Abstract
Introduction
Coronaviruses (family
A novel virus in the
Materials and methods
Challenge inoculum preparation
A pilot study was conducted to generate sufficient infectious inoculum for the primary experiment. Preliminary virus inoculums were prepared from 2 field specimens positive for PDCoV by qRT-PCR assay (South Dakota State University Diagnostic Laboratory, Brookings, SD): inoculum A from a dam with a history of diarrhea (PDCoV quantification cycle [Cq] = 18.66) and inoculum B from a neonatal pig (PDCoV Cq = 19.59). The pilot study was conducted using 3 groups of three 1-day-old piglets, with 1 group receiving inoculum A, 1 group receiving inoculum B, and 1 serving as the source-matched negative control. The PDCoV preliminary inoculum material tested negative for other common enteric viruses (TGEV; PEDV;
Piglets were sourced from a closed commercial farrowing facility that was negative for PEDV by serology and PCR testing, and negative for other enteric viruses by PCR. After 1 day of acclimation, piglets were gavaged with 5.0 mL of clarified inoculum. Twenty-four hours after inoculation, the animals receiving inoculum A were vomiting, passing large amounts of liquid feces, and were significantly dehydrated. All of the piglets that received inoculum A were euthanized, and tissues and intestinal contents were collected for qRT-PCR. The intestinal contents were stored at 4ºC for use in the primary experiment. The piglets administered inoculum B did not generate a productive infection, as determined with negative fecal qRT-PCR results and a lack of clinical disease.
The primary experiment inoculum was prepared by pooling intestinal contents with low PDCoV qRT-PCR Cq values from those inoculated animals in the preliminary study exhibiting clinical disease. Each animal sample tested qRT-PCR negative for TGEV, PEDV, toroviruses, and
Metagenomic sequencing was performed on the primary challenge inoculum and a clinical sample from an experimental pig post-inoculation to determine both the PDCoV genome sequence and identify any extraneous viruses present in the sample. The library was prepared using a method previously described.
18
Approximately 2.5 × 106 reads (challenge material) and 3.2 × 106 reads (clinical sample) were generated on a commercial instrument
a
using paired-end 150-bp reads. Reads mapping to host DNA were subtracted, and the remaining sequences were assembled de novo using bioinformatics software
b
into 135 contigs and classified based on BLASTN (http://www.ncbi.nlm.nih.gov/blast/Blast.cgi) expectation (
Animal infection
Nine near-term gestating dams were obtained from the same source as the pilot study, and were housed in the same facility and maintained in separate standard farrowing stalls. The dams were farrowed in a window of 2 days following a standard induction protocol. The dams and piglets of their subsequent litters were randomly divided into 4 groups: group A, PDCoV inoculated (47 piglets, 6 dams); group B, contact control (8 piglets originating from 4 group A dams); group C, aerosol control (20 piglets, 2 dams); and group D, negative control (12 piglets, 1 dam). All litters were kept intact, and there was no cross-fostering. The pigs and dams in group A were inoculated with the developed challenge inoculum at the start of the study (day 0), when piglets were ~2–4 days of age. The pigs in group B (~2 per litter) served as contact controls that were not inoculated and were littermates of the pigs of the inoculated group A animals. The group C aerosol control pigs were not inoculated, were housed in a separate pen in the same common animal room as a group A dam, and were separated by a distance of 0.6 m and a floor-to-ceiling length solid plastic curtain. Air movement was possible around the periphery of the curtain, but particles could not directly pass between farrowing stalls of the dams. Manipulations were performed on aerosol control (group C) animals first by individuals wearing clean disposable coveralls, gloves, masks, and head covers, and disinfected boots that were left in their respective rooms. The curtain and floor space between farrowing stalls was disinfected after each manipulation. Source-matched negative control animals were designated as group D, and were housed in a separate designated room. Animal-care staff was assigned to exclusively collect samples in each room to minimize the possibility of cross-contamination.
Clinical evaluation
The animals were observed daily for the following clinical signs: diarrhea, dehydration, wasting, coughing, sneezing, lethargy, ocular discharge, and any other unexpected clinical manifestation of disease. Fecal scoring was recorded using a 3-tiered system: 1 = normal feces, 2 = soft but formed feces, and 3 = diarrhea.
Autopsy and sample collection
A small subset of pigs from group A was sequentially euthanized on predetermined days post-inoculation (dpi), and a thorough postmortem examination was conducted. Pigs from groups B, C, and D were more variably sampled during select study days (Fig. 1). Severely diseased pigs were selectively chosen for euthanasia and autopsy. Criteria for euthanasia included severe dehydration, recumbency, weakness, and severe lethargy. A complete set of tissues, including tonsil, nasal turbinate, esophagus, stomach, trachea, cranial lung lobe, middle lung lobe, caudal lung lobe, submandibular lymph node, duodenum, proximal jejunum, distal jejunum, ileum, cecum, spiral colon, descending colon, mesenteric lymph node, thymus, tracheobronchial lymph node, spleen, liver, kidney, and inguinal lymph node, was collected from each pig, including fresh-frozen and formalin-fixed specimens. Serum and whole blood was also collected from each autopsied pig. Nasal and fecal swabs were collected from all available piglets and dams at each sampling time point postinoculation, and whole blood and serum were collected more variably (Fig. 1). Oral fluids were collected from each litter at 14, 21, 28, 35, and 39 dpi. Oral fluid collection ropes from a commercial kit c were applied to each litter’s farrowing stall and collected and processed according to the manufacturer’s directions. Blood and serum were collected from dams at 8, 14, 21, 28, and 35 dpi, and all dams were euthanized and examined by autopsy at 35 dpi. Mortality data were analyzed using Kaplan–Meier survival curve analysis generated with commercially available software. d Pigs noted as moribund and euthanized or dead were considered as deaths in analysis. All animal experiments were approved and performed in accordance with the guidelines of the Institutional Animal Care and Use Committee (IACUC protocol no. 1012).
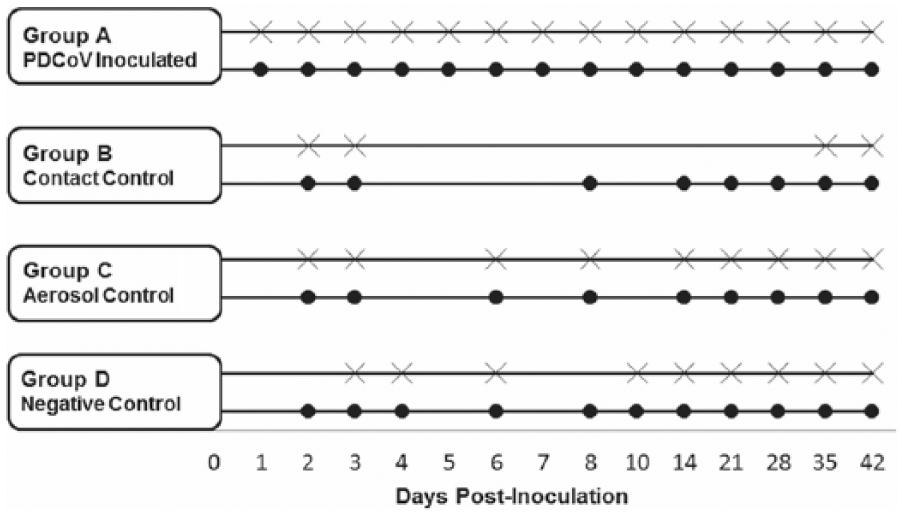
Experimental design indicating piglet blood sample collection and sequential euthanasia. Within each experimental group, circles denote collection of blood samples, and the crosses indicate autopsy was performed on a representative sampling of the group.
