Abstract
Control of bovine viral diarrhea virus (BVDV) relies on resource-intensive sampling to detect and remove persistently infected (PI) cattle. Herd-level surveillance tools would be useful for herds with unknown BVDV status and for monitoring herds with BVDV-free status. Our objective was to determine the feasibility of using stable flies as a sampling tool to detect BVDV at the herd level. Stable flies (Stomoxys calcitrans) were fed citrated blood from either BVDV-PI or BVDV-free cattle to establish pools of 100 flies with various proportions of BVDV-fed flies (0%, 1%, 10%, 20%, 40%, or 100% in each pool). BVDV-fed flies in these pools were harvested either 1, 2, or 3 d after consuming BVDV-PI blood to determine the impact of time after feeding. Two replicates of a 3-d by 6-dilution level matrix were produced. BVDV RNA was consistently detected on day 1 when ≥10% of the flies in the pool consumed PI blood. On days 2 and 3, positive BVDV RNA detection was variable and became less consistent. Our results demonstrate that BVDV RNA can be detected in stable flies after feeding on blood from PI cattle. Successful use of stable flies as a surveillance tool will require validation under field conditions.
Bovine viral diarrhea virus (BVDV; Flaviviridae, Pestivirus) is an important, prevalent pathogen of cattle around the world.5,8 An estimated 91% of U.S. cow–calf operations have at least one seropositive animal as a result of vaccination, maternal antibody, or exposure. 6 Calves infected with BVDV during the first trimester of gestation become persistently infected (PI). These calves shed BVDV for their entire lives and serve as the most important source of new infections in cattle. 7 Thus, effective disease control relies on detecting and removing BVDV-PI cattle from herds. 3 Identifying herds harboring BVDV-PI cattle currently requires resource-intensive, individual animal sampling, followed by individual or pooled testing. 10 True herd-level surveillance strategies for beef herds rely on evidence of seroconversion and are not feasible in vaccinated herds. Consequently, routine surveillance for BVDV is not conducted commonly. Developing methods to classify BVDV status at a herd level without the need to handle the cattle would remove many obstacles that limit adoption of BVDV control. Previous research has shown that stable flies (syn. biting flies; Stomoxys calcitrans) can harbor the virus for at least 96 h after feeding on an infected animal and can transmit infection to susceptible animals. 9 Our objective was to determine the feasibility of using stable flies as a sampling tool to detect BVDV RNA in cattle herds. We hypothesized that BVDV RNA would be detectable in stable flies that had fed on blood from BVDV-PI calves.
Stable fly pupae were transported from the West Central Research and Extension Center (North Platte, NE) to the Great Plains Veterinary Educational Center (University of Nebraska–Lincoln, Clay Center, NE) and placed in mesh-covered cages at room temperature. Approximately 1,000 flies emerged during a 2- to 5-d period while being continuously exposed to a sanitary napkin on the top of the cage each day saturated with 20 mL of citrated whole blood from BVDV-PI calves (provided by the Department of Pathobiology, Auburn University, Auburn, AL). Blood samples from several BVDV-PI calves were collected, shipped on ice, and then frozen at −20°C. At the beginning of the experiment, blood from both BVDV-PI calves and BVDV-free calves was thawed for feeding to the flies and stored at 4°C until the experiment was completed. Aliquots of BVDV-PI blood and BVDV-free blood used during the experiment were stored at −80°C for use as positive and negative controls for further analysis. Flies were briefly chilled at −20°C (< 5 min) to immobilize them and facilitate handling. Unhatched pupae were moved to a separate cage to hatch control (BVDV-free) flies. After the non-eclosed pupae were removed, flies in the BVDV-fed cage were offered blood from BVDV-PI calves for an additional 24 h. The day after BVDV-fed and BVDV-free fly populations were separated was considered day 1 and both cages received BVDV-free blood for the remainder of the study. Each day from day 1 to 3, enough BVDV-fed and BVDV-free flies were collected to establish the following pools (percentage of BVDV-fed flies of 100 total flies per pool): 0%, 1%, 10%, 20%, 40%, and 100%. Each pool of 100 flies was suspended in 2 mL of phosphate-buffered saline and homogenized using tissue dissociation tubes (gentleMACS M tubes; Miltenyi Biotec, Auburn, CA). Homogenates were clarified by centrifuging for 5 min at 1,500 × g, and 140 µL of the supernatant was used for RNA extraction using a spin-column viral RNA isolation kit (QIAamp viral RNA mini kit; Qiagen, Germantown, MD) per the manufacturer’s instructions.
