Abstract
Accurate and timely identification of infectious etiologies is of great significance in veterinary microbiology, especially for critical diseases such as strangles, a highly contagious disease of horses caused by Streptococcus equi subsp. equi. We evaluated a matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) platform for use in species- and subspecies-level identification of S. equi isolates from horses and compared it with an automated biochemical system. We used 25 clinical isolates each of S. equi subsp. equi and S. equi subsp. zooepidemicus. Using the MALDI-TOF MS platform, it was possible to correctly identify all 50 isolates to the species level. Unique mass peaks were identified in the bacterial peptide mass spectra generated by MALDI-TOF MS, which can be used for accurate subspecies-level identification of S. equi. Mass peaks (mass/charge, m/z) 6,751.9 ± 1.4 (mean ± standard deviation) and 5,958.1 ± 1.3 were found to be unique to S. equi subsp. equi and S. equi subsp. zooepidemicus, respectively. The automated biochemical system correctly identified 47 of 50 of the isolates to the species level as S. equi, whereas at the subspecies level, 24 of 25 S. equi subsp. equi isolates and 22 of 25 S. equi subsp. zooepidemicus isolates were correctly identified. Our results indicate that MALDI-TOF MS can be used for accurate species- and subspecies-level identification of S. equi.
Keywords
Introduction
Streptococcus equi is an important equine pathogen classified under Lancefield group C streptococci with 2 subspecies of major clinical relevance in horses: S. equi subsp. equi and S. equi subsp. zooepidemicus. S. equi subsp. equi is the etiologic agent of strangles, a highly contagious and severe nasopharyngeal disease of horses.3,13,19,22 S. equi subsp. zooepidemicus causes respiratory tract infections in foals and uterine infections in mares. 6 Another member of Lancefield group C, S. dysgalactiae subsp. equisimilis, is an opportunistic pathogen of horses.11,21,20 The subspecies of S. equi are closely related and are often not reliably identified even by 16S ribosomal DNA sequencing.2,9 Classical sugar fermentation testing is the gold standard for subspecies-level identification of S. equi, and is routinely employed in veterinary diagnostic laboratories. 16 However, sugar fermentation tests can be time-consuming and cause undue delay in obtaining a confirmatory diagnosis.2,8
Although identification of microorganisms using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) has been reported since the1990s, routine implementation as an analytical tool in diagnostic microbiology laboratories is very recent.12,15 The major advantage of MALDI-TOF MS platforms compared to conventional automated biochemical systems is the rapid turnaround time in obtaining bacterial identifications. Bacterial identification by MALDI-TOF MS is based on the generation of unique protein mass spectra profiles from whole cell bacterial samples. Ribosomal proteins form a major component of the mass spectra and have been recognized to produce species-specific spectral patterns that can be used as biomarkers for bacterial identification. 18 Because ribosomal proteins are subject to less variation, bacterial culture conditions have very little effect on the data generated.15,18 The successful use of MALDI-TOF MS for species- and subspecies-level identification of different bacteria has been reported.1,17 A 2015 study reported the identification of equine S. equi isolates to the subspecies level using MALDI-TOF MS. 10 We evaluated a MALDI-TOF MS platform for use in species- and subspecies-level identification of S. equi isolates from horses and compared it with an automated biochemical system.
Materials and methods
Bacterial isolates and culture conditions
Fifty S. equi isolates, 25 each of S. equi subsp. equi and S. equi subsp. zooepidemicus, were used in this study. All bacterial isolates were obtained from equine clinical specimens submitted for testing to the Oklahoma Animal Disease Diagnostic Laboratory (Stillwater, Oklahoma). Bacteria were grown on 5% sheep blood agar (Hardy Diagnostics, Santa Maria, CA) at 37°C in 5% CO2. Classification as S. equi subsp. equi or S. equi subsp. zooepidemicus was based on Lancefield grouping (StrepPRO grouping kit, Hardy Diagnostics, Santa Maria, CA) and classical sugar fermentation tests (maltose, sorbitol, trehalose, salicin, and lactose). 16 Fresh overnight cultures were used for all biochemical and MALDI-TOF MS testing.