Polymerase chain reaction
Serum, fecal swabs, nasal swabs, fresh-frozen tissues, and oral fluids collected from the experiment were subjected to qRT-PCR analysis. For virus RNA extraction, 50 µL of each sample were loaded into a deep-well plate and extracted using a magnetic particle processor e and a viral RNA isolation kit f according to the manufacturer’s instructions, with a single modification, reducing the final elution volume to 60 µL. At least 1 negative extraction control consisting of all reagents, except sample, was included in each extraction. The extracted RNA was frozen at −20°C until assayed by qRT-PCR.
A duplex qRT-PCR was designed for the dual purpose of detecting PDCoV in samples by targeting the virus polymerase (RdRp) gene and monitoring extraction efficiency by targeting the 18S ribosomal RNA subunit gene as an internal control. Primers and probes for PDCoV (PDCoV-F: 5′-GTGCATGCATCTTTGTGGAT-3′, PDCoV-R: 5′-TAGGGTCAACCTTGGTGAGG-3′, PDCoV probe: 5′-FAM-TTCAGAACCTTGAACGTTACATCTCA-BHQ1-3′) and 18S (18S-F: 5′-GGAGTATGGTTGCAAAGCTGA-3′, 18S-R: 5′-GGTGAGGTTTCCCGTGTTG-3′, 18S probe: 5′-Cy5-AAGGAATTGACGGAAGGGCA-BHQ2-3′) were used in conjunction with the PCR kit g in a 20-µL reaction volume. The qRT-PCR reaction mix consisted of 3.5 µL of nuclease-free water, 10 µL of 2× reaction buffer, 1.0 µL of PDCoV forward and reverse primers (10 µM each), 1.0 µL of 18S forward and reverse primers (10 µM each), 0.5 µL of PDCoV FAM-labeled probe (10 µM), 1.0 µL of 18S Cy5-labeled probe (10 µM), 1 µL of enzyme mix, g and 2.0 µL of extracted RNA. Each qRT-PCR plate was run on a real-time PCR detection system g under the following conditions: 48°C for 10 min; 95°C for 10 min; followed by 45 cycles of 95°C for 20 s, 58°C for 20 s, and 72°C for 30 s. A synthetic PDCoV positive amplification control, and negative extraction and negative amplification controls were included in each run. The real-time Cq bar was set to the middle of the linear range of the amplification curves under log view. If a sample crossed the Cq before 38 cycles, the sample was considered positive. Based on quantification data estimating RNA copy numbers, this Cq value represents ~1–10 copies of virus RNA/reaction.
PDCoV antibody
An anti-PDCoV antibody was generated for use in IHC and other serologic assays. Briefly, the antibody was developed by cloning the full-length nucleoprotein (NP) of PDCoV and expressing it in
Histopathology and immunohistochemistry
For all pigs, representative formalin-fixed samples from the complete set of tissues collected at autopsy were routinely processed and embedded in paraffin blocks within 10 days after collection. Tissues were sectioned at 4 µm, stained with hematoxylin and eosin, and examined with light microscopy at the Veterinary Diagnostic Center, University of Nebraska–Lincoln.
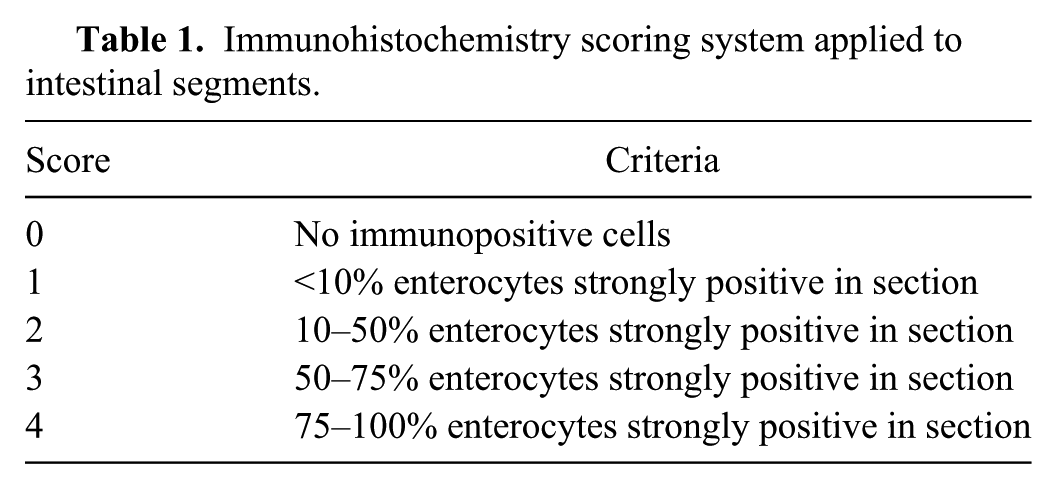
The formalin-fixed, paraffin-embedded tissues examined histologically were also stained for IHC. One section was evaluated for each tissue. There were some nonuniform sample sizes based on the pig age and available tissue from collection. The sections were cut at 4 µm and applied to slides, which were deparaffinized and stained using an automated immunohistochemical stainer. h The primary antibody consisted of the anti-PDCoV polyclonal rabbit serum previously developed. Positive and negative controls for PDCoV staining consisted of a slide containing known positive tissue collected in the pilot study along with slides of test samples using an irrelevant antibody (normal rabbit polyclonal immunoglobulin h ). The positive control tissue was examined with standard laboratory methods by IHC for PEDV and TGEV prior to use, and was immunonegative for either agent. After deparaffinization, the slides were incubated with a cell-conditioning solution h for 36 min. Before application of the primary antibody (optimally diluted at 1:10,000), a blocking step using normal goat serum i diluted at 1:10 was applied for 15 min. Primary antibody incubation was for 40 min at 36°C. Secondary antibody incubation and staining were conducted with commercial reagents using the manufacturer’s recommended protocols. h Tissues were counterstained with hematoxylin for 4 min and coverslipped with glass coverslips. The slides were examined with light microscopy for positive immunoreactivity. Intestinal enterocyte immunoreactivity was semiquantitatively scored based on the estimated percent of villus enterocytes with positive intracellular staining across the section examined, using a scale of 0–4 (Table 1).
Immunohistochemistry scoring system applied to intestinal segments.