Reverse-transcription, real-time PCR (RT-rtPCR) was carried out on a real-time PCR thermocycler and detector (CFX96 touch real-time PCR detection system; Bio-Rad Laboratories, Hercules, CA) using a kit according to the manufacturer’s instructions (OneStep RT-PCR kit; Qiagen). Each 25 µL of RT-rtPCR reaction contained 1× reaction buffer, 1 μL of enzyme solution, 400 µM of NTP, 0.4 µM each of forward and reverse primers (5′-GGGNAGTCGTCARTGGTTCG-3′ and 5′-GTGCCATGTACAGCAGAGWTTTT-3′, respectively), 4 0.1 µM of oligonucleotide probe (5′-/56-FAM/CCAYGTGGACGAGGGCAYGC/3BHQ_1/-3′), and 0.005 µM of magnesium. The reverse-transcription step was carried out at 50°C for 30 min, followed by a 15-min PCR activation step at 95°C, and then 40 cycles of 94°C for 30 s, 55°C for 1 min, and 72°C for 1 min. The final step was a 5-min extension at 72°C. Each extracted RNA sample was assayed in duplicate on the same RT-rtPCR plate.
The experimental process of feeding and collecting flies, extracting viral RNA from the fly pool homogenates, and RT-rtPCR analysis was repeated to obtain 2 complete replicates of the 3-d by 6-dilution level matrix (Supplementary Fig. 1). Each experimental feeding trial replicate had a positive control comprised of RNA extracted from the blood from the BVDV-PI calves and multiple negative controls, including RNA extracted from the 0% PI flies, RNA extracted from the BVDV-free blood, and nuclease-free water. Results were analyzed with software provided by the manufacturer according