Bacterial identification by automated biochemical testing system
The TREK Sensititre automated bacterial identification system (Thermo Fisher Scientific, Oakwood Village, OH) was used for biochemical testing. Bacterial colonies grown overnight were inoculated into a gram-positive identification plate (Thermo Fisher Scientific) using an automated inoculation delivery system (Thermo Fisher Scientific), according to the manufacturer’s recommendations. Plates were incubated for 22 h at 37°C. An autoreader (Thermo Fisher Scientific) was used to optically read plates, and bacterial identification was made by the on-board software. As per the manufacturer’s guidelines, probability scores of >90%, >80%, and >60% denote excellent, good, and acceptable levels of confidence in the results, respectively. Probability scores <60% indicate low selectivity or low result confidence.
Bacterial identification by MALDI-TOF MS
The Microflex LT MALDI-TOF mass spectrometer (Bruker Daltonics, Billerica, MA) was used in this study. The mass spectrometer was calibrated for molecular weights with a range of 3,637–16,952 Da prior to sample testing using the bacterial test standard (Bruker Daltonics), as per manufacturer’s recommendations. For sample testing, bacterial colonies were applied to a spot on the MALDI-TOF MS target plate and overlaid with freshly made 70% formic acid (Sigma-Aldrich, St. Louis, MO) and α-cyano-4-hydroxycinnamic acid matrix solution (Bruker Daltonics), following the manufacturer’s recommended protocol. Sample peptide spectra were collected using commercial software (Bruker Daltonics). The spectra were captured in positive linear mode in a mass range of 2–20 kDa, with a laser frequency of 60 Hz (IS1: 20 kV; IS2: 18 kV; lens: 6 kV; extraction delay time: 100 ns). Spectra were acquired in automatic mode, by accumulating a maximum of 240 profiles (6 × 40 laser shots from different positions of the target spot). Bacterial identification was performed using default settings of the software (Biotyper RTC v. 3.1 and microbial peptide mass spectra database v. 4.0.0.1, Bruker Daltonics) provided with the Microflex LT system. A score value of 0–3 is assigned by the software to each bacterial identification based on similarity of the acquired spectra to reference spectra in the database. As per the manufacturer’s guidelines, scores >2.3 indicate highly probable species classification, whereas scores >2 indicate secure genus and probable species classification.
Mass spectra analysis for subspecies-specific spectral signatures
Peptide mass spectra generated by MALDI-TOF MS were analyzed using FlexAnalysis software v. 3.4 provided with the Microflex LT (Bruker Daltonics) to identify unique spectral peaks. A molecular mass list was generated for each of the bacterial isolates tested. Only mass peaks with intensity >2,000 arbitrary units were selected and compared to identify unique spectral signatures.
Validation of mass peaks
A second MALDI-TOF MS platform of the same model (Microflex LT, Bruker Daltonics), located at the Animal Health Diagnostic Center (AHDC; Cornell University, Ithaca, New York), was also used to analyze and confirm unique peptide mass peaks that were identified for the 2 S. equi subspecies. Peptide mass spectra from 50 S. equi isolates (25 each of S. equi subsp. equi and S. equi subsp. zooepidemicus) that were independently isolated from equine clinical specimens submitted to AHDC were analyzed following similar protocols as described above.
Confirmation of S. equi subspecies by PCR
S. equi isolates, which differed in their subspecies identification on the automated biochemical platform and the MALDI-TOF MS system, were subsequently tested by PCR following a previously published protocol 2 with minor modifications. Total DNA was extracted from bacterial colonies using the DNeasy blood and tissue kit (Qiagen, Valencia, CA), following the manufacturer’s recommended protocol. For each S. equi isolate, 2 separate real-time PCR reactions were performed targeting either the sodA gene or seeI gene. 2 Both PCRs were performed in a 25-µL final reaction volume. The sodA PCR reaction mix contained 0.6 µM each of the forward and reverse primers (Integrated DNA Technologies, Coralville, IA), 0.2 µM probe (Promega, Madison, WI), and 2 µL of template DNA. The seeI PCR reaction mix contained 0.9 µM each of the forward and reverse primers (Integrated DNA Technologies), 0.2 µM hydrolysis probe (Promega), and 2 µL of template DNA. For both PCR assays, 1× buffer (Promega) and 0.5 µL of Taq DNA polymerase (Promega) were used. The thermocycling conditions were set at 95°C for 8 min, followed by 45 cycles of 95°C for 20 s, and 60°C for 60 s. The PCR was performed on a real-time PCR system (Applied Biosystems, Thermo Scientific, Foster City, CA).