Results
Clinical data
Group A and B pigs developed soft to diarrheic feces by 2 dpi, with many affected pigs having notable liquid fecal staining of the perineum. From 2–4 dpi, widespread severe diarrhea was evident in these pigs, with average fecal scores of 2.58–2.83. Groups A and B transitioned to soft, formed stools and then to normal stools by 12 dpi. Emesis was observed in 4 pigs from group A on day 2, but no emesis was observed in the other experimental groups or on subsequent days. Group C pigs developed diarrhea with perineal fecal staining on days 3 and 4. The litter that developed diarrhea at day 3 was exposed by direct contact with the inoculated dam of the room that escaped her farrowing stall overnight during day 1 post-inoculation. Corresponding to the onset of clinical disease, several individuals in the exposed litters (groups A–C) developed moderate to severe dehydration at 2 dpi. Lethargy was noted and peaked in groups A and B on day 2 in 8 of 55 pigs, and on day 3 in 8 of 46 pigs. Clinical status improved with the resolution of soft to diarrheic feces among the litters over the next 9 days. All surviving exposed pigs had normal feces and were otherwise clinically normal by 12 dpi.
Group A dams developed soft feces at 2 dpi, which progressed to diarrhea on day 3. Group C dams developed soft to diarrheic feces on day 5. During the first few days of diarrheal disease, dams were inappetent. Emesis was not observed in the dams, and no significant dehydration or lethargy was detected in dams during the study. All dams were clinically normal by 9 dpi. The negative control dams and piglets remained clinically normal throughout the study.
Morbidity was 100% among the exposed pigs. Mortality was variable among the litters, but particularly impacted 2 of the 6 inoculated litters (including group A and B pigs), as evidenced by the lowest documented litter survival at 63.5% and 50%. Two other inoculated litters had higher survival percentages of 88.9%, with 3–4 pigs surviving to the end of the study. Survival of the negative control pigs was 90.9% to the end of the study; overall survival of PDCoV-exposed pigs (groups A–C) was 72.9%. Group A and B pig survival was roughly similar at 69.6% and 62.5%, respectively. Group C pigs had a survival rate of 78%. Mortality in all exposed groups was distributed over the clinical phase of infection, as demonstrated in Figure 2, with the highest mortality occurring on day 3. Survival was also analyzed by the age of pigs at inoculation (groups A and B). Pigs ~2 days old at inoculation had a survival rate of 49.9%, and pigs that were 3 days or older at inoculation had a survival rate of 78.5%. All dams survived to their planned euthanasia and autopsy near the termination of the study.
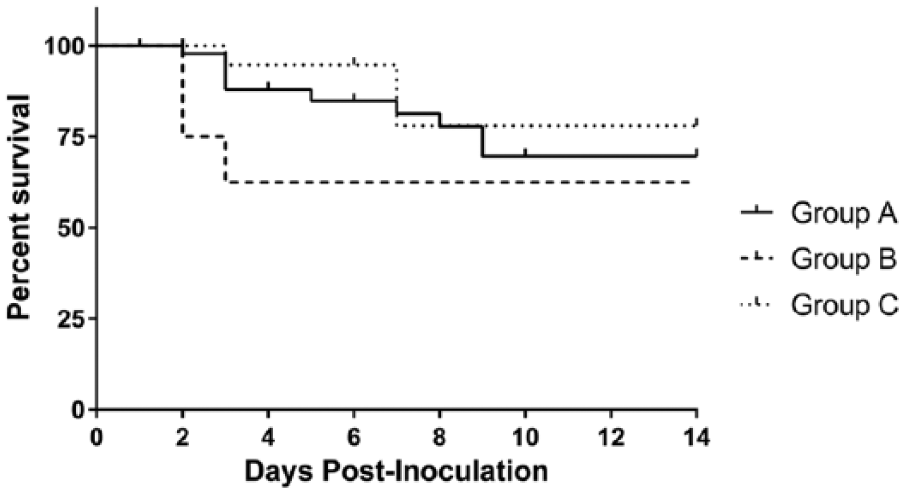
Survivability of pigs in each
Polymerase chain reaction
Fecal swabs from all group A and B pigs were qRT-PCR positive by 2 dpi. In group C, half of the pigs were fecal qRT-PCR positive by 2 dpi (1 litter of 2), and all were positive by day 3. The duration of fecal virus shedding as detected by qRT-PCR varied among individual pigs. Fecal swabs from all pigs of groups A and B had undetectable virus RNA by 28 dpi and after. One group C pig remained weakly positive (Cq 35.2, weak positive 35–38) at 28 dpi, but was negative by the next testing period at day 35 (Fig. 3A). The lowest mean fecal Cq value of positive inoculated pigs, indicative of peak fecal virus shedding, occurred at 2 dpi (Fig. 3B). For nasal shedding, all inoculated and contact control pigs were qRT-PCR positive by nasal swab on day 2, and all samples had undetectable virus on day 28 and thereafter. Generally, fecal swab qRT-PCR results had lower Cq values than nasal swabs (Fig. 3C). The aerosol control pigs were positive on nasal and fecal swabs qRT-PCR on the same day, except for 7 pigs from a single litter, which were nasal swab qRT-PCR positive at day 2 and fecal qRT-PCR positive at day 3. Virus RNA was undetectable in aerosol control pigs by 35 dpi. Throughout the study, the negative control pigs had undetectable PDCoV RNA in fecal and nasal swabs by qRT-PCR.
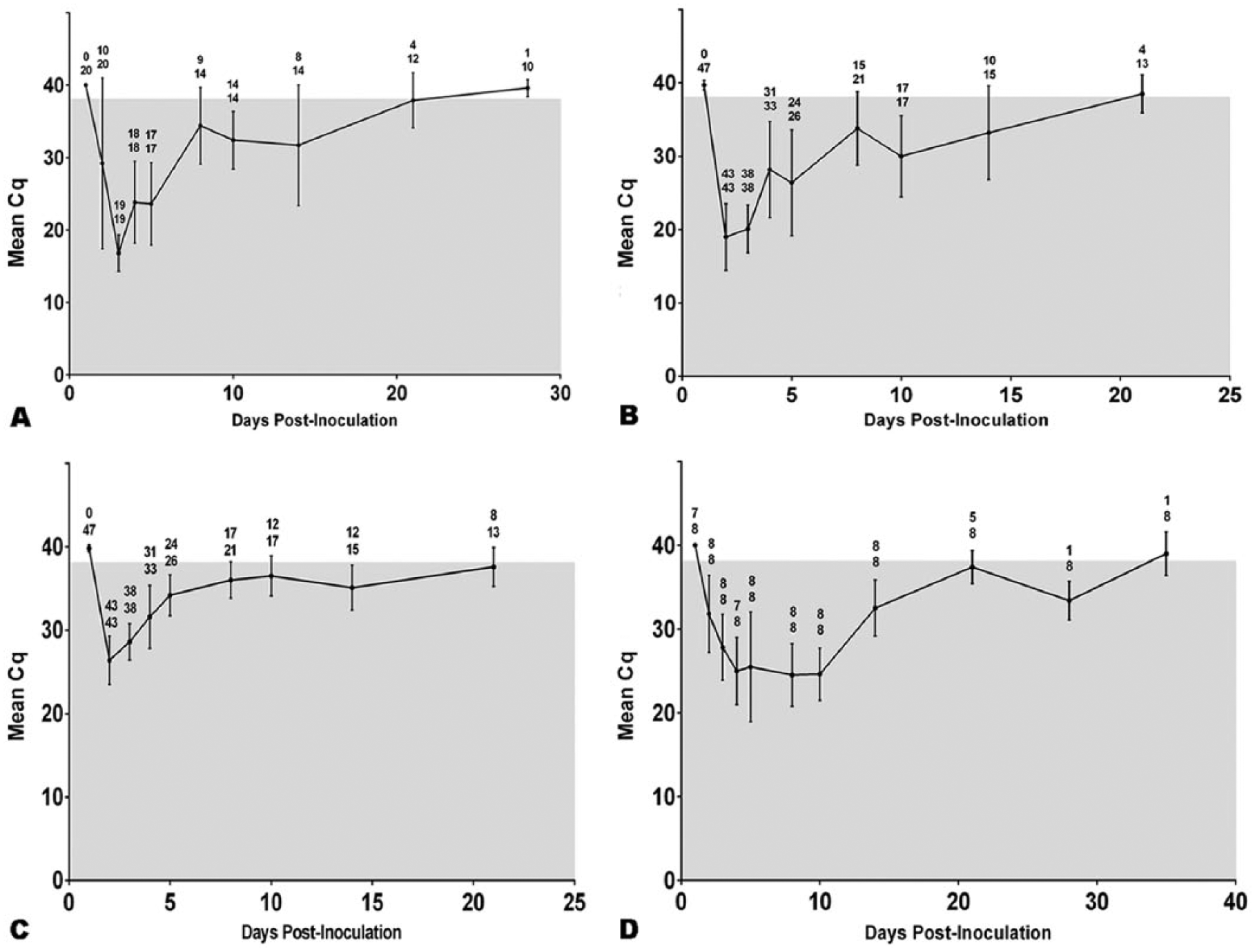
Mean quantification cycle (Cq) values and standard deviation of quantitative real-time reverse transcription PCR (qRT)-PCR for experimental groups are presented. Values above the linear data points indicate the number of individual animals that were qRT-PCR positive over the total animals tested at that time sampling point on the X-axis.
