Abstract
We compared the nested internal transcribed spacer (ITS) PCR and the 18S PCR-RFLP (restriction-fragment length polymorphism) pan-trypanosome assays in a cross-sectional survey of bovine trypanosomiasis in 358 cattle in Kwale County, Kenya. The prevalence of trypanosomiasis as determined by the nested ITS PCR was 19.6% (70/358) and by 18S PCR-RFLP was 16.8% (60/358). Of the pathogenic trypanosomes detected, the prevalence of
Trypanosomiasis is a severe chronic wasting disease of domestic animals. Wildlife are also susceptible to infection with trypanosomes,
1
but they are less likely to succumb to the disease
17
and instead act as reservoirs.
14
The important animal trypanosomes include
The distribution of pathogenic animal trypanosomes, affected animal hosts, and mode of transmission.
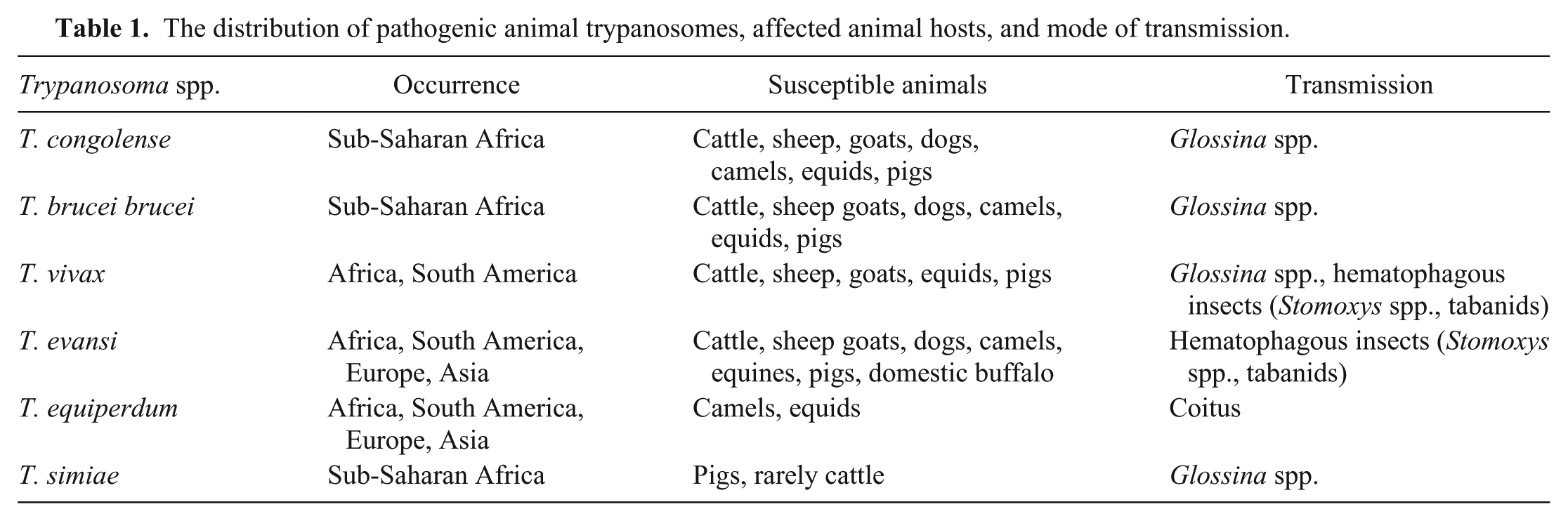

Animal trypanosomiasis causes low milk output, poor carcass quality, inability of animals to provide draught power, and high mortality rates in some cases. 10 Affected countries, in collaboration with international development agencies, have intervened by introducing various control measures, such as vector control, chemoprophylaxis and chemotherapy, and community education. The impact of these interventions requires periodic assessment through surveillance with a robust detection test. Documentation of the performance of detection tests in different geographic locations where animal trypanosomiasis is endemic should be encouraged in order to understand their feasibility as tools for surveillance of the disease.
