Abstract
Bovine viral diarrhea virus 1 (BVDV-1) is associated with mild or subclinical infections, whereas BVDV-2 is frequently implicated in outbreaks of severe thrombocytopenia and acute fatal disease. In the present study, the carcass of a beef breed cow and tissue samples of a beef calf were received for laboratory diagnosis. Both animals exhibited severe clinical signs compatible with thrombocytopenia or hemorrhagic syndrome. Direct fluorescent antibody test (DFAT) failed to detect BVDV antigen in the tissue specimens of both cases. However, immunohistochemistry (IHC) revealed the presence of BVDV antigen in oral and esophageal mucosa and Peyer patches of the beef breed cow. Real-time reverse transcription polymerase chain reaction (real-time RT-PCR) detected BVDV-2 in selected tissues of both animals. Subsequently, BVDV was isolated from both cases and subjected to genetic and serologic characterizations. Mutations in the 5′–untranslated genomic region (5′-UTR) primer and probe binding sites and the E2 gene were associated with reduced efficiency of an established real-time RT-PCR assay and amino acid alterations in the E2 glycoprotein, respectively. Both viral isolates were classified by real-time RT-PCR and phylogenetic analysis as BVDV-2 subgenotype a. Unlike BVDV reference strains Singer and 125c, the isolates cross-reacted with anti–BVDV-1 and anti–BVDV-2 reference sera, indicating antigenic variations in field isolates. The isolates also showed reduced reactivity to porcine anti-BVDV antiserum (the raw serum used to produce BVDV DFA conjugate). In summary, data from the present investigation indicated that genetic and antigenic variations affected the performance of detection assays, especially DFAT, highlighting the need for regular evaluation and modification of BVDV tests.
Keywords
Introduction
Bovine viral diarrhea virus (BVDV), along with Classical swine fever virus (CSFV) and sheep Border disease virus (BDV), is classified in genus Pestivirus in the Flaviviridae family. 14 BVDV has a broad tissue tropism, and the infection is associated with a wide variety of clinical manifestations, such as respiratory distresses, diarrhea, reproductive disorder, encephalitis, and congenital abnormalities. 2 BVDV transmission can occur horizontally through the oronasal route or vertically from a pregnant cow to its fetus.15,21 Infection in the first trimester of gestation may lead to immune tolerance, and fetuses that survive are born as persistently infected (PI) calves. 2 PI animals do not have detectable antibodies against BVDV, but shed the virus continuously in their excretions and secretions, serving as a primary source of BVDV infection. Therefore, prompt identification and subsequent removal of PI animal(s) from exposed herds are critically important to a successful control and prevention program.2,3
The 5′–untranslated genomic region (5′-UTR) of BVDV is highly variable. Phylogenetic analysis of 5′-UTR sequences led to the recognition of BVDV genotypes 1 and 2.12,19 BVDV-1 is often associated with mild or subclinical infections, whereas BVDV-2 is frequently isolated from outbreaks of hemorrhagic syndrome in North America.4,20 The 2 genotypes are serologically distinguishable, and serotyping has been performed in BVDV diagnosis. 6 The E2 gene is another hypervariable region in the BVDV genome. 7 The E2 glycoprotein plays important roles in the induction of host immune response. Antigenic changes due to mutations in the E2 gene can significantly affect the sensitivity of diagnostic tests, such as immunohistochemistry (IHC) and enzyme-linked immunosorbent assay (ELISA).9,11 In the present study, we report that the direct florescence antibody test (DFAT), with proper positive and negative controls, failed to detect BVDV antigen in 2 cases of the severe hemorrhagic form of BVD. The potential causes of DFA detection failure were investigated via genetic and serologic approaches.