to their instructions (CFX Manager v.3.1; Bio-Rad). Relative fluorescence units (RFU) were examined for the negative controls, and a single threshold line at 150 RFU was set to classify all negative controls as negative, including the fly pools with BVDV-free flies. Experimental samples failing to exceed the 150 RFU threshold line were classified as negative. A cycle threshold (Ct) value was provided when any sample exceeded the 150 RFU threshold line.
Results showed that BVDV RNA was readily detected by RT-rtPCR in fly pools with ≥ 10% exposed flies immediately after their last 24-h exposure to BVDV-PI blood (Fig. 1). BVDV RNA was detected in all pools except the 20% pool on day 2 (Table 1). On day 3, the 100% pool yielded positive results, whereas the other pools were either not detectable or had only one duplicate classified as positive. No amplification of BVDV RNA was detected in any of the negative controls in either replicate (Fig. 1).
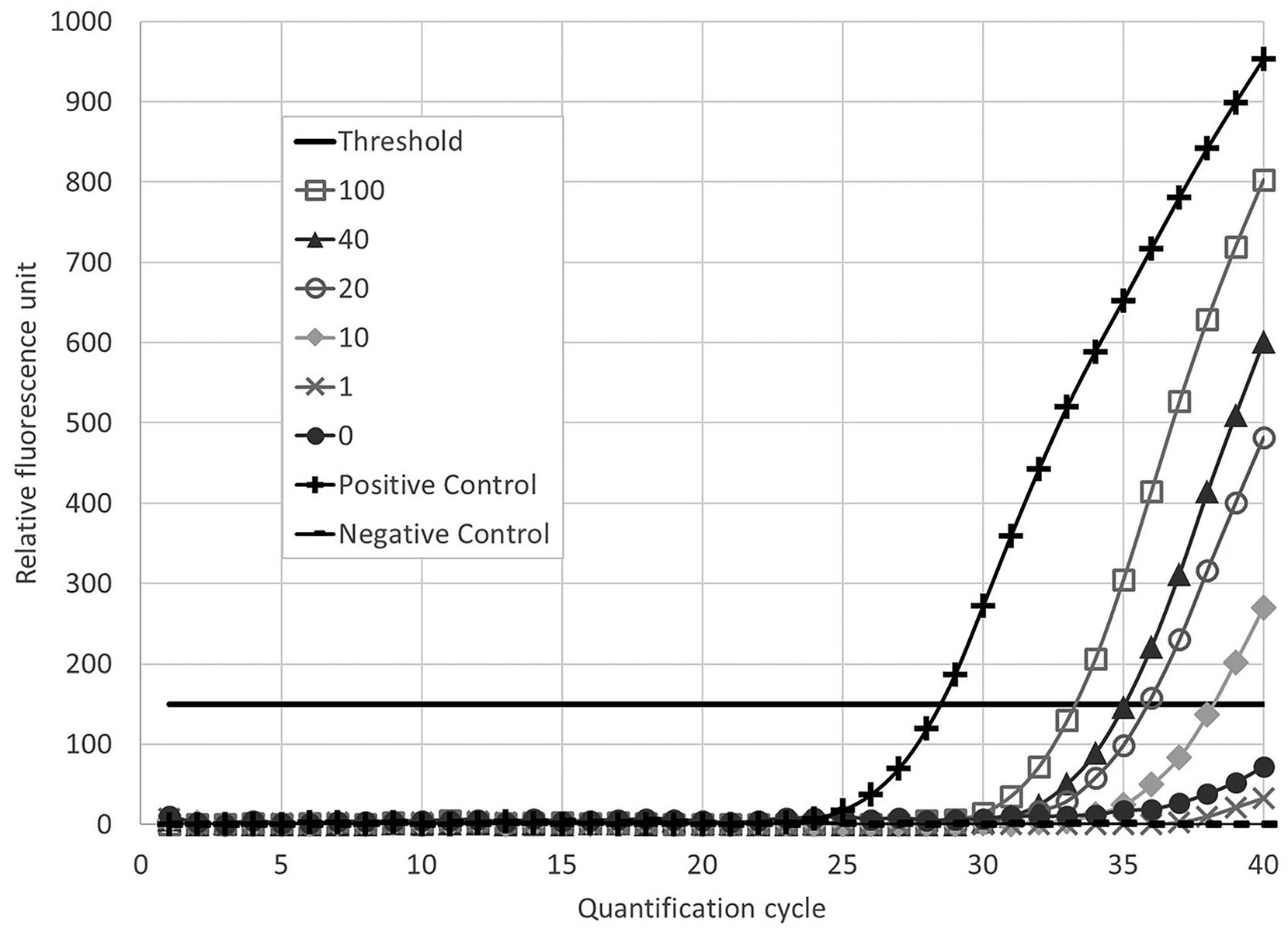
Representative reverse-transcription, real-time PCR cycle threshold data on day 1 from 2 replicates of a 3-d by 6-dilution level matrix of stable fly homogenates, with 1–100% of flies that were fed bovine viral diarrhea virus–containing blood.
Summary of bovine viral diarrhea virus (BVDV) reverse-transcription, real-time PCR (RT-rtPCR) cycle threshold (Ct) data from 2 replicates of a 3-d by 6-dilution level matrix of stable fly homogenates.
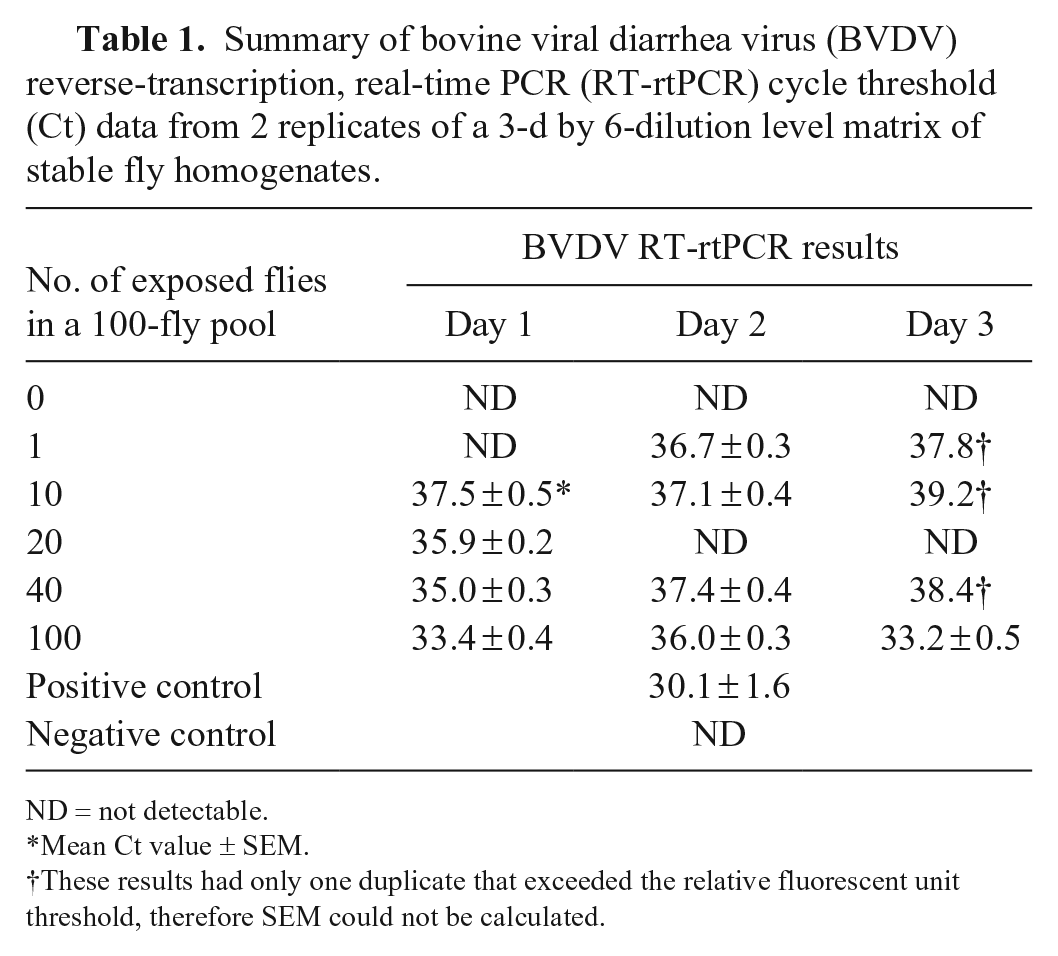
ND = not detectable.