Detection tests for animal trypanosomiasis include parasitologic, immunologic, and molecular methods, 7 but, for economic reasons, parasitologic techniques are still favored in sub-Saharan Africa despite their lack of sensitivity and labor intensiveness. Undoubtedly, PCR is the most appropriate tool for sensitive screening of animal trypanosomiasis and retrospective analysis. Species-specific PCRs require multiple reactions per sample to detect the different species of trypanosomes, 11 which is overwhelming during large-scale operation. Thus, pan-trypanosome PCRs are the preferred assays for epidemiologic survey targeting multiple species of trypanosomes.3,6,8,15
The ribosomal (r)DNA-based pan-trypanosome PCRs (Table 2) are commonly used for detection of animal trypanosomes; however, only a few of these assays have been comparatively evaluated on field samples.15,18 The nested internal transcribed spacer (ITS) PCR and the 18S PCR-RFLP (restriction-fragment length polymorphism) pair lacks such comparative data. Thus, we compared the 2 assays on field samples obtained from Kwale County, Kenya, where trypanosomiasis is endemic. Kwale County is covered with dense vegetation and all-season streams, consequently providing the required humidity for proliferation of the vectors. The occurrence of the various species of animal trypanosomes in the region has been reported,12,16 thus making the region attractive for evaluation of these tests. Additionally, it was important to understand the current burden of bovine trypanosomiasis in the region as a follow-up to ongoing control efforts.