The first case involved an adult beef breed cow (M10-3432) that was found dead by the attending veterinarian and submitted to the Mississippi Veterinary Research and Diagnostic Laboratory (MVRDL; Mississippi State University, Pearl, MS) for autopsy. Prior to death, the animal developed acute respiratory signs and bloody diarrhea. The death was the seventeenth fatality in a herd of 40 animals within a 3-month period of time. Necropsy was performed, and samples were collected from major organs, including brain, ear notches, oral and esophageal mucosa, small and large intestine, mesenteric lymph node, lung, liver, spleen, and kidney. The second case (M10-5347) involved a feedlot calf from a different farm that exhibited antemortem clinical signs similar to those described above. Tissue samples of the lung, mesenteric lymph node, and small intestine from the calf were submitted to MVRDL for laboratory diagnosis. In both cases, the clinical manifestations and fatal outcomes were compatible with hemorrhagic syndrome.5,22
Gross examination of the adult beef cow revealed multifocal shallow erosions and ulcers in the oral cavity, predominantly along the soft palate and tongue. Numerous erosions and ulcers were also present along the entire length of the esophagus. The lung contained multiple areas of hemorrhage. Mucosa in the abomasum was hyperemic. The uterus contained an ~5–6-month-old bull calf fetus. No significant lesions were observed in the rumen, reticulum, omasum, small intestine, large intestine, liver, kidney, adrenal gland, spleen, and brain. With regard to the beef calf, the lung was congested and multifocal hemorrhage was present. No significant gross lesions were seen in mesenteric lymph node and intestine tissue samples. Histological examination of the tissues from the beef cow revealed multifocal ulcerative and lymphocytic stomatitis, multifocal ulcerative esophagitis, moderate eosinophilic and lymphocytic enteritis, multifocal lymphocytic abomasitis, and multifocal moderate to severe nonsuppurative perivascular encephalitis. Due to autolysis, histologic examination was not performed on tissue samples from the beef calf.
Routine aerobic and anaerobic bacterial cultures isolated Escherichia coli and Clostridium perfringens from the intestine of the beef cow. No bacteria were isolated from any other tissues of this animal. Light growth of E. coli was isolated from the intestine, lung, and lymph nodes of the beef calf, suggesting either postmortem contamination or antemortem bacteremia. No parasite ova or oocysts were found in fecal samples of these animals.
As requested by the clients, DFAT was performed to detect BVDV, infectious bovine rhinotracheitis virus (IBRV, Bovine herpesvirus 1), Bovine parainfluenza virus 3 (BPIV-3), Bovine respiratory syncytial virus (BRSV), and Bluetongue virus (BTV) antigens in tissue specimens. Briefly, cryostat frozen sections of fresh tissues were cut at 5 µm and fixed in cold acetone for 10 min at room temperature. After air drying, the sections were stained with appropriate FA conjugates for BVDV, IBRV, BPIV-3, BRSV, and BTV a for 30 min at 37°C, counterstained with Evans blue, b and examined under a fluorescent microscope. c Due to lack of specific fluorescence in the tissue sections examined, both cases were recorded as DFAT negative for BVDV, IBRV, BPIV-3, BRSV, and BTV. IHC, which was conducted simultaneously as described previously, 23 revealed the presence of BVDV antigen in the epithelial cells lining the tubuloacinar glands of the oral and esophageal mucosa of the beef cow. Positively stained macrophages were also observed in the Peyer patches. The IHC result on the beef calf was considered invalid due to compromised tissue structure caused by autolysis.
To resolve the discrepancy between DFAT and IHC results, we performed reverse transcription polymerase reaction (RT-PCR) as described previously. 23 Real-time RT-PCR detected BVDV in the pooled tissue samples of the beef cow as well as pooled samples of the beef calf. Real-time RT-PCR–based genotyping 23 was performed, which indicated that the viral isolates belonged to BVDV-2. 23 Real-time RT-PCR on individual tissue samples detected the presence of BVDV-2 in the esophagus, lung, and oral mucosa, but not in the brain, ear notch, small intestine, and spleen, of the beef cow and in the lung of the beef calf. The real-time RT-PCR results confirmed IHC and clinical diagnosis of BVDV infection in the affected animals (Table 1).