Results
Identification of isolates by automated biochemical system
The TREK Sensititre system (Thermo Fisher Scientific) identified 24 of 25 S. equi subsp. equi isolates, with a probability score range of 40–100%. One isolate was misidentified as Streptococcus pneumoniae, with a probability score of 63%. The automated system identified 22 of 25 S. equi subsp. zooepidemicus isolates, with a probability score range of 99–100%. Two isolates were misidentified as S. pneumoniae, with a probability score of 100%, whereas 1 isolate was misidentified as S. equi subsp. equi, with a probability score of 61%. Identities of the 4 isolates that were misidentified on the automated system were confirmed by PCR.
Identification of isolates by MALDI-TOF MS
The Microflex LT MALDI-TOF mass spectrometer (Bruker Daltonics) correctly identified all isolates tested as S. equi. Among the S. equi subsp. equi isolates tested, 22 of 25 were correctly identified to the subspecies level, with a score range of 2.3–2.6, whereas 3 isolates were misidentified as S. equi subsp. zooepidemicus, with scores of 2.2, 2.4, and 2.5 (Table 1). The 3 isolates misidentified as S. equi subsp. zooepidemicus by the Microflex LT platform were further tested and confirmed as S. equi subsp. equi by PCR. The MALDI-TOF platform correctly identified all (25 of 25) S. equi subsp. zooepidemicus isolates, with a score range of 2.3–2.6 (Table 2).
Comparison of TREK Sensititre system (Thermo Fisher Scientific) and the Microflex LT matrix-assisted laser desorption/ionization time-of-flight mass spectrometer (Bruker Daltonics) for identification of 25 isolates of SESE (n = 25).
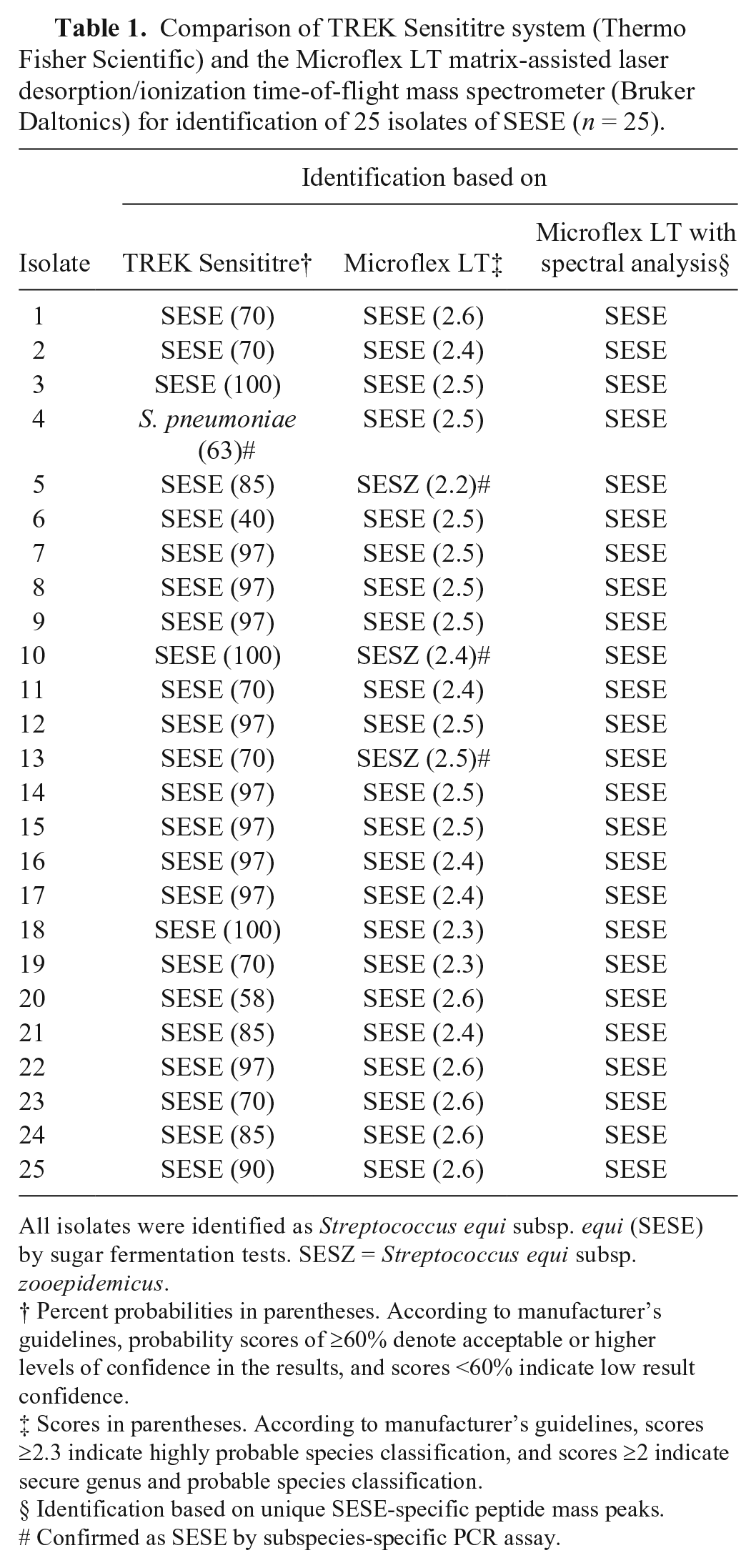
All isolates were identified as Streptococcus equi subsp. equi (SESE) by sugar fermentation tests. SESZ = Streptococcus equi subsp. zooepidemicus.