Different ribosomal (r)DNA-based pan-trypanosome PCR tests for African trypanosomiasis.
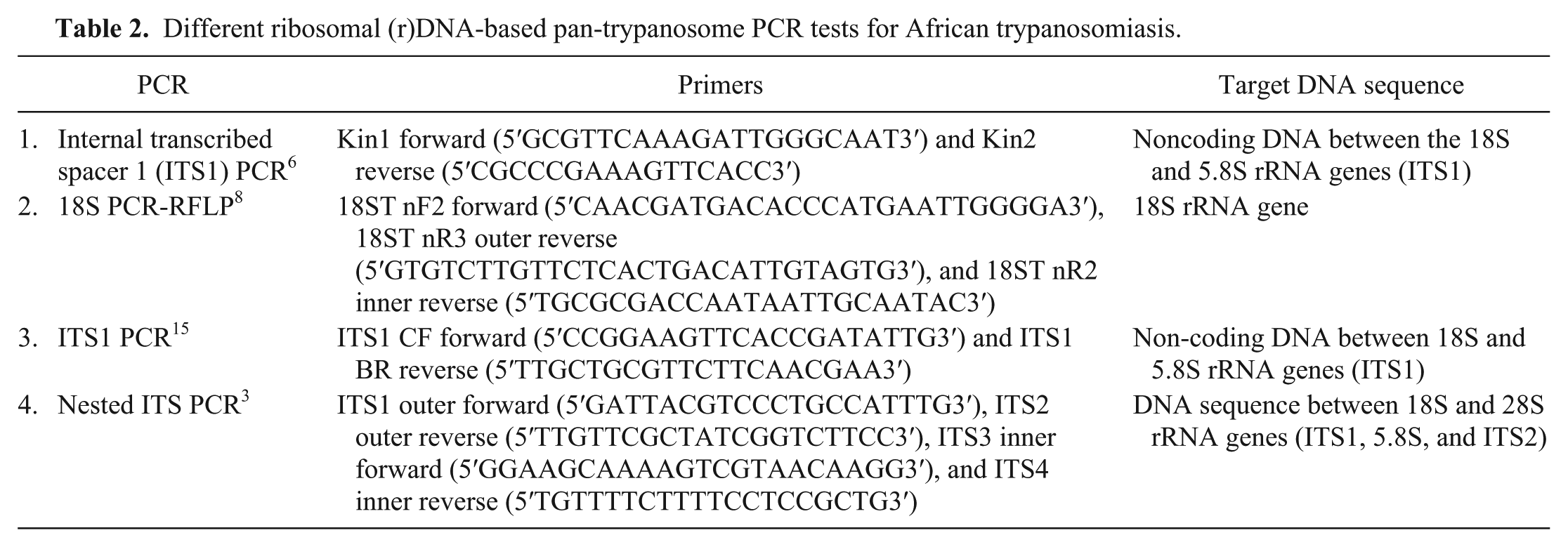

Reference DNA samples were prepared from
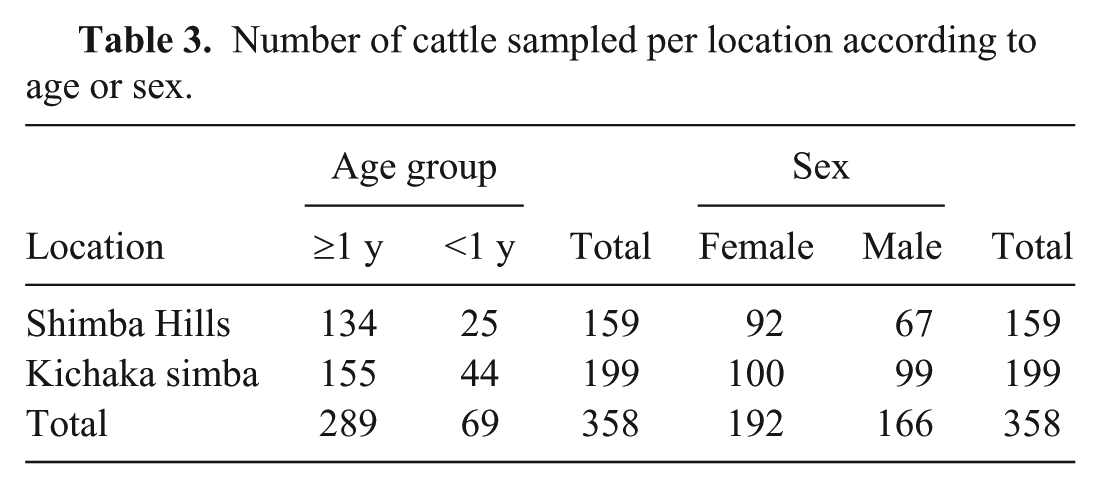
Number of cattle sampled per location according to age or sex.
DNA was extracted from whole blood (10 µL) using a commercial kit b ; the concentration was determined using a spectrophotometer, c and samples were stored at −20°C until analyzed. The samples were examined for trypanosome DNA by the nested ITS PCR as described previously. 3 Briefly, the first run consisted of 5 µL of the template, 1× Taq DNA polymerase buffer, d 0.2 mM of each dNTP, 0.4 µM of each primer (ITS1 and ITS2), 1.25 U Taq DNA polymerase, e and adjusted with distilled water to 25 µL/reaction. For the second run, the product of the first run (1 µL) was seeded in 24 µL of a fresh master mix prepared as above except the ITS3 and ITS4 primers were used in the place of ITS1 and ITS2. The cycle conditions were the following: lid temperature 105°C; initial denaturation 95°C for 5 min; 94°C for 30 s, 57°C for 30 s, 72°C for 60 s for 35 cycles; postcycle 72°C for 10 min; and paused at 12°C. The amplified products of the second run were separated by electrophoresis on 1.5% (w/v) agarose f at 100 V for 40 min. The profiles on gels were scored according to the standard profiles obtained from reference trypanosomes and a previous study. 3
The detection of trypanosome DNA using the 18S PCR-RFLP was performed as described.
