmitted from healthy vaccinated animals. The single BRSV PCR-positive result is difficult to explain because even unattenuated BRSV has been shown to undergo minimal replication when administered systemically. 14 However, these results should be repeated using other parenterally administered MLV vaccines as the results may vary because of differences in strains, attenuation, and viral amounts in different vaccines. 15
Footnotes
Acknowledgements
We thank Dr. Amy Glaser for providing technical expertise associated with assays performed at the Animal Health Diagnostic Center, Cornell University. We also thank Ms. Patricia Galik and Ms. Yijing Zhang for technical support with assays performed at Auburn University, and Mr. George Fincher and Mr. David Powell at Auburn University for animal care assistance.
Declaration of conflicting interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Daniel Scruggs and Victor Cortese are employees of Zoetis Inc. None of the authors directly benefited financially from completion of this work.
Funding
Funds for this research project were provided by Zoetis Inc., Florham Park, NJ.