Detection of Bovine viral diarrhea virus (BVDV) by direct fluorescent antibody test (DFAT), immunohistochemistry (IHC), and real-time reverse transcription polymerase chain reaction (real-time RT-PCR).*
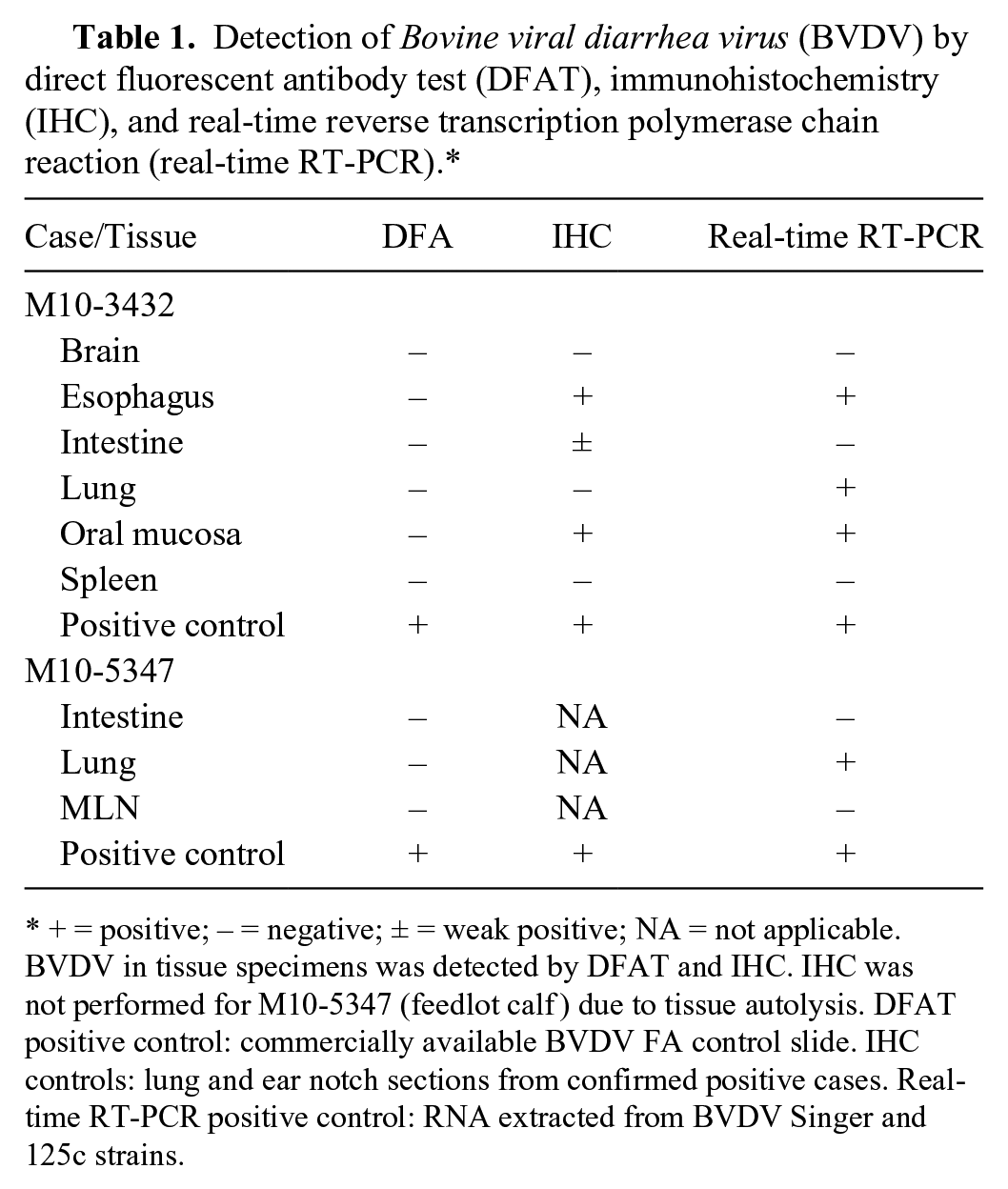
+ = positive; − = negative; ± = weak positive; NA = not applicable. BVDV in tissue specimens was detected by DFAT and IHC. IHC was not performed for M10-5347 (feedlot calf ) due to tissue autolysis. DFAT positive control: commercially available BVDV FA control slide. IHC controls: lung and ear notch sections from confirmed positive cases. Real-time RT-PCR positive control: RNA extracted from BVDV Singer and 125c strains.
As part of the ongoing BVDV research, virus isolation was performed using Madin–Darby bovine kidney (MDBK) cells as previously described. 23 BVDV-1 (Singer) and BVDV-2 (125c) were obtained from a reference laboratory d and served as positive controls. Briefly, the tissue samples were pooled and homogenized in 10 mL of Hanks buffered saline solution (HBSS) containing penicillin (100 U/mL), streptomycin (100 µg/mL), and amphotericin B (2.5 µg/mL). One milliliter of supernatant from each clarified homogenate was used to inoculate MDBK cells at 70–90% confluence in T-25 tissue culture flasks. As the negative control, MDBK cells were mock-infected with 1 mL of HBSS. The inoculated cells were incubated at 37°C in 5% CO2 and inspected daily for 