8
Briefly, the master mix was prepared from 1× Taq DNA polymerase buffer, 0.2 mM of each dNTP, 0.5 µM forward primer 18ST nF2 and 0.5 µM reverse primer 18ST nR3 for the first PCR, or 0.5 µM reverse primer 18ST nR2 for the semi-nested PCR and 1.25 U Taq DNA polymerase. The master mix was adjusted with distilled water to the required volume. Whereas the first PCR master mix (20 µL) was seeded with an extracted DNA sample (5 µL), the second PCR master mix (24 µL) was seeded with the first PCR product (1 µL). The cycle conditions were the following: lid temperature 105°C; precycle 95°C for 4 min; 92°C for 30 s, 58°C for 45 s, 72°C for 60 s for 40 cycles; postcycle 72°C for 10 min; and paused at 4°C. Amplified products of the second run were separated by electrophoresis on 2% (w/v) agarose. PCR amplicons of ±600 bp were double digested with the restriction enzymes
Conflicting profiles were cross-validated through sequencing of the PCR products. For this, 10 µL of the positive PCR products were processed by incubating with 2 U alkaline phosphatase k and 20 U exonuclease I l on a thermal cycler m programmed at 37°C for 15 min and then at 80°C for 15 min. The sequencing of the processed sample was outsourced to a VIB genetic service facility (University of Antwerp). Homology search in the databank was performed using a bioinformatic software. n
Statistical analysis was performed,
o
and frequencies were compared using the chi-square (χ2) test. Chosen confidence level (α) was 0.05, and
The prevalence of animal trypanosomiasis by the nested ITS PCR was 19.6% (70/358). The nested ITS PCR detected
Single and mixed trypanosome infections detected with the nested internal transcribed spacer (ITS) PCR according to location, animal sex, or age group.*
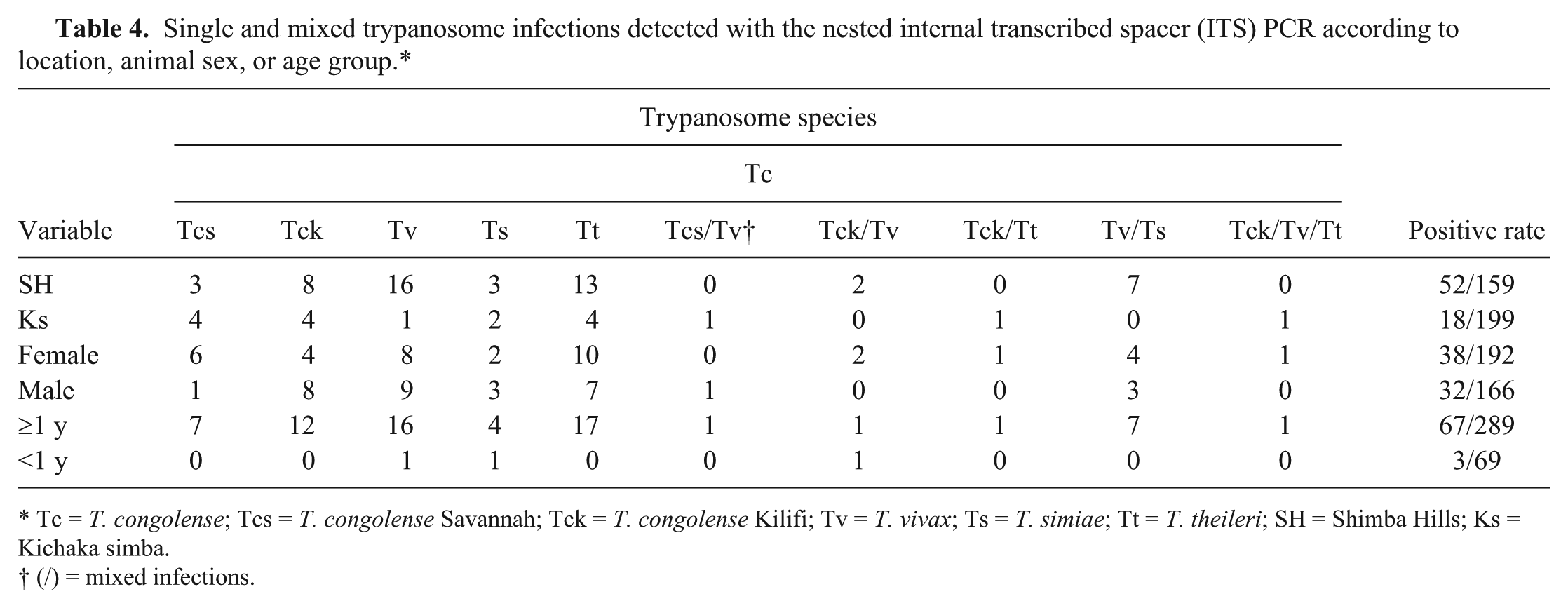
Tc =
(/) = mixed infections.