Mean Ct value ± SEM.
These results had only one duplicate that exceeded the relative fluorescent unit threshold, therefore SEM could not be calculated.
We failed to reject our hypothesis that BVDV RNA would be detectable in stable flies after feeding on blood from PI calves. These results are consistent with a report in which flies allowed to feed directly from a PI calf were able to transmit BVDV to naïve cattle, and BVDV remained detectable by virus isolation for 96 h following feeding on a PI calf. 9 In that previous study, all flies used to isolate BVDV had been exposed to a PI calf; however, in a field situation, the mostly likely scenario would be one in which only a few flies of the total population captured would have fed on any PI cattle present. The prevalence of PI cattle has been estimated to be 0.5–2%, although the number of PI animals within a herd can vary.2,11 Therefore, we examined the impact of both diluting BVDV-fed flies with BVDV-free flies and the effect of feeding BVDV-free blood to BVDV-fed flies over the course of 3 d. Although our study provides evidence that BVDV RNA can be detected in stable flies exposed to blood from BVDV-PI cattle, successful use of stable flies as a surveillance tool will depend on validation under field conditions.
Substantial variation across experimental replicates, days, and pools was noted. Although many sources of variation could contribute, one notable source could be the feeding status of the flies. Flies in both the BVDV-fed group and the group fed only BVDV-free blood were offered the appropriate type of blood. In the context of our experiment, flies offered the opportunity to feed on blood were assumed to have done so. No attempt was made to confirm that the flies that had been offered blood consumed it. In the 1% pools, whether the fly selected for the pool had consumed blood was not known, and the feeding status of the fly may have impacted our results. In the larger pools, this explanation is less likely given that a significant number of flies would have had to forgo blood meals to generate the results seen. Specifically, the results from the 20% pools on days 2 and 3 were anomalous in that those pools appear to have less BVDV RNA amplification than more dilute pools. The reason for these anomalous results is unknown. Overall, the feeding status of the flies included may have impacted the presented results. In any case, any use of stable flies as an effective surveillance tool will depend on identifying and managing sources of variation.
Our results indicate that both dilution of BVDV-fed flies with BVDV-free flies and time after feeding increased the number of RT-rtPCR cycles needed to detect BVDV RNA. The dilution of BVDV resulting from lower numbers of BVDV-fed flies can be expected to have a direct effect on the Ct values for the PCR assay. The delay associated with flies tested 1, 2, or 3 d after their last exposure to BVDV-PI blood may have increased Ct values either by degradation of viral RNA or through dilution associated with consuming additional blood from BVDV-free cattle. Given our results, detecting BVDV RNA in stable flies that have fed on a PI calf may be most probable immediately following the feeding event. Sensitivity of the technique may be better under field conditions if flies can be captured immediately after feeding given that a 24-h delay existed between the final PI blood meal and PCR testing. Our findings are consistent with a 2011 study wherein BVDV was found in horn fly (Haematobia irritans) homogenates for up to 48 h after collection from PI calves. 1
Supplemental Material
Supplementary_material – Supplemental material for Detection of bovine viral diarrhea virus in stable flies following consumption of blood from persistently infected cattle
Supplemental material, Supplementary_material for Detection of bovine viral diarrhea virus in stable flies following consumption of blood from persistently infected cattle by Jaden M. Carlson, Brian L. Vander Ley, Sang I. Lee, Dale M. Grotelueschen, Paul H. Walz, Aspen M. Workman, Michael P. Heaton and David J. Boxler in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The use of product and company names is necessary to accurately report the methods and results; however, the United States Department of Agriculture (USDA) neither guarantees nor warrants the standard of the products. The use of names by the USDA implies no approval of the product to the exclusion of others that may also be suitable. The USDA is an equal opportunity provider and employer.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