The majority of exposed dams (groups A and C) became qRT-PCR positive on fecal swabs synchronously with her litter, except dam 4 from group A was fecal negative on day 2 when her piglets were all strongly fecal positive, and dam 8 of group C became weakly positive (Cq 37) at day 2 prior to her litter becoming positive on day 3. Typically, the dams had notably higher Cq values on fecal qRT-PCR than the majority of their piglets the first day or two of infection, but the Cq values appeared similar to the piglets thereafter (Fig. 3D). The range of days that dams were detected shedding was 12-34 days, with an average of 21 days. One inoculated dam demonstrated the longest duration of PCR positive feces, from 2 to 35 dpi, at which time the dams were removed from the study and euthanized.
Blood and/or serum samples were subjected to qRT-PCR analysis. Day 1 post-inoculation, viremia was detected in 50% of the sampled group A pigs (Cq 31.1–34.4). Viremia was detectable in most group A pigs from days 2–5 post-inoculation (Cq 27.4–37.7), where the prevalence of viremia is demonstrated graphically (Fig. 4). At day 6, all group A pigs sampled had undetectable levels of virus. All sampled group A pigs (
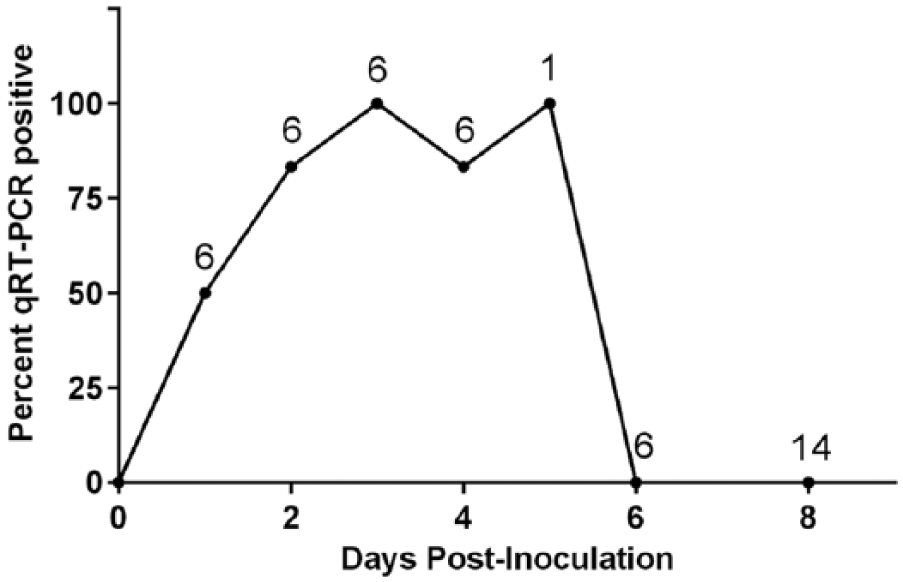
Percent of positive serum and/or whole blood samples from individual inoculated pigs days 1 – 8 post-inoculation. Values above the linear data points indicate the number of animals sampled, and the Y-axis position indicates the percent of those samples positive.
In group B, 100% (2/2) sampled were qRT-PCR positive on serum on day 2 (Cq 28.1 and 33.7). In group C, a pig sampled on day 2 was qRT-PCR positive on serum (Cq 33), and 1 pig on day 3 had undetectable levels of virus. The group B and C pigs had undetectable virus RNA on day 8 and thereafter for the remainder of the study. All negative control pigs had undetectable levels of virus in serum and/or whole blood throughout the study.
Nearly all tissues collected during the first 2 days of the study were variably positive. On day 1 (only group A pigs selected), the small intestine was consistently strongly positive (Cq ≤ 22); the spiral colon, cecum, and lymph node samples were strongly to moderately positive (22 < Cq < 35; Supplemental Fig. 1, available online at http://vdi.sagepub.com/content/by/supplemental-data). Samples from the descending colon on day 1 were moderately to weakly positive (Cq ≥ 35). On day 2, the large and small intestine and lymph node were strongly to moderately positive to 6 dpi. Following day 6, lymph node and small intestinal samples were mostly moderately positive out to the end of the study at day 42. Some random individual samples of descending colon, spiral colon, and cecum from group C pigs only were weakly to moderately positive from day 28 to termination (Supplemental Fig. 3, available online at http://vdi.sagepub.com/content/by/supplemental-data). The negative control pigs euthanized and tissues sampled had no viral RNA detected throughout the study.
A corresponding set of tissue samples collected from dams at 35 dpi was subjected to qRT-PCR analysis. All exposed dams were qRT-PCR positive in lymph node (Cq 27.3–34.6, Supplemental Fig. 4, available online at http://vdi.sagepub.com/content/by/supplemental-data). Dam 1 (group A) was positive in the descending and spiral colon. Three inoculated dams were detected as qRT-PCR positive in the small intestine (Cq 35.4–37.9). Both group C dams were qRT-PCR positive in the small intestine (Cq 31.2 and 32.7). Spleen samples from one group A and one group C sow were weakly positive. All other tissue samples from the dams were qRT-PCR negative, including all tissues from the negative control dam (S9, Supplemental Fig. 4).