Single and mixed trypanosome infections detected with the 18S PCR-RFLP (restriction-fragment length polymorphism) according to location, animal sex, or age group.*
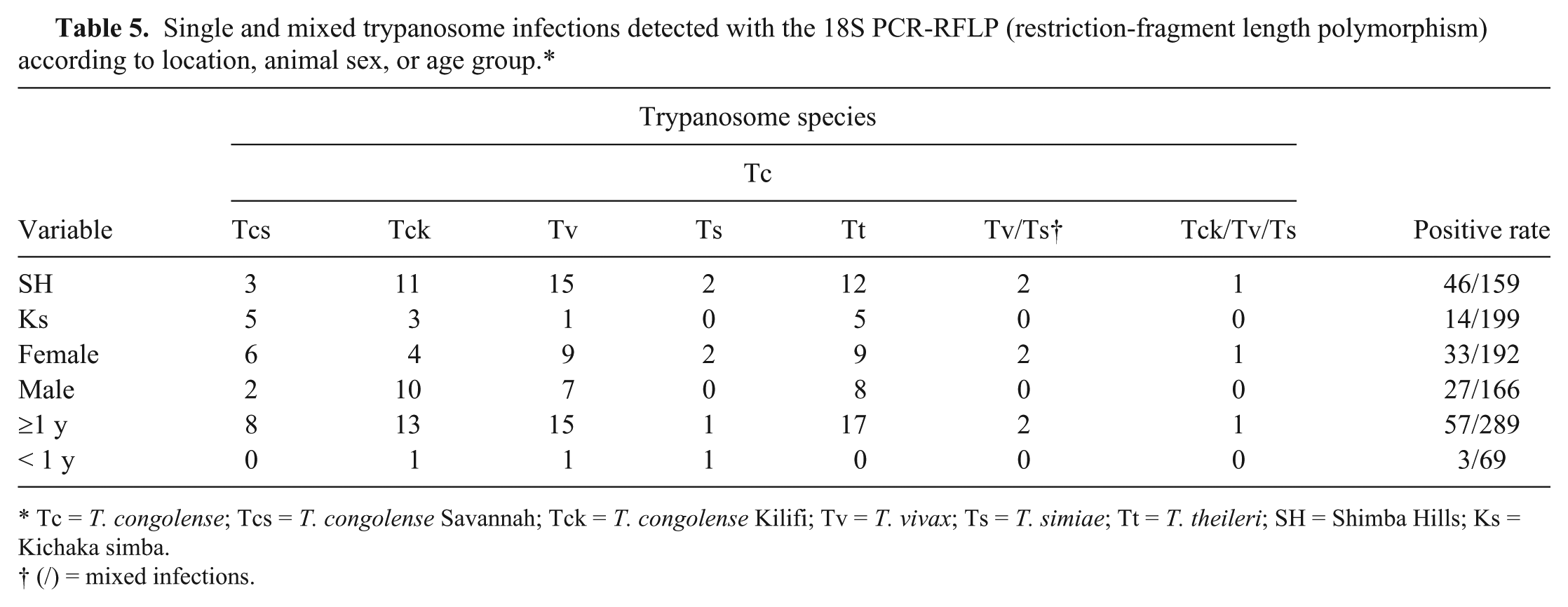
Tc =
(/) = mixed infections.
