Abstract
We compared different methods for their ability to isolate Mycobacterium bovis from tissue samples from animals with lesions resembling bovine tuberculosis. In the first trial, M. bovis was isolated from 86 of 200 tissue samples that were cultured using 2 liquid media, BACTEC 12B and BBL mycobacteria growth indicator tube (MGIT), and a solid medium, Middlebrook 7H11 supplemented with pyruvate (7H11P). M. bovis was isolated from 2 samples with MGIT but not BACTEC 12B. M. bovis was isolated from 9 samples with BACTEC but not MGIT; these 9 samples came from the North Canterbury/Marlborough region of New Zealand. The proportion of tissues from which M. bovis was isolated with BACTEC 12B or MGIT and the mean time for isolation was different for samples from the North Canterbury/Marlborough region but not the rest of New Zealand. In the second trial, M. bovis was isolated from 401 of 1,033 tissues that were cultured using MGIT, Middlebrook 7H9 broth, or solid 7H11P. The proportion of isolates of M. bovis and the mean time for their isolation with MGIT was different for the North Canterbury/Marlborough and the rest of New Zealand. The reason for this difference was not determined but may be related to the genotypes present in this region. Genotyping using variable number tandem repeats (VNTRs) of 197 isolates of M. bovis revealed that the 44 isolates from North Canterbury/Marlborough were represented by 2 closely related VNTR types that were not found in 153 isolates from the remainder of New Zealand.
Introduction
The ability to isolate Mycobacterium bovis from tissue samples is an essential component of a bovine tuberculosis (TB) control program.4,9 Culture provides a postmortem test that is both sensitive and specific, qualities that support the claims for culture being the “gold standard” test in a bovine TB control program. 18 Although DNA amplification methods are increasingly used in bovine TB control programs, culture provides isolates of M. bovis that are necessary for DNA typing methods, such as restriction endonuclease analysis, variable number tandem repeats (VNTRs), and whole genome sequencing. Typing produces unique epidemiologic information on the spread of infection, and most importantly on the role of wildlife reservoirs in the persistence of infection in cattle and farmed deer. 1
Classical methods for the isolation of M. bovis used a solid medium, which had the major disadvantage of requiring media to be incubated for up to12 wk. 4 Liquid culture systems have superseded the use of solid media and have enabled culture times to be reduced by at least one-half. 17 The principal disadvantage of liquid culture systems is that they are prone to overgrowth by rapidly growing microbes. 17 For many years, the BACTEC 12B liquid culture systema,b was used by human mycobacterial laboratories for the isolation of M. tuberculosis 5 and by veterinary laboratories for the isolation of M. bovis. 11 The disadvantages of the BACTEC 12B were that it used radioactive media to detect growth, and syringes and needles were required for inoculating the vials.2,10 In order to address these disadvantages, in the 1990s, a new liquid culture system known as the BBL mycobacteria growth indicator tube (MGIT)c-e became available.2,10,14,15 A fluorescence compound for the detection of growth is present in the MGIT and a needle is not required for inoculation. The use of MGIT has also been driven by the fact that BACTEC 12B medium is no longer available. 19 We focused on the evaluation of MGIT as a liquid system for the isolation of M. bovis from cattle and farmed deer.
Materials and methods
The tissue samples for our study were those submitted for the isolation of M. bovis as part of the routine procedures of a bovine TB control program. The samples were principally lymph nodes collected at abattoirs from throughout New Zealand from cattle with macroscopic lesions that were either suspicious or typical of those caused by M. bovis. A small number of samples were from farmed deer and wildlife.
All samples were processed using the same procedure. Tissues were homogenized using grinders f and then decontaminated with 1.5% NaOH. Decontaminated tissue homogenates were neutralized, centrifuged, and then inoculated into the various media. The volume of processed homogenate used to inoculate liquid and solid media was 0.5 mL and 0.2 mL, respectively.