Oral fluids were subjected to qRT-PCR analysis. Two of the group A litters were not sampled, as there were no remaining piglets in either litter at day 14. All exposed litters sampled (groups A–C) were qRT-PCR positive on oral fluids taken on days 14 and 21 (Cq 28.1–36.9). On day 28, 4 of 6 litters were PCR positive on oral fluids. Two of these were group C litters (Cq 34.5 and 36.6), and the other 2 positive litters were comprised of group A and B pigs (Cq 30.3 and 37.0). Oral fluids from all litters had undetectable RNA by qRT-PCR on 35 dpi, and 1 group C litter was weakly positive 39 dpi with a Cq of 37.9.
Histopathology
By 2 dpi, all group A pigs exhibited lesions of mild to severe atrophic enteritis within the jejunum and ileum. These lesions included villus enterocyte swelling with intracytoplasmic vacuolation, enterocyte attenuation and loss, mild to severe villus blunting and fusion, and submucosal edema with an occasional infiltrate of neutrophils (Fig. 5). Not all samples of small intestine from each exposed pig were affected, and the severity of lesions often varied between the small intestinal segments examined in individual pigs. On days 3–6, all group A pigs examined had lesions of mild to moderate atrophic enteritis in the jejunum and ileum, although 6 of 49 samples were too autolyzed to determine the full extent of the lesions. Of the 3 group A pigs examined on day 7, all were too autolyzed to examine in detail. Two of the 4 group A pigs examined on day 8 had lesions of mild to moderate atrophic enteritis in the jejunum and ileum. No significant enteric lesions were observed in group A pigs at 10 dpi or thereafter.
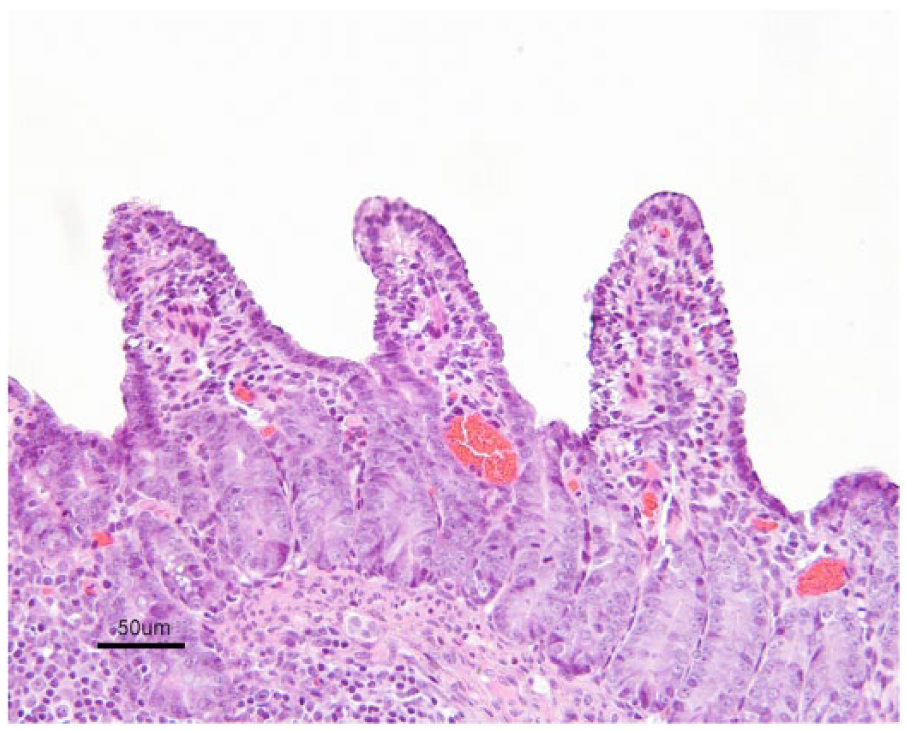
Section of ileum of
Pigs from groups B and C were examined on select days. The 2 group B pigs examined at day 2 had lesions of atrophic enteritis in the jejunum, and one of these pigs had detectable lesions within the ileum. No lesions were observed in the group C pig examined at 2 dpi. The group B pig examined at day 3 had mild lesions of atrophic enteritis in the jejunum and ileum. One group C pig examined on each of day 3 and 6 displayed atrophic enteritis in the jejunum and ileum. Three group C pigs were examined on day 8, but autolysis obscured identification of any lesions of atrophic enteritis. No enteric lesions were observed in group B pigs examined at 28 and 42 dpi, and no enteric lesions were observed in group C pigs examined at 14 dpi and each sampling day thereafter. No lesions of atrophic enteritis were observed in the negative control pigs throughout the study.
Enteric lesions other than characteristic atrophic enteritis were infrequently noted. In the duodenum from 3 to 8 dpi, 8 pigs from group A and 1 from group C had scattered foci of mild to moderate, and rarely severe, necrosuppurative lesions in the tips of the villi with occasional fibrin thrombi within small vessels of the villi, and local enterocyte attenuation. Four of these duodenal samples had minimal IHC staining (score 1), and the others were IHC negative.
No significant enteric lesions were observed by microscopic examination in the dams. One dam had a grossly and histologically evident chronic infarct affecting 75% of the spleen.
Immunohistochemistry
At 1 dpi, all 3 euthanized group A pigs displayed strong positive intracytoplasmic staining in villus enterocytes of the jejunum and ileum (Table 2, Fig. 6). All group A pigs sampled continued to be IHC positive in villus enterocytes of the jejunum and ileum on days 2–6. Occasional scattered interdigitating cells within the Peyer patches were also positive after day 2. The percentage of group A individuals with positive staining in the jejunum and ileum quickly declined through days 7 and 8, and no positive staining was observed in samples from 10 dpi and thereafter. No mesenteric lymph node samples from group A pigs were positive at day 1 or 7. On days 2–5 and 8, the majority (71–100%) of the mesenteric lymph nodes sampled were positive, characterized by scattered interdigitating cells with strong intracytoplasmic staining (Fig. 7). No mesenteric lymph nodes were positive on day 10 or thereafter. Other immunopositive tissues included duodenum, cecum, and spiral colon. A lower percentage of pigs had positive staining in these other intestinal segments (Table 2), and often the staining affected a low number of scattered enterocytes. Positive staining in the duodenum was observed at 2–6 dpi. The spiral colon only stained positive on days 2 and 3, and cecum samples were variably positive on days 2–4 and 6. Pigs from groups B and C were also euthanized and sampled at their selected time points (Fig. 1). Similar to group A, the duodenum, jejunum, ileum, and mesenteric lymph node samples were consistently positive at 2–8 dpi (Table 2). No immunopositive tissues were observed after day 8 in group B and C pigs. No positive immunoreactivity was observed in tissues other than the intestinal tract and mesenteric lymph node. The euthanized negative control pigs remained negative by IHC analysis throughout the study. A mean IHC score was calculated from fixed tissue samples collected from the groups at each dpi (Table 2). The highest mean IHC scores in group A pigs were observed at 1–6 dpi in the jejunum and ileum. A low mean IHC score (≤0.5) was observed in the jejunum and ileum of group A pigs on day 8. A low mean IHC score was observed in the duodenum, spiral colon, and cecum at 2–6 dpi. A similar trend was observed for the data set obtained for groups B and C, although the sample of pigs from these groups was notably smaller. PDCoV was not detected by IHC in any tissue sample from the dams at day 35.