The agreement between the nested internal transcribed spacer (ITS) PCR and the 18S PCR-RFLP (restriction-fragment length polymorphism).*
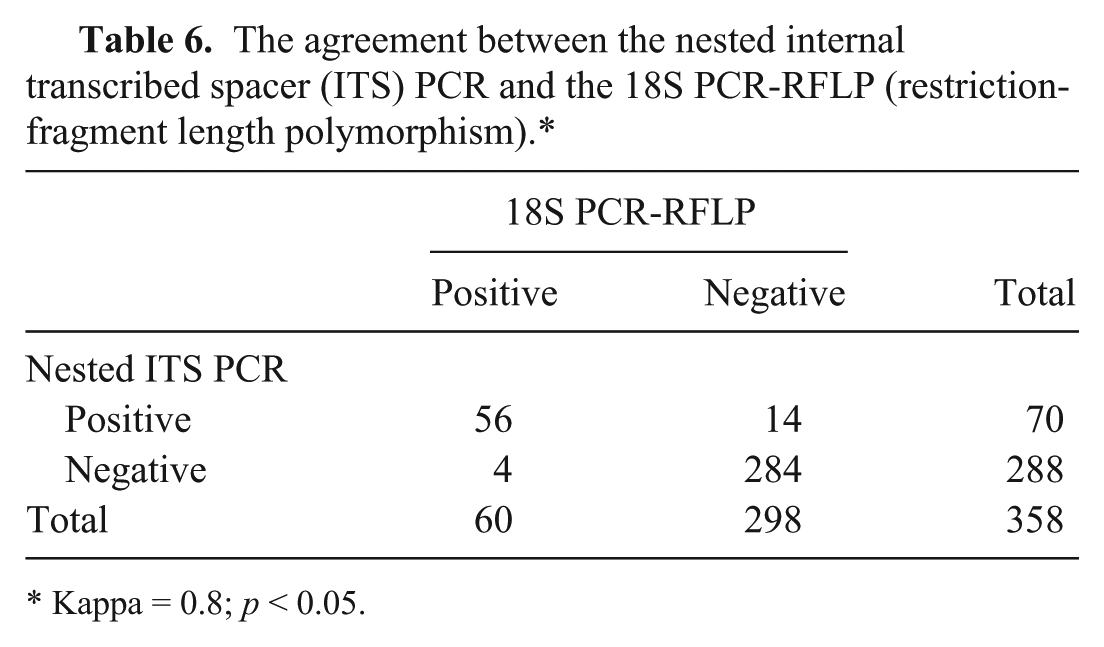

Kappa = 0.8;
The nested ITS PCR detected 83 parasite events, whereas the 18S PCR-RFLP detected 64, but the difference was not significant (Table 7; χ2 = 2.5, df = 1,
Events of the different species (subspecies) of trypanosomes detected by the nested internal transcribed spacer (ITS) PCR and the 18S PCR-RFLP (restriction-fragment length polymorphism).*
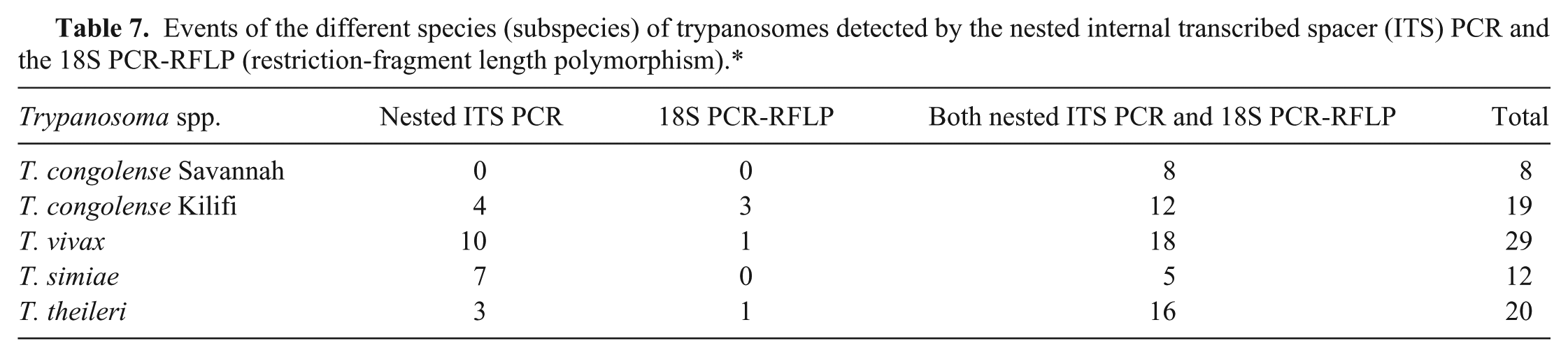

In both assays, infections encountered were caused by
Of these 3 pathogenic trypanosomes (
We recommend the use of the nested ITS PCR for detection of trypanosomes as it proved to be a sensitive technique for detection of trypanosomes occurring in both single and mixed infections. Practically, the inability of the assay to classify certain species (subspecies) is of less importance if the aim of detection is therapeutic intervention because these trypanosomes are treated with the same drugs. The methodological disadvantage of the 18S PCR-RFLP is the additional step that involves digestion of amplified product, which makes the assay time consuming and costly, and affects its sensitivity in cases where the concentration of amplified product is low. Nonetheless, unique restriction patterns provided by the 18S PCR-RFLP allow easy characterization of trypanosomes because only a few species have conflicting profiles by this assay.
Footnotes
Acknowledgements