5 days to observe virus-induced cytopathogenic effect (CPE). To detect noncytopathic BVDV, inoculated cells were stained with fluorescein isothiocyanate (FITC)–conjugated anti-BVDV antiserum a according the manufacturer’s instruction. Noncytopathic BVDV and cytopathic BVDV were isolated from the beef cow and beef calf, respectively. For downstream serologic characterizations, the viral stock was titrated using the Reed–Muench method. 18
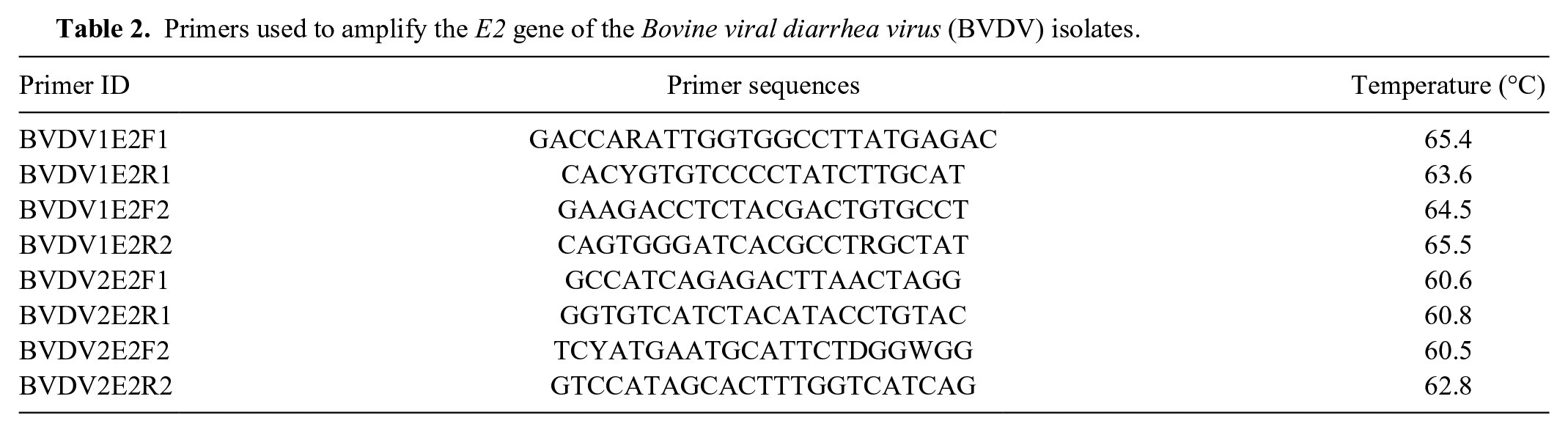
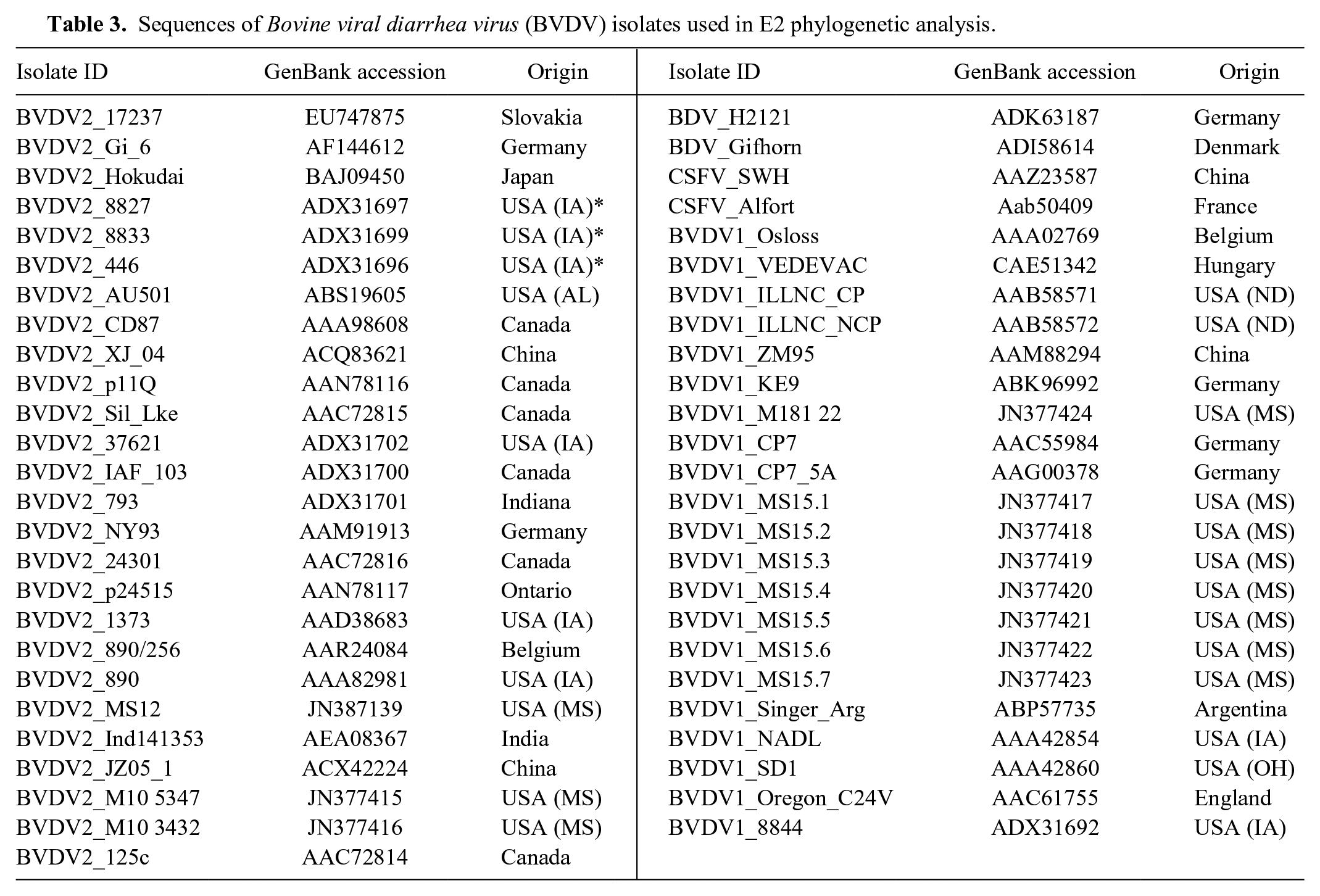
Subsequently, the 5′-UTR and E2 gene of the isolates was amplified by PCR and cloned into the TOPO (topoisomerase I) TA (Taq-amplified) cloning vector e as described previously. 23 Plasmid DNA was extracted using a commercial kit, f and the inserts were sequenced by a commercial DNA company. g A primer walking technique was used to obtain the complete sequence of the E2 gene. The primer sequences are listed in Table 2. Phylogenetic analysis was performed based on the 5′-UTR nucleic acid sequences and the deduced E2 amino acid sequences. Sequence alignments were performed using the Muscle program. 8 The best-fit model of nucleotide and amino acid sequences were selected using the Modeltest 3.716,17 and the ProtTest 2.4, 1 respectively. Phylogenetic trees were constructed using the Bayesian and maximum likelihood (ML) methods.10,13 The reliability of phylogenetic tree was estimated