Percent immunohistochemistry (IHC)-positive pigs and mean IHC score (out of 4).*
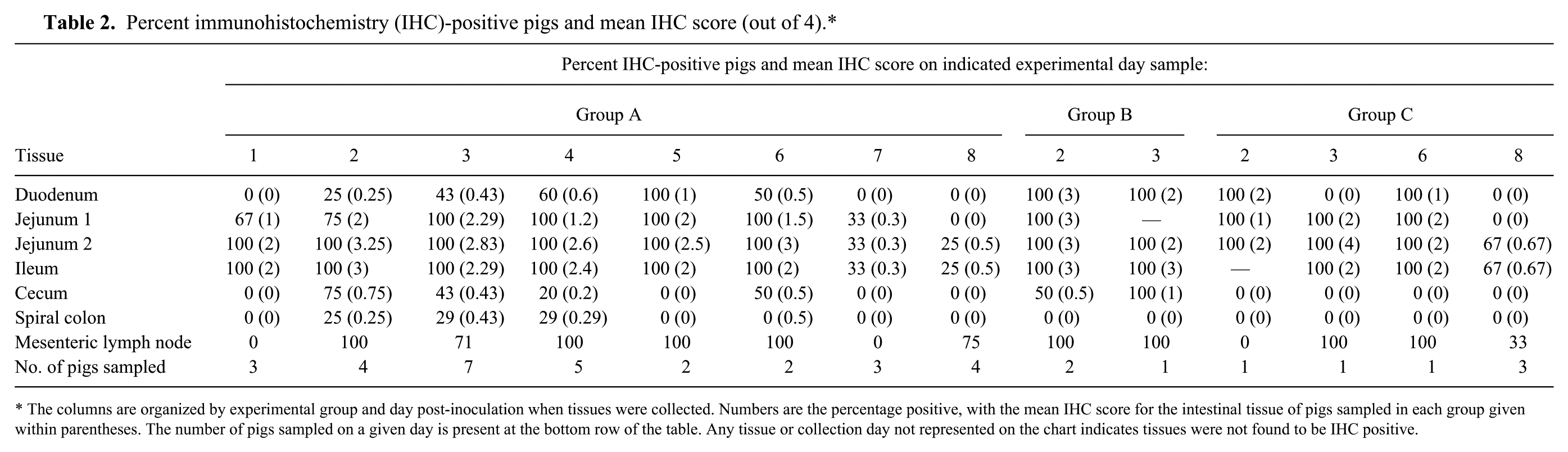
The columns are organized by experimental group and day post-inoculation when tissues were collected. Numbers are the percentage positive, with the mean IHC score for the intestinal tissue of pigs sampled in each group given within parentheses. The number of pigs sampled on a given day is present at the bottom row of the table. Any tissue or collection day not represented on the chart indicates tissues were not found to be IHC positive.
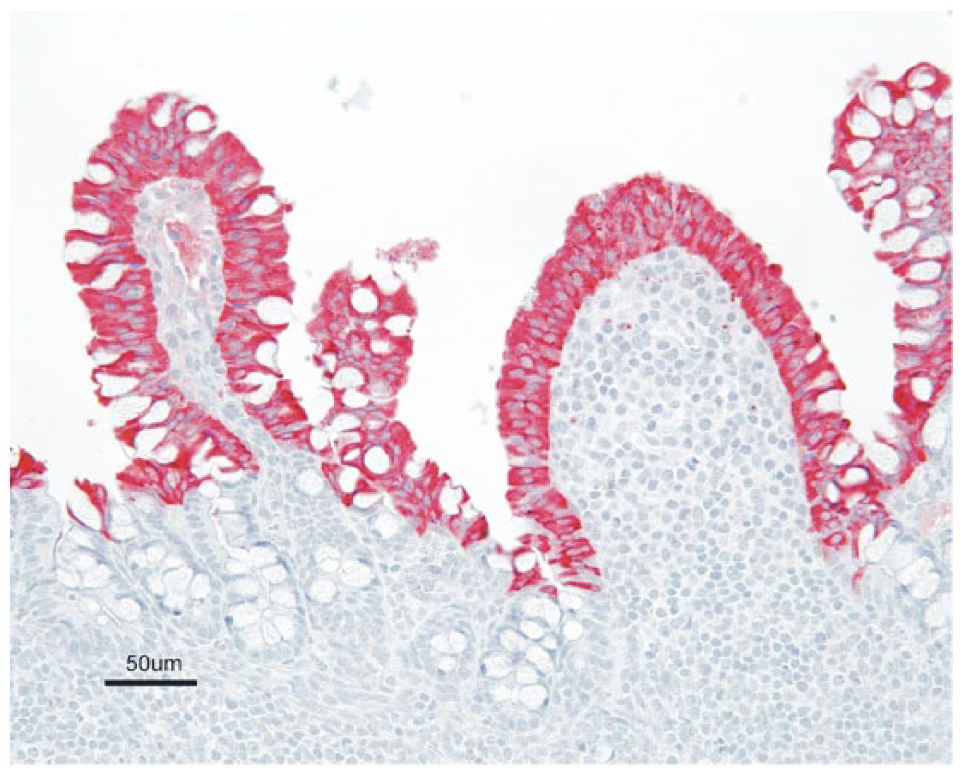
Section of ileum of
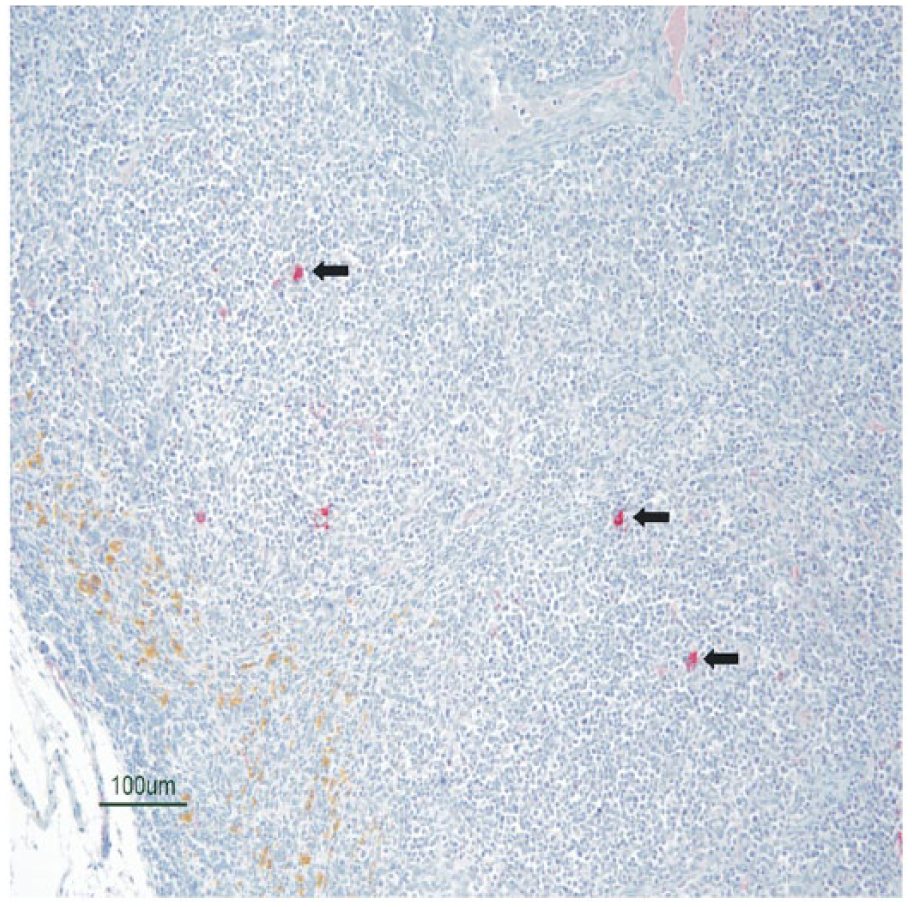
Section of mesenteric lymph node of
Discussion
Experimental inoculation demonstrated that PDCoV infection in the absence of other enteric pathogens is capable of producing clinical diarrhea, emesis, and dehydration, as well as appreciable mortality in conventional neonatal pigs. Clinical signs were much less severe in infected dams compared to piglets. Disease in the dams was characterized by transient soft to diarrheic feces and inappetence in the absence of clinically significant dehydration or mortality. The use of this challenge model of conventional piglets and dams in lieu of gnotobiotic or conventional weaned pigs may provide altered host–pathogen interaction and different observations about lesions. Work with human noroviruses has shown that the microbiota of the gut can have a critical effect on virus pathogenesis. 7 The lack of a cell culture–adapted PDCoV at the time of trial necessitated the generation of an infectious inoculum from a field sample, which was demonstrated to be free of other etiologic agents of porcine enteric disease by all available modalities, including bacterial culture, PCR, and metagenomic sequencing. Importantly, no significant extraneous or confounding viruses were detected in the challenge inoculum, although partial genome coverage of porcine kobuvirus was demonstrated in a fecal swab from a PDCoV-infected animal. As porcine kobuvirus has been isolated from healthy pigs and those with diarrhea, the virus has no distinct role as a gastrointestinal pathogen of pigs and is considered an insignificant finding in our study. 23 The use of a sizable experimental sampling of conventional pigs and a non–cell culture–adapted virus inoculum in our study closely mimics natural field conditions.