BACTEC 12B medium was prepared as described previously 11 by adding the recommended amount of the antibiotic supplement. b The MGIT c - e medium was formulated according to the manufacturer’s instructions. Middlebrook 7H9 (7H9) g broth consisted of 4 mL of 7H9 supplemented with sodium pyruvate and oleic albumin dextrose and the addition of the antimicrobial additive e used in MGIT (at the same concentration as present in MGIT). The solid Middlebrook 7H11 (7H11P) h medium was supplemented with pyruvate and prepared as described previously. 8 All media were incubated at 37°C. Maximum times for incubation were 30 d for BACTEC 12B vials and liquid 7H9, 42 d for MGITs, and 90 d for solid media. The BACTEC 12B vials were examined twice per week for evidence of microbial growth in a BACTEC 460 machine, i and MGITs were incubated in a MGIT 320 j machine that read the samples every hour. A growth index >15 was considered positive for BACTEC 12B and >75 for MGITs. Incubation of BACTEC 12B vials with positive indices was continued until they exceeded 800. Positive BACTEC 12B and MGIT cultures were checked for contamination by inoculating and incubating blood agar plates and for mycobacteria by the examination of Ziehl–Neelsen-stained smears. Where a pure growth of salmon-pink colonies was found, a Gram stain, and then catalase, urease, and glucose utilization tests were carried out to identify Rhodococcus equi. After 30 d of incubation, 7H9 cultures were inspected for mycobacteria by the examination of Ziehl–Neelsen-stained smears. A presumptive identification of M. bovis was based on a positive reaction in a M. tuberculosis complex commercial probe k or a M. tuberculosis lateral flow test. l Molecular typing of presumptive M. bovis isolates was carried out using a previously described method 16 that used 9 VNTRs and 2 direct repeats (DR1, DR2, Table 3).
The paired data were analyzed using a McNemar test; the proportion of M. bovis isolated in the different media was compared using a Fisher exact test. The means times to detection were analyzed using analysis of variance.
Results
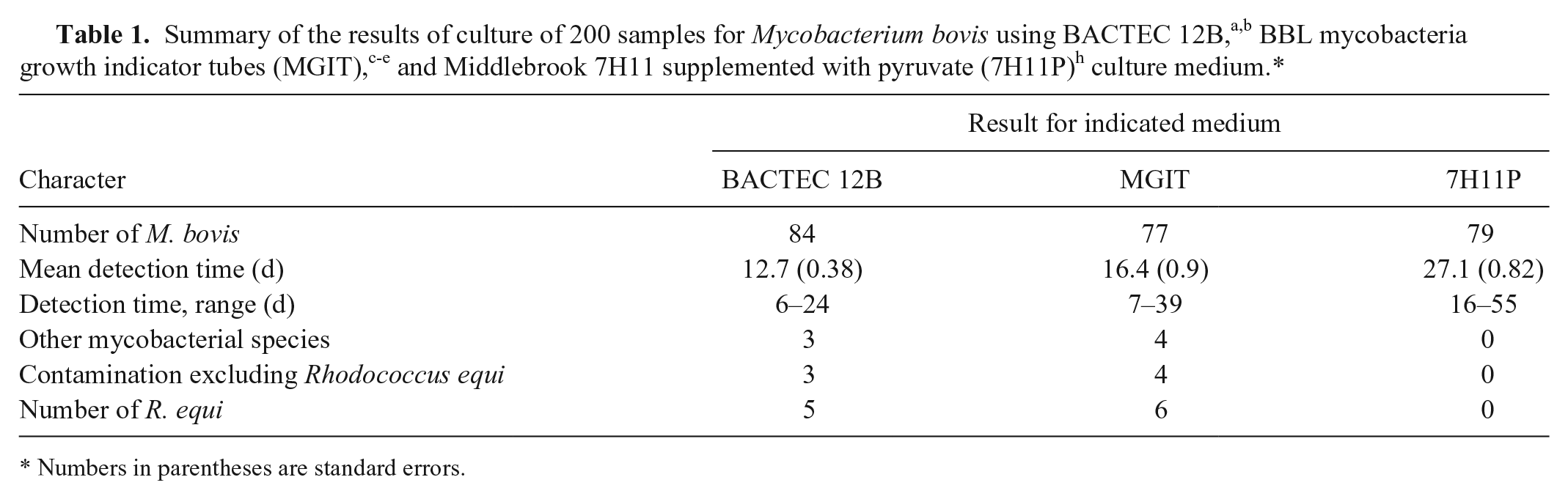
In trial 1, a single vial of BACTEC 12B, MGIT, and solid 7H11P per sample were compared for their ability to isolate M. bovis. A total of 200 tissues, consisting of samples from 183 cattle, 13 farmed deer, 2 cats, and 2 brushtail possums were cultured, and M. bovis was isolated by 1 or more of the methods from 86 of the samples (Table 1). M. bovis was isolated from 2 samples with MGIT but not BACTEC 12B. M. bovis was isolated from 9 samples with BACTEC 12B but not MGIT; these 9 samples came from the North Canterbury/Marlborough region on the northeast coast of the South Island of New Zealand. Of the samples from this region, M. bovis was isolated from 16 samples with BACTEC 12B and only 7 with MGIT. The differences in the number of cultures of M. bovis isolated by MGIT and BACTEC 12B was significant for samples from North Canterbury/Marlborough (p = 0.004, McNemar test) but not for those from the rest of New Zealand. The mean detection times for the 3 media used in trial 1 are summarized in Table 1 and Figure 1.