using the bootstrap analysis with 1,000 replicates, and a value >70% (70/100 for ML and 0.7/1.00 for Bayesian) supports the phylogenetic position. Phylogenetic analysis of the 5′-UTR and E2 gene classified both isolates as BVDV-2 subgenotype a. Figure 1 shows the dendrogram based on the E2 amino acid sequence of various BVDV isolates. Isolate ID, GenBank accession number, and the origin of the isolate are provided in Table 3. Our analysis showed that isolate M10-3432 (beef breed cow) was closely related to MS12 (BVDV-2) and BVDV-2a 890, with the former being associated with an acute BVDV infection in Mississippi and the latter being a highly virulent strain isolated from a yearling heifer in Canada. Isolate M10-5347 (feedlot calf) was closely related to cytopathic strains JZ05-1 and 125c. Strain JZ05-1 was isolated from China, and strain 125c was isolated from an animal with mucosal disease after vaccination.
Primers used to amplify the E2 gene of the Bovine viral diarrhea virus (BVDV) isolates.

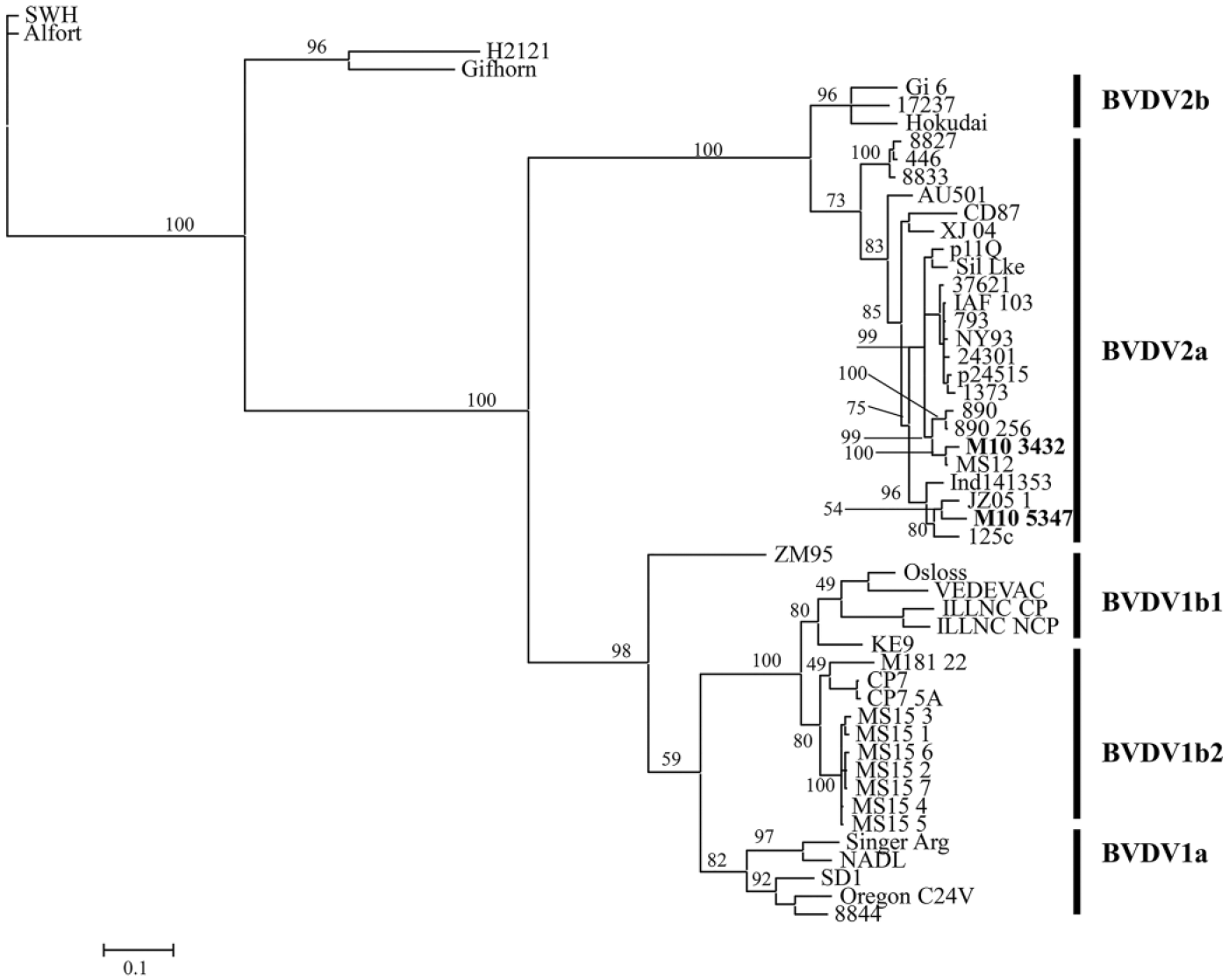
Phylogenetic analysis based on the deduced E2 amino acid sequences (373 aa) of 51 Bovine viral diarrhea virus (BVDV) strains (isolates). The virus ID, GenBank accession number, and the origin of the isolates are provided in Table 2. Both M10-3432 (adult beef breed cow) and M10-5347 (feedlot calf ) were classified as BVDV-2a, a subtype consisting of a number of virulent BVDV strains. M10-3432 was closely related with a high virulence strain 890, whereas M10-5347 was grouped with other virulent strains CD87, 125c, and JZ05-1.