The PDCoV clinical disease course of diarrhea, emesis, and inappetence observed in piglets and dams in this study is similar to TGEV and PEDV. 20 The incubation period prior to the onset of clinical diarrhea in our study also resembles experimental and natural infections with TGEV and PEDV in neonatal pigs, where the onset of clinical signs can vary from 24 to 48 h and may depend on pig and virus strain characteristics.3,5,9 Other experimental inoculations of PDCoV in 19-day-old conventional piglets and in 5-day-old gnotobiotic piglets demonstrate that the onset of diarrhea can vary from 24 h to 2–4 dpi.2,8,11 Emesis, an inconsistent feature of both PEDV and TGEV infections, was documented to occur in a few PDCoV-inoculated pigs of our study only on day 2 postinoculation, and may be an early transient feature of PDCoV infection. The contact control pigs in this study developed diarrhea synchronously with the inoculated pigs, and the aerosol control pigs developed disease 1–2 d after, indicating rapid and efficient horizontal transfer of PDCoV in pigs. It is possible that virus inoculum from the group A pigs contaminated the contact control pigs, rather than spread from fecal virus, resulting in this rapid development of disease. One aerosol control litter had no direct contact with inoculated animals, whereas the other aerosol control litter did have incidental direct contact with an inoculated dam. These data indicate PDCoV infection would likely spread rapidly throughout modern farrowing and gestation barns, despite minor barriers such as farrowing stalls and physical barriers. PEDV has been demonstrated to have the potential for airborne transmission of infectious virus, and PDCoV may also have similar characteristics. 1
Despite many similarities between the demonstrated PDCoV infection and TGEV and PEDV, some differences may exist in the severity of clinical disease and mortality during infection of neonatal pigs. Mortality caused by epidemic TGEV and PEDV in neonatal pigs of naive herds is high, often approaching 90–100% in the youngest pigs and decreasing with age.5,22 The first field reports of PDCoV associated with diarrheal disease in pigs noted that the mortality in affected piglets was ~30–40%, much lower than typically observed with PEDV infection. 24 In our study, it was not possible to calculate a true mortality rate because of the rigorous serial sequential euthanasia of pigs and selection bias of severely diseased pigs, thus survival analysis was conducted using dead and moribund pig numbers. The survival of all PDCoV-exposed groups was less than the negative control pigs, with mortality distributed over the diarrhea phase, demonstrating an increase in mortality of clinically ill PDCoV-infected neonates. The single pig that died in the negative control group was runted with evidence of septicemia, and appears to represent unrelated neonatal mortality. A notable survival effect was demonstrated by comparing the different ages of pigs at inoculation. Pigs ~2 days of age at inoculation had lower survival rates than those 3 days or older at inoculation. This indicates that disease severity and mortality of infected pigs likely decreases with age, similar to TGEV and PEDV. This is important to note as recent experimental studies all used older conventional pigs (5–10 days of age) for challenge.2,11
Histologic examination revealed lesions characteristic of atrophic enteritis, which appeared to be confined to the jejunum and ileum for 7 experimental days during the clinical disease course. Overall, these lesions were clearly impactful but did not appear to demonstrate the severity seen in PEDV infections in naive neonatal piglets, similar to observations in other studies. 26 The onset of histologic lesions corresponded to the onset of diarrhea, and was associated with the lowest qRT-PCR Cq values of PDCoV in blood, tissues, and feces, and IHC detection of PDCoV in the small intestine. Other studies in 5-day-old conventional piglets demonstrated a more gradual onset of diarrhea and fecal virus shedding by PCR from 2 to 5 dpi. 2 Intestinal lesions resolved 2–3 days prior to the complete resolution of diarrheal disease at 12 dpi. Viral RNA was detected in serum and whole blood samples in animals of this study, in contrast to other studies which found none. 8 By IHC, the jejunum and ileum were the primary sites of PDCoV replication, as these were the first tissues to be positive and were the most consistently positive throughout the experiment, which is consistent with other experimental PDCoV infections. 2 Viral antigen was less frequently detected by IHC in the duodenum, cecum, and spiral colon, and was not associated with any significant lesions in these areas of the gastrointestinal tract. In contrast to these findings, other studies have not demonstrated PDCoV IHC staining in sections of cecum and colon. 2 The mild to moderate necrosuppurative lesions detected in some duodenal samples were inconsistent with viral atrophic enteritis, and no significant IHC detection of virus was associated with these lesions. These lesions appeared to be limited to duodenum, and were not significantly associated with mortality. Therefore, the duodenal lesions are considered to be likely a result of intestinal dyshomeostasis. The mesenteric lymph nodes were the only nonintestinal tissue found to be positive by IHC, and no specific lesions were associated with IHC staining. One PDCoV study, which used gnotobiotic pigs, demonstrated epithelial degeneration and necrosis and occasional syncytial cell formation in the stomach; however, no significant lesions or IHC positive staining of the stomach were observed in our study. 11 Overall, the lesions and virus distribution by IHC observed in our experiment resembles that observed in TGEV and PEDV.12,21 Additional studies are needed to further investigate potential mechanisms underlying the strong tropism of PDCoV for the jejunum and ileum, and lesser tropism for other intestinal segments.