Numbers in parentheses are standard errors.
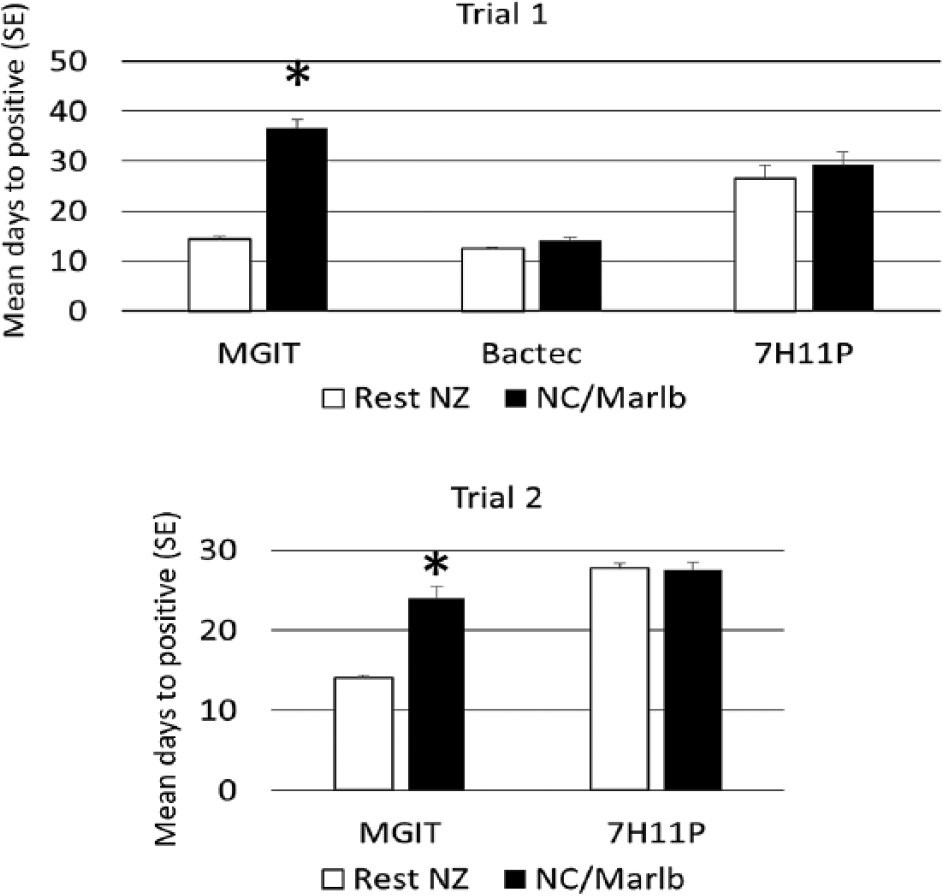
Growth rates for the isolation of Mycobacterium bovis expressed as mean number of days (standard error [SE]) of different culture media. North Canterbury/Marlborough (NC/Marlb) is indicated by open bars, and the rest of New Zealand (NZ) is indicated by closed bars. Asterisk (*) indicates significant difference from the mean of days to positive for the M. bovis strains from NC/Marlb grown on the same culture medium (analysis of variance, p < 0.05). BACTEC = BACTEC 12Ba,b; MGIT = BBL mycobacteria growth indicator tubec-e; 7H11P = Middlebrook 7H11 supplemented with pyruvate. h
Contaminants, as defined by rapidly growing, acid-fast–negative microbes, were isolated from 8 (4%) and 10 (5%) cultures from BACTEC 12B and MGIT. respectively; these results were not statistically significant (McNemar test). Included in these figures were 5 (2.5%) and 6 (3%) pure cultures of R. equi from BACTEC 12B and MGIT, respectively.