Sequences of Bovine viral diarrhea virus (BVDV) isolates used in E2 phylogenetic analysis.
A close inspection of the 5′-UTR sequences identified mutations in the probe binding site (base 16, T to C) of M10-3432 and primer binding site of M10-5347 (base 11, C to T). The impact of 5′-UTR nucleotide mutations on the real-time RT-PCR assay was then evaluated by using in vitro transcribed RNA as described previously. 23 Standard curves analysis revealed a reduced real-time RT-PCR efficiency, as evidenced by the low R2 (correlation coefficient of PCR) of 0.95 for both isolates, compared with 0.99 for the reference strain BVDV 125c (Fig. 2). Analysis of the deduced E2 amino acid sequences revealed multiple unique mutations in the E2 glycoprotein of M10-3432 and M10-5347, compared to BVDV 125c and other BVDV isolates (Fig. 3). Four unique mutations in the E2 glycoprotein of M10-3432 were located at positions 140 (A to V), 162 (R to K), 190 (D to G), and 328 (D to G). Six unique amino acid alterations in the E2 glycoprotein of M10-5347 were identified at the following positions: 29 (T to A), 102 (L to P), 237 (S to P), 272 (M to L), 278 (E to V), and 352 (K to R). Additional E2 mutations common to both isolates were also identified (Fig. 3).