Percent probabilities in parentheses. According to manufacturer’s guidelines, probability scores of ≥60% denote acceptable or higher levels of confidence in the results, and scores <60% indicate low result confidence.
Scores in parentheses. According to manufacturer’s guidelines, scores ≥2.3 indicate highly probable species classification, and scores ≥2 indicate secure genus and probable species classification.
Identification based on unique SESE-specific peptide mass peaks.
Confirmed as SESE by subspecies-specific PCR assay.
Comparison of TREK Sensititre system (Thermo Fisher Scientific) and the Microflex LT matrix-assisted laser desorption/ionization time-of-flight mass spectrometer (Bruker Daltonics) for identification of 25 isolates of SESZ (n = 25).
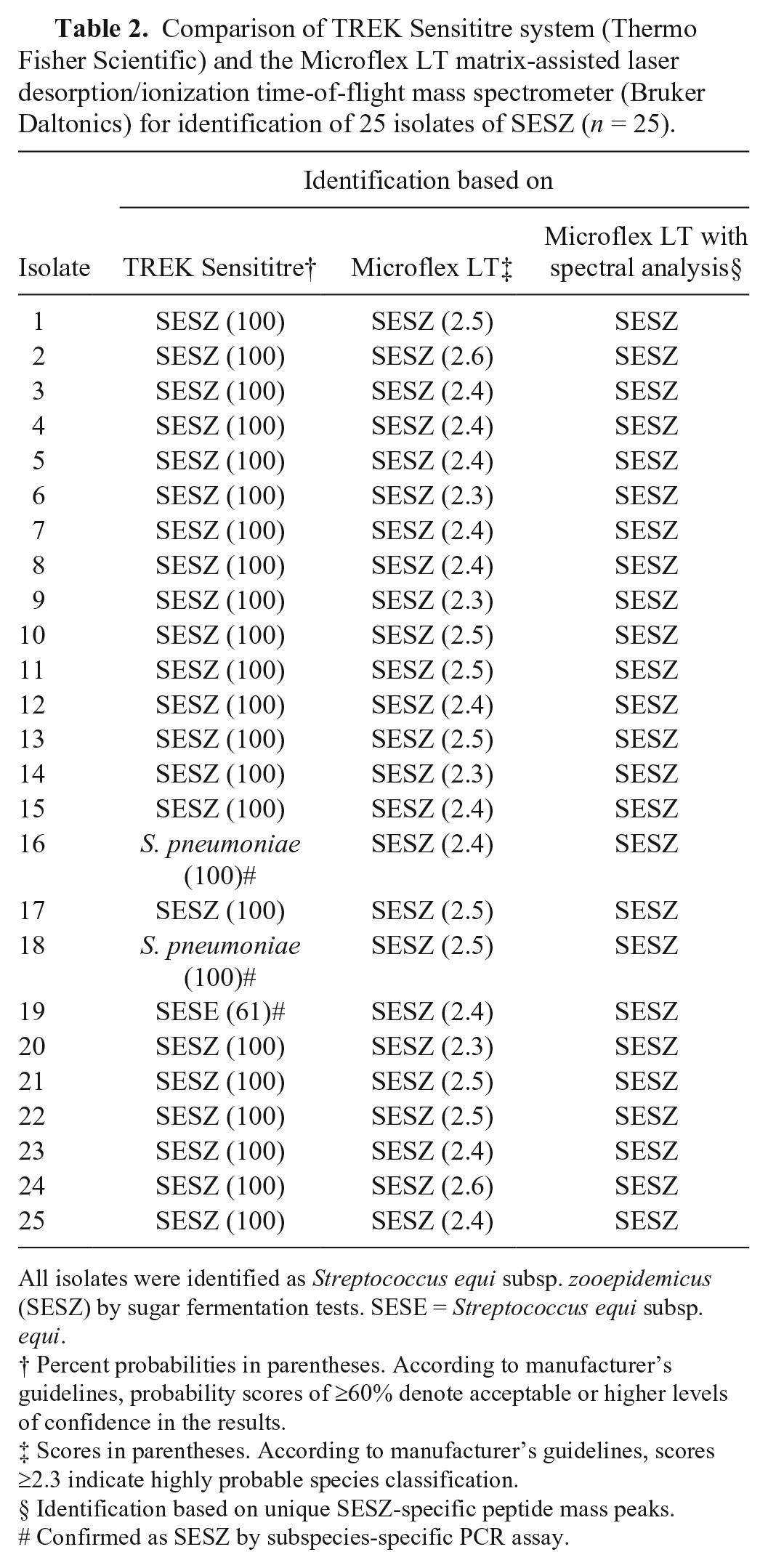

All isolates were identified as Streptococcus equi subsp. zooepidemicus (SESZ) by sugar fermentation tests. SESE = Streptococcus equi subsp. equi.