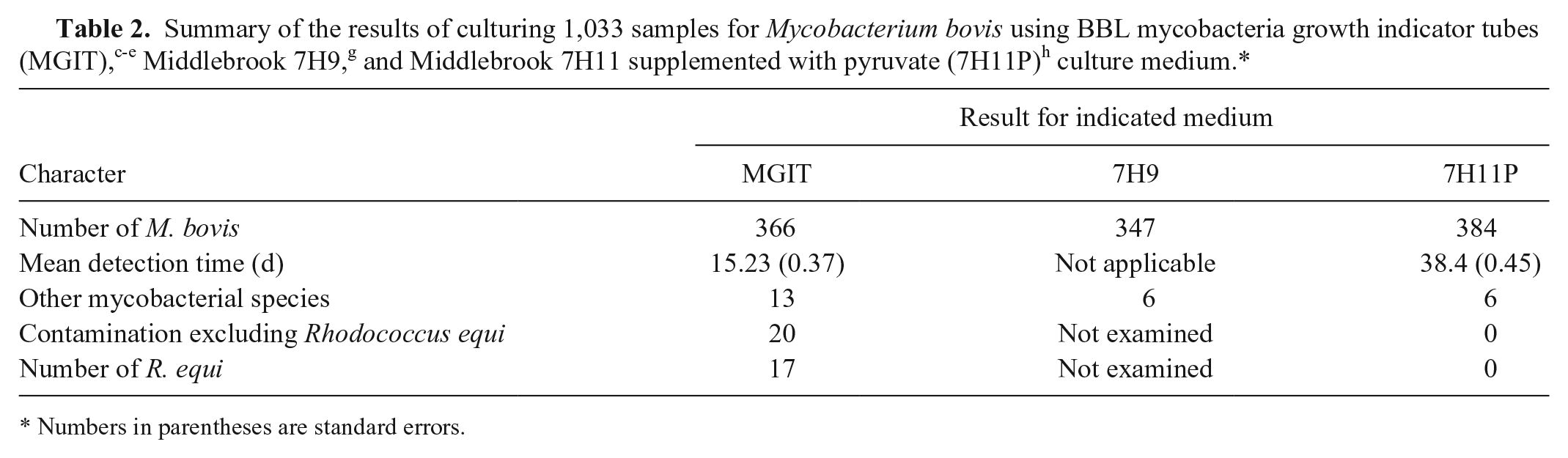
For trial 2, single vials of MGIT, liquid 7H9, and solid 7H11P per sample were compared for their ability to isolate M. bovis. The BACTEC 12B vials were no longer available for the second trial, and liquid 7H9 supplemented with MGIT antimicrobial additive e was used as an alternative. Over a period of 14 mo, M. bovis was isolated by 1 or more of the culture methods from 401 of the 1,033 samples, of which 1,011 came from cattle, 21 from farmed deer, and 1 from a cat (Table 2). There were 67 isolates of M. bovis grown from samples from North Canterbury/Marlborough, and of these, 45 (67%) were isolated with MGIT, 55 (82%) with 7H9, and 65 (97%) with 7H11P. In contrast, 312 of 334 (93.4%) of the M. bovis isolated from all other areas of New Zealand were recovered using MGIT, 292 of 334 (87.4%) for 7H9, and 319 (95.5%) with 7H11P. The difference in the proportion of M. bovis isolated with MGIT from the 2 regions is highly significant (p < 0.001, Fisher exact test). The mean detection times for MGIT and 7H11P are summarized in Figure 1. Rapidly growing, non–acid-fast bacteria were isolated from 42 MGIT cultures (4%). Included in this figure were 23 pure cultures (2.2%) of R. equi. There were 2 samples for which R. equi was isolated from the MGIT medium, but M. bovis was isolated from solid 7H11P.