We appreciate the valuable contributions from Dr. Hamisi Mwalonya, Kwale County Veterinary Officer; Dr. George Adinoh Omondi, Dawn Maranga, and Thomas Adinoh who are all research assistants at the Institute of Primate Research (Nairobi); and Nancy Nyambura Ndumia who was an intern in the laboratory of Prof. Dr. ir. Stefan Magez. Finally, we acknowledge Dr. ir. Stijlemans Benoît for critically reviewing the entire manuscript.
Authors’ note
S Odongo is a research fellow at Vrije Universiteit Brussel.
Authors’ contributions
S Odongo contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; and drafted the manuscript. V Delespaux contributed to acquisition, analysis, and interpretation of data. M Ngotho and S Magez contributed to conception and design of the study, and contributed to acquisition, analysis, and interpretation of data. SM Bekkele contributed to acquisition, analysis, and interpretation of data, and drafted the manuscript. All authors critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Vacutainer tube EDTA (K2) 6 mL, BD, Franklin Lakes, NJ.
b.
DNeasy blood and tissue (50), Qiagen GmbH, Hilden, Germany.
c.
NanoDrop, Thermo Fisher Scientific, Wilmington, DE.
d.
GoTaq buffer, Promega Corp., Madison, WI.
e.
GoTaq DNA polymerase, Promega Corp., Madison, WI.
f.
SeaKem LE agarose, Lonza Inc., Rockland, ME.
g.
h.
i.
CutSmart buffer, New England Biolabs Inc., Leiden, The Netherlands.
j.
SAM buffer, Thermo Fisher Scientific Inc., Erembodegem–Aalst, Belgium.
k.
FastAP, Thermo Fisher Scientific Inc., Erembodegem–Aalst, Belgium.
l.
Exonuclease I, Thermo Fisher Scientific, Erembodegem–Aalst, Belgium.
m.
MWG BIOTECH Primus 96plus 220 VAC, Cole-Parmer, Belgium.
n.
Clone Manager Professional 9, Sci-Ed Software, Morrisville, NC.
o.
SPSS16 statistical program, Polar Engineering and Consulting, Nikiski, AK.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the EU/FP7 HEALTH 2007-2.3.4-1 NANOTRYP project.