Percent probabilities in parentheses. According to manufacturer’s guidelines, probability scores of ≥60% denote acceptable or higher levels of confidence in the results.
Scores in parentheses. According to manufacturer’s guidelines, scores ≥2.3 indicate highly probable species classification.
Identification based on unique SESZ-specific peptide mass peaks.
Confirmed as SESZ by subspecies-specific PCR assay.
Analysis of protein mass spectra generated by MALDI-TOF MS
Several subspecies-specific peptide mass peaks were detected in the protein mass spectra generated from the different S. equi isolates tested. Based on the parameters described in the Materials and methods section, we identified 2 subspecies-specific peptide mass peaks at 6,751.9 ± 1.4 mass/charge (m/z; mean ± standard deviation; n = 25) and 5,958.1 ± 1.3 m/z (n = 25) that were prominent and consistently present in the peptide mass spectra of all S. equi subsp. equi and S. equi subsp. zooepidemicus isolates tested, respectively (Fig. 1; Table 3). Two additional unique mass peaks were also identified at 5,915 ± 1.3 m/z (n = 24) and 6,739.4 ± 1.4 m/z (n = 23) for the S. equi subsp. equi and S. equi subsp. zooepidemicus isolates tested, respectively. However, the 2 latter mass peaks were not consistently present and were not detected in 3 of the 50 isolates tested (Table 2).
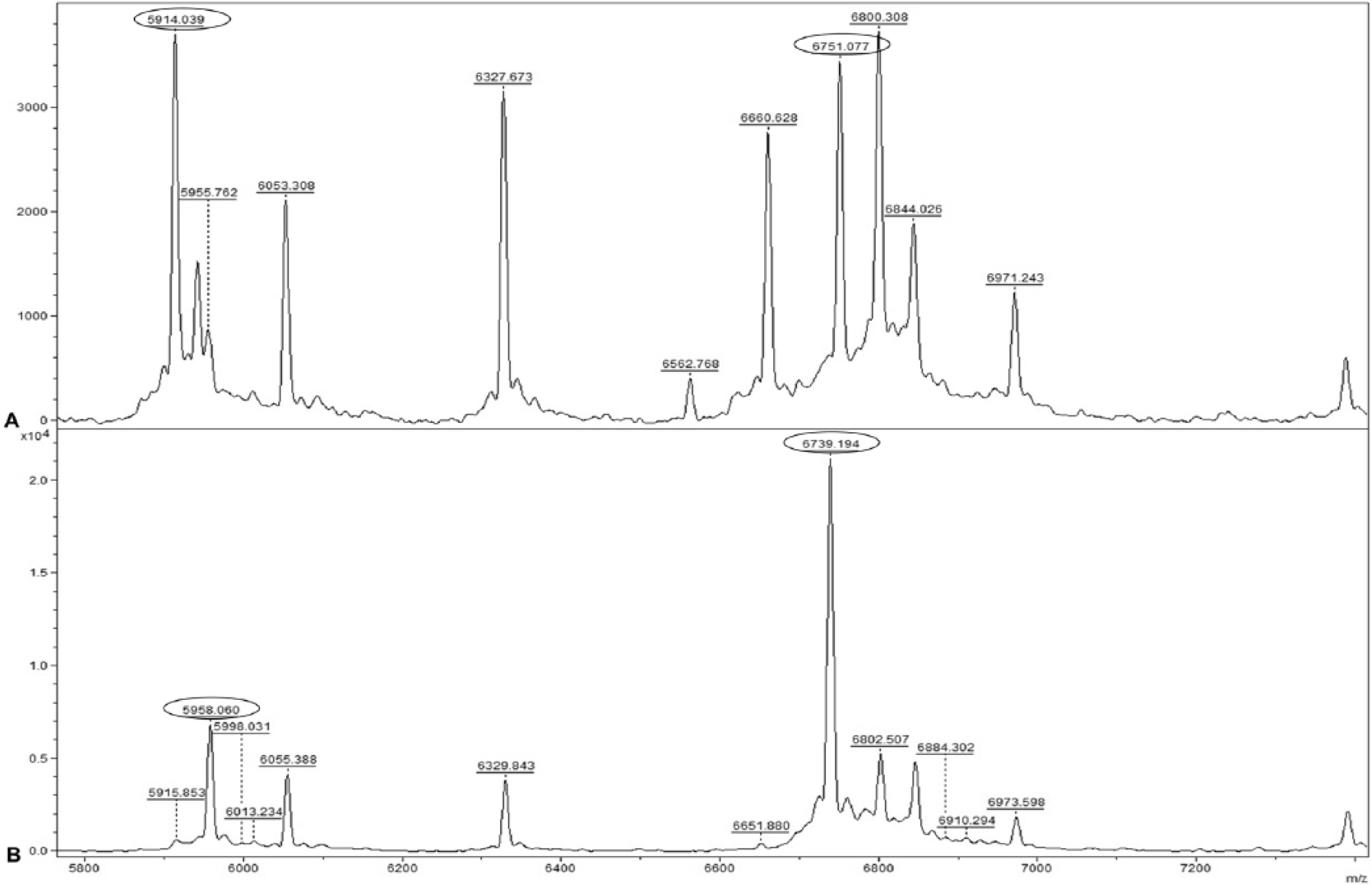
Representative peptide mass spectra of (A) Streptococcus equi subsp. equi and (B) Streptococcus equi subsp. zooepidemicus showing the 4 unique peptide mass peaks for the respective isolates (circled). The x-axis and y-axis represent mass/charge (m/z) and peak intensity in arbitrary units (a.u.), respectively. For panel B, the y-axis is in 104 a.u.
Frequency of occurrence of subspecies-specific peptide mass peaks in the different isolates of SESE (n = 25) and SESZ (n = 25) tested by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry.
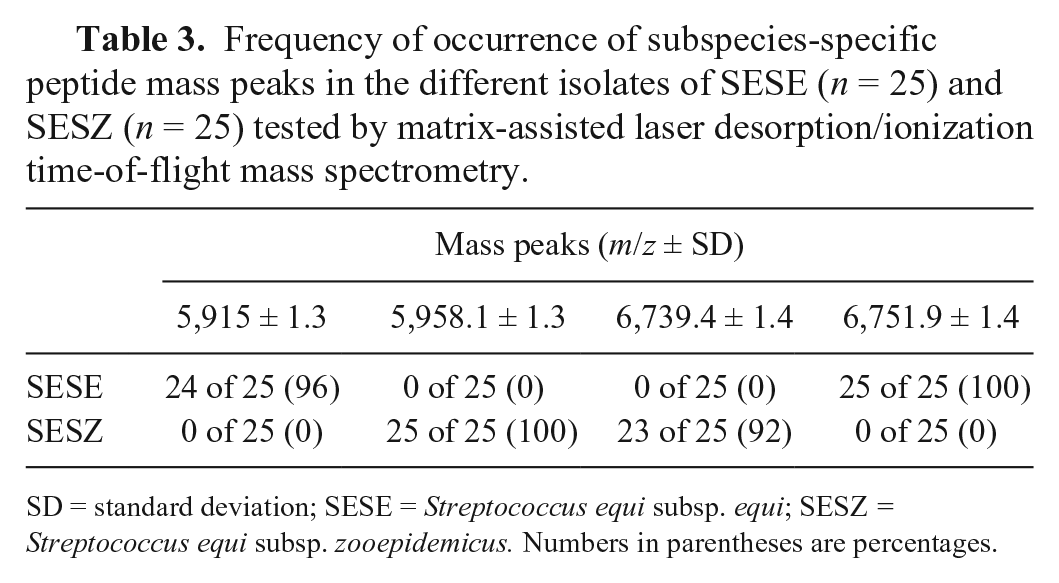

SD = standard deviation; SESE = Streptococcus equi subsp. equi; SESZ = Streptococcus equi subsp. zooepidemicus. Numbers in parentheses are percentages.