Numbers in parentheses are standard errors.
Using VNTR, we typed 197 isolates of M. bovis from trial 2, comprising 153 from the rest of New Zealand and 44 from North Canterbury/Marlborough. Isolates for typing were selected from newly infected herds and from herds of interest for disease control purposes. The isolates from North Canterbury/Marlborough had 2 VNTR patterns that were not present in isolates from the remainder of New Zealand (Table 3). The isolates from the remainder of New Zealand were represented by 22 different VNTR patterns.
Molecular typing using variable number tandem repeats (VNTRs) of 197 isolates of Mycobacterium bovis recovered from trial 2.*
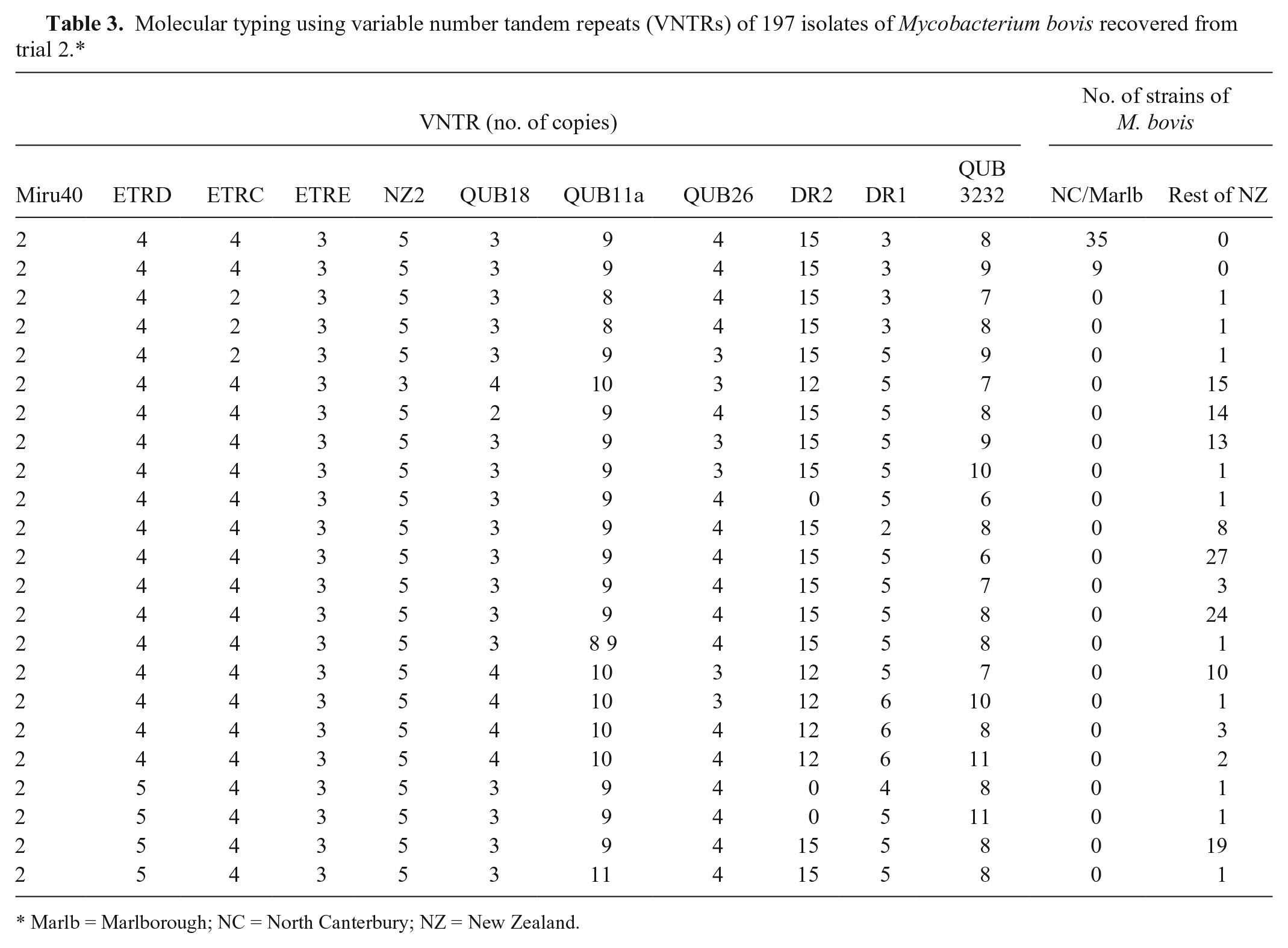
Marlb = Marlborough; NC = North Canterbury; NZ = New Zealand.