Although the onset and resolution of lesions correlate with detection of virus by IHC, PDCoV was detected in tissues and feces by qRT-PCR for many days after diarrhea ended. In general, most PDCoV-exposed pigs and dams were qRT-PCR positive in the small intestine and lymph node out to termination. In comparison, PEDV has been demonstrated to persist in lymph node and small intestine such that animals are PCR-positive for at least 28 dpi (personal communication, R Hesse). The positive fecal swabs from PDCoV-inoculated pigs up to 21 dpi is comparable to similar studies of 3-week-old pigs experimentally challenged with PEDV, where fecal shedding of virus was demonstrated extending out to 24 dpi. 12 In the current study, 1 dam demonstrated fecal shedding out to 35 dpi, and 1 aerosol control pig demonstrated fecal shedding to 28 dpi. This suggests that fecal shedding of PDCoV could extend longer, particularly in adults, which is important as they may serve as reservoirs infecting naive dams and piglets in farrowing facilities. However, it should be noted that PCR detection of virus does not necessarily indicate the presence of viable virus or indicate that there is viral replication in the tissues. Detection of PDCoV RNA concurrently with negative IHC may be related to the larger sampling and greater sensitivity of the PCR method, or to persistence of detectable nucleic acids versus replicating virus in the samples. In a natural disease outbreak, IHC may be useful to determine if an affected pig with acute onset of diarrhea has an active PDCoV infection or to detect PDCoV when only fixed tissues are available for examination. Currently, a number of PEDV-positive farms also have detectable PDCoV in the herd, and IHC could be used to determine the contribution of PDCoV to active enteric disease. This is clearly important as, demonstrated in our study, fecal shedding and tissue PCR detection can be prolonged weeks beyond the clinical phase of disease. However, the PCR results obtained from a positive herd may be important in order to monitor the reservoir of infected animals shedding fecal-borne virus into the environment and to screen for recently infected animals.
Oronasal virus detection was successful during the study. Nasal swabs and oral fluids were qRT-PCR positive and correlated with fecal qRT-PCR results temporally. Average Cq values indicated that nasal swabs contained less viral RNA than fecal swabs. No virus was demonstrated within the nasal turbinate, trachea, or lung by IHC, and no respiratory tract lesions were observed. These observations indicate that experimental PDCoV infection in our study did not yield a detectable respiratory infection even though pigs were exposed to virus via an intranasal route. Previous studies also did not observe lesions or detect antigen by immunostaining reactions in respiratory tissue of PDCoV-challenged 11–14-day-old gnotobiotic piglets or 5-day-old conventional piglets.2,8 In contrast, related viruses PRCV and TGEV have been shown capable of replication in alveolar cells and epithelial cells of the respiratory tract from the nasal mucosa to bronchioles, as well as in the ileum. 4 PEDV has also been described to replicate in alveolar macrophages. 19 The current findings also contrast a PDCoV experimental inoculation of gnotobiotic pigs in which interstitial pneumonia was noted along with PDCoV antigen detection in bronchial mucosal epithelial cells using IHC. 11 However, the investigators utilized hyperimmune sow serum for the primary antibody, and this may result in nonspecific or background staining. In the current study, the positive PCR detection on nasal swabs is interpreted to be environmental contamination of external nares. However, nasal swab and oral fluid PCR may be complementary tests to monitor PDCoV infection and virus shedding in infected pigs on a herd level.
In our study, PDCoV has been demonstrated as a pathogenic agent causing atrophic enteritis and diarrhea in conventional neonatal piglets and dams, confirming other PDCoV experimental inoculations and providing a comprehensive pathogenesis model to enable comparisons to the other important enteric coronaviruses of swine, PEDV and TGEV. PDCoV should be considered in a differential diagnosis in diarrhea outbreaks in neonatal as well as adult pigs. The presented pathogenesis described in our study using conventional pigs serves as a model for the expected acute disease course in a typical naive swine production system infected with PDCoV. It is important to note that while disease was reproduced successfully in this study from a PDCoV field case, a second inoculum generated from a separate field case failed to cause disease in the primary piglet challenge. This discrepancy may highlight an important feature of PDCoV in that pathogenicity may depend on uncharacterized viral virulence properties or other host factors. This study also demonstrated rapid transmission to aerosol control animals, suggesting that infective virus can readily spread under confinement conditions and may have the capacity to spread in an airborne manner as demonstrated with PEDV. 1 The data generated in the present study regarding virus detection with IHC and qRT-PCR enables development of guidelines for PDCoV laboratory testing, particularly when investigating outbreaks of diarrhea in pigs and monitoring closed herds.
Footnotes
Acknowledgements
We thank Travis Clement for coordination of PCR testing and inoculum preparation, as well as Steve Lawson, Aaron Singrey, and Faten Okda for preparation of anti-PDCoV antisera. We also thank Dr. Kelly Heath, Vicky Samek, Jon Kolman, Sheryl Hemmer, Brandon Stewart, Kevin Brodersen, Taylor Engle, Tyler Harris, and the University of Nebraska–Lincoln Veterinary Diagnostic Center staff and animal care staff for all their efforts in helping to complete many tasks of sample collection and processing.
Authors’ contributions
S Vitosh-Sillman contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; and critically revised the manuscript. JD Loy contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; and critically revised the manuscript. B Brodersen, C Kelling, C Topliff, E Nelson, and R Hesse contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; and critically revised the manuscript. A Doster contributed to conception of the study and to acquisition of data. J Bai contributed to conception and design of the study; contributed to acquisition and analysis of data; and critically revised the manuscript. E Schirtzinger contributed to acquisition and analysis of data and critically revised the manuscript. E Poulsen, B Meadors, J Anderson, and B Hause contributed to acquisition and analysis of data. G Anderson contributed to acquisition of data. S Vitosh-Sillman, JD Loy, B Brodersen, C Kelling, C Topliff, E Nelson, and R Hesse gave final approval and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
MiSeq, Illumina Inc., San Diego, CA.
b.
CLC Genomics Workbench version 7.5, Qiagen Inc., Valencia, CA.
c.
TEGO Swine Oral Fluids kit, ITL Biomedical Animal Healthcare, Reston, VA.
d.
GraphPad Prism version 6.05 for Windows, GraphPad Software, La Jolla, CA.
e.
Kingfisher 96 magnetic particle processor and AgPath-ID one-Step RT-PCR kit, Fisher Scientific, Pittsburgh, PA.
f.
MagMAX-96 viral RNA Isolation kit, Life Technologies, Grand Island, NY.
g.
CFX96 Touch real-time PCR detection system, Bio-Rad Laboratories, Hercules, CA.
h.
Benchmark ULTRA IHC/ISH slide staining platform, CONFIRM negative rabbit Ig, Cell conditioner 1, and proprietary reagents; Ventana Medical Systems Inc., Tucson, AZ.
i.
Goat serum, Colorado Serum Co., Denver, CO.
Declaration of conflicting interests
The author (s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Pork Board (#14-182). This work was supported in part by funds through USDA Animal Health and Disease Research Program under the provisions of Section 1433 of Subtitle E, Title XIV of Public Law 95-113 and the Kansas State Veterinary Diagnostic Laboratory.