Validation of mass peaks
Subspecies-specific mass peaks (m/z) corresponding to 6,751.9 ± 1.4 and 5,958.1 ± 1.3, which were identified in all of the isolates of S. equi subsp. equi and S. equi subsp. zooepidemicus, respectively, were also detected on a second MALDI-TOF MS platform at AHDC. Mass peaks (m/z) corresponding to 5,915 ± 1.3 was detected in all 25 S. equi subsp. equi isolates; however, the 6,739.4 ± 1.4 mass peak was detected only in 21 of 25 isolates (84%) of S. equi subsp. zooepidemicus.
Discussion
The MALDI-TOF MS platform has been previously demonstrated to successfully identify beta-hemolytic Streptococcus isolates from humans and animals.4,10 MALDI-TOF MS was found to be superior in accuracy at species-level identification when compared to an automated biochemical-based bacterial identification system. 4 As well, MALDI-TOF MS has been used for the subspecies-level identification of other pathogens, such as Salmonella sp. and Francisella sp.7,17
We evaluated MALDI-TOF MS for identification of S. equi isolates from horses. Lancefield grouping and classical sugar fermentation tests were used to establish the correct identity of all isolates tested. Selected isolates were also tested using a S. equi–specific PCR. The MALDI-TOF MS platform showed 100% accuracy for species-level identification, whereas automated biochemical system only correctly identified 47 of 50 isolates tested, with an additional 2 isolates reporting below the acceptable probability score. At subspecies level, although the percent probability of identification for some of the isolates was low (Table 1), the automated biochemical system identified more S. equi subsp. equi isolates (24 of 25) compared to the MALDI-TOF MS platform (22 of 25). However, accurate subspecies-level identification by MALDI-TOF MS was possible on all isolates tested when the presence of unique peptide mass peaks was also taken into consideration. Multiple peptide mass peaks were identified that were unique to the 2 subspecies of S. equi. In addition, the analytical repeatability of these subspecies-specific unique mass peaks was also tested and validated on a different MALDI-TOF MS platform.
The use of MALDI-TOF MS for differentiating subspecies of S. equi was reported in 201510 using a different instrument model (Vitek MS RUO, bioMérieux, Marcy l’Etoile, France). As in our study, subspecies-specific unique mass peaks were identified. The major subspecies-specific peaks identified included mass peaks (m/z) for S. equi subsp. equi (6,659) and S. equi subsp. zooepidemicus (3,368; 5,975; 6,735; and 10,397). None of these peaks were consistently detected in the isolates tested in our study. Furthermore, the 2 consistently detected subspecies-specific peptide mass peaks (6,751.9 ± 1.4 and 5,958.1 ± 1.3) found in the bacterial isolates tested in our study were not present or consistent in the above reported study. This discrepancy in results may be attributed to differences in the composition of the reagents used for bacterial isolate processing, differences in instrument parameters, and possibly other intrinsic design differences associated with different MALDI-TOF MS platform models.5,14
MALDI-TOF MS is a promising technology for the rapid and reliable identification of S. equi subsp. equi and S. equi subsp. zooepidemicus isolates. Bacterial peptide spectrum analysis for the presence of subspecies-specific mass peaks will help in the accurate identification of S. equi isolates and can significantly improve turnaround time in bacteriology laboratories and reduce costs associated with additional confirmatory testing protocols.
Footnotes
Acknowledgements
We thank Brooke Horinek and LaRinda Holland, OADDL, for performing the S. equi PCR assays.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors declared that they received no financial support for their research and/or authorship of this article.