Discussion
The MGIT culture system is widely used for the isolation of mycobacteria, especially in human laboratories for the isolation of M. tuberculosis. 17 Although there are limited numbers of reports on the efficacy of the MGIT system for the isolation of M. bovis, 2 studies have reported that this culture system was comparable to using BACTEC 12B medium.11,17 With one exception, the results of our study are in agreement with previous findings. The success of using MGIT for isolating M. bovis and the time to positivity were both significantly less for samples coming from the North Canterbury/Marlborough region compared with those coming from the remainder of New Zealand. The brushtail possum (Trichosurus vulpecula) is the principal wildlife reservoir host of M. bovis in this area of New Zealand. 12 The reason for this regional difference in the ability of MGIT to isolate M. bovis is unknown but may reflect the unique genotypes in this region. The unique genotypes were defined using VNTRs and were consistent with previous findings. 16
We based the presumptive identification of the isolates as M. bovis on a positive lateral flow test or a commercial probe that detected the M. tuberculosis complex. Confirmation of selected isolates as M. bovis was based on VNTR typing, which is able to distinguish M. bovis from other members of the M. tuberculosis complex.3,16 On very rare occasions, M. pinnipedii and M. orygis have been isolated from cattle in New Zealand.6,13
The level of contamination as measured by the presence of rapidly growing, nonmycobacterial organisms in BACTEC and MGIT cultures was 4–5% and was similar for both trials. R. equi was isolated in pure culture in >50% of the BACTEC and MGIT cultures that yielded rapidly growing, nonmycobacterial cultures. This organism is of relatively low virulence in cattle but can cause granulomatous lymphadenitis, which macroscopically resembles lesions caused by M. bovis. 7 On 2 occasions in trial 2, R. equi was isolated in MGIT, and M. bovis was isolated from the accompanying solid medium. The level of contamination observed in these trials was at an acceptable level and lower than the levels of ≤24% observed by others. 17 The low level of contamination observed in our study may reflect the care taken in collecting the samples and their rapid delivery to the laboratory.
MGITs are a valuable method for the isolation of M. bovis, but they should be complemented with the use of another medium, such as 7H11P. The additional medium will provide for situations in which MGITs are overgrown by rapidly growing, non–acid-fast bacteria or for subtypes of M. bovis that may grow poorly in MGIT.
Footnotes
Acknowledgements
We thank Dongwen Luo for reviewing the statistical analyses, and Neil Wedlock and Natalie Parlane for their help in the preparation of this article.
a.
BACTEC 12B medium, Becton Dickinson, Sparks, MD.
b.
BACTEC PANTA Plus, Becton Dickinson, Sparks, MD.
c.
BBL MGIT, Becton Dickinson, Sparks, MD.
d.
BACTEC MGIT growth supplement, Becton Dickinson, Sparks, MD.
e.
BBL MGIT PANTA, Becton Dickinson, Sparks, MD.
f.
Tenbroeck tissue grinder, Wheaton Science Products, Millville, NJ.
g.
Middlebrook 7H9, Becton Dickinson, Sparks, MD.
h.
Middlebrook 7H11, Becton Dickinson, Sparks, MD.
i.
ACTEC 460 radiometric system, Becton Dickinson, Sparks, MD.
j.
MGIT 320, Becton Dickinson, Sparks, MD.
k.
AccuProbe Mycobacterium tuberculosis complex culture identification test, Gen-Probe, San Diego, CA.
l.
SD Bioline, TB Sg MPT64 Rapid; Standard Diagnostics, Gyeonggi-do, Republic of South Korea.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding for this project was provided by TBFree New Zealand.