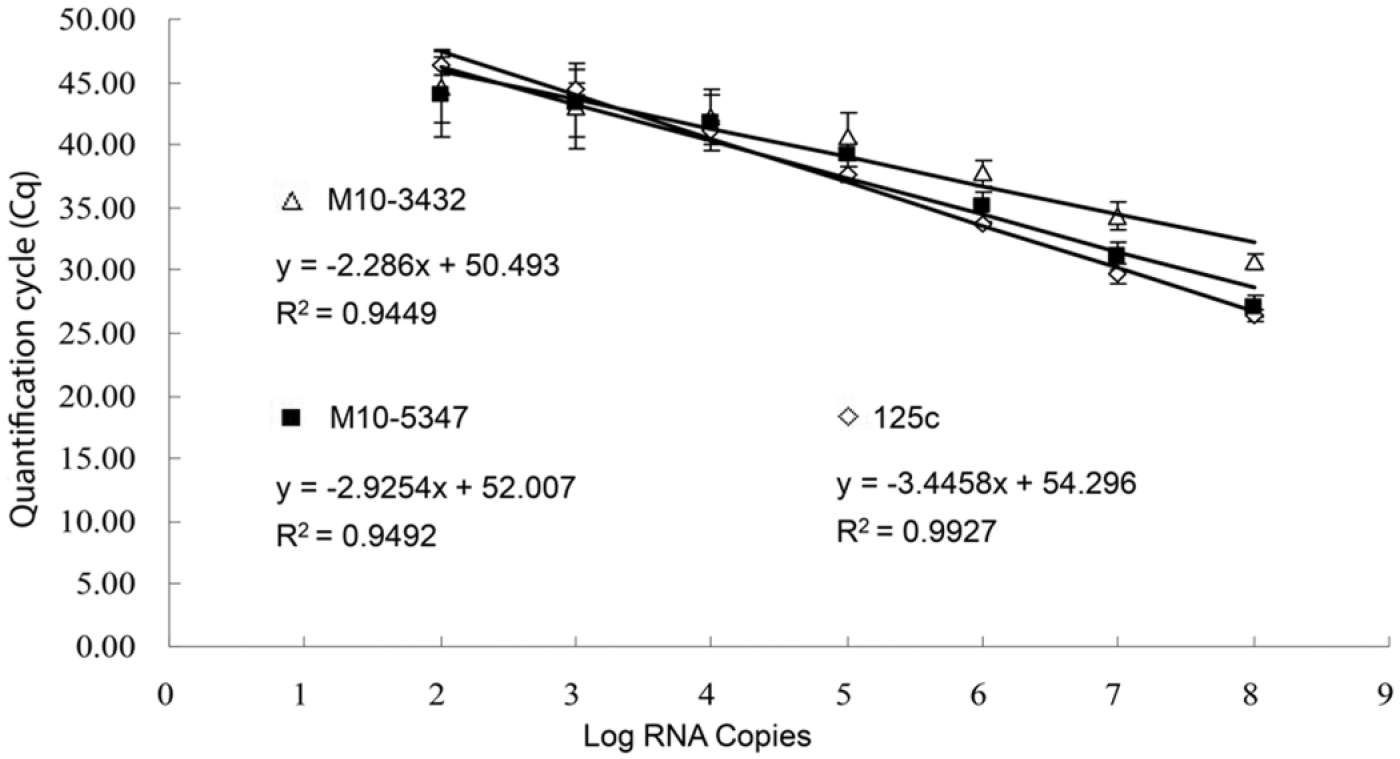
Evaluation of real-time reverse transcription polymerase chain reaction efficiency. Standard curves were constructed based on the quantification cycle (Cq) values and input 5′-UTR RNA copies. RNA was in vitro transcribed from Bovine viral diarrhea virus 2 (BVDV-2) 125c, M103432, and M105347 isolates. Data presented are geometric means of 3 independent experiments with duplicate reactions ± standard deviation.
E2 amino acid sequence alignments. Multiple sequence alignment was performed using the ClustalW program of SDS Biological Workbench (http://workbench.sdsc.edu/). Altered amino acid resides are shown in letters, and consensus sequence are indicated by dots. Alterations unique to Bovine viral diarrhea virus (BVDV) M10-3432 and M10-5347 are indicated by circles and square boxes, respectively.
Because multiple lines of evidence suggested that the beef cow and calf were infected with BVDV-2a, the initial DFAT results were considered as false negative. Based on the extensive mutations in the E2 gene, we hypothesized that E2 antigenic variations interfered with the binding of the isolates to anti-BVDV antisera. To test our hypothesis, serum neutralization was performed using anti–BVDV-1 and anti–BVDV-2 reference sera. d In brief, sera were 2-fold serially diluted in 96-well plates followed by addition of testing viruses (100 TCID50 per well). After 1-hr incubation, MDBK cells (5,000/well) were seeded into the wells and incubated at 37°C. The inoculated cells were observed daily for CPE for 5 days or stained with FITC-conjugated anti-BVDV antiserum a on day 5 to detect noncytopathic BVDV. Duplicate wells were included for each serum dilution, and the experiments were repeated 3 times. Serum neutralization titer was defined as the highest dilution at which the infectivity of 100 TCID50 of the isolates was neutralized. A box-and-whisker plot (Fig. 4) was constructed to display the mean, quartiles, and minimum and maximum of neutralization titers (Fig. 4).
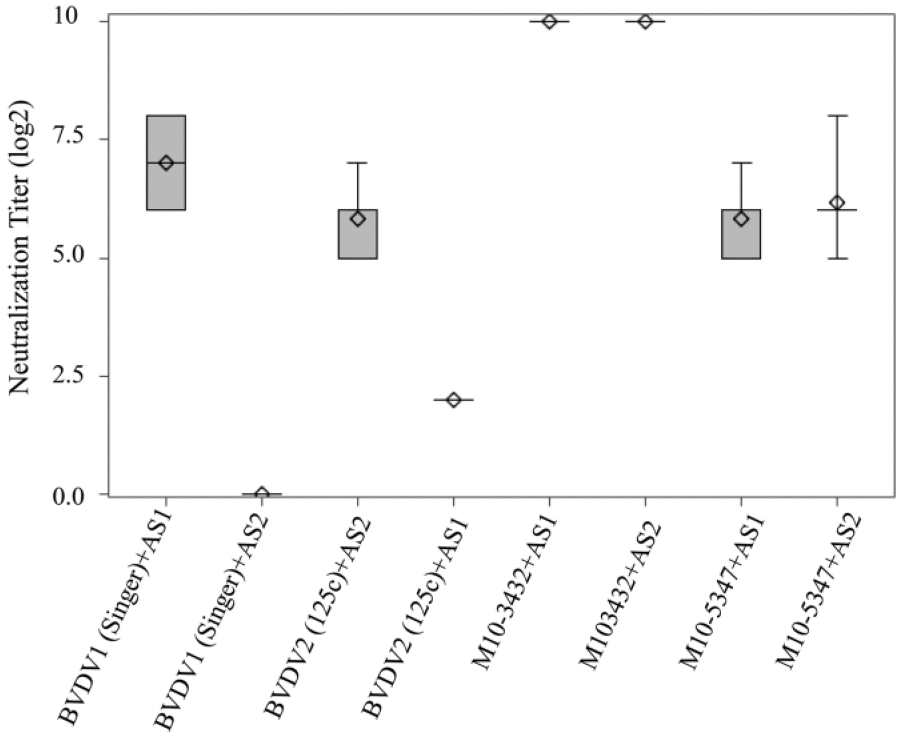
Cross-neutralization of Bovine viral diarrhea virus (BVDV) isolates with anti–BVDV-1 and anti–BVDV-2 antisera. Neutralization titer was determined as the highest dilution of a serum that was able to neutralize 100 TCID50 virus. Titers from 3 independent experiments were presented in the box-and-whisker plot. The plot displays the mean, quartiles, and minimum and maximum observations for each group. The whiskers were drawn from the quartiles to the extreme values of the group. AS1: anti–BVDV-1reference serum; AS2: anti–BVDV-2 reference serum.
Our results showed that reference BVDV-1 and BVDV-2 strains were neutralized specifically by their respective antisera. There was no or minimal cross-reaction between genotype 1 virus and anti–BVDV-2 antiserum or genotype 2 virus and anti–BVDV-1 antiserum. Interestingly, isolates M10-3432 and M10-5347 were neutralized by both anti–BVDV-1 and anti–BVDV-2 sera, suggesting that the BVDV isolates had undergone significant antigenic variations.
Next, we obtained the raw porcine anti-BVDV antiserum from the same manufacturer a that produced the FITC-conjugated porcine anti-BVDV antiserum. The reactivity of the porcine anti-BVDV antiserum to the isolates was evaluated by performing a series of indirect (I)FATs. In brief, MDBK cell monolayers in 96-well tissue culture plates were inoculated with BVDV at a dose of 100 TCID50 per well. BVDV-1 Singer and BVDV-2 125c strains as well as cell culture medium were included as positive and negative controls, respectively. At 48-hr postinoculation, the monolayers were fixed in a mixed solution (50 µL/well) of acetone and methanol (1:1 volume) for 10 min at room temperature. The fixed monolayers were incubated with 2-fold serially diluted porcine anti-BVDV antiserum (25 µL/well) for 30 min at 37°C followed by 3 washes with FA buffer. a The monolayers were treated with FITC-conjugated anti-porcine immunoglobulin G a (25 µL/well) for 30 min at 37°C and then examined under a fluorescent microscope. c The reactivity of porcine anti-BVDV antiserum to different virus strains was expressed as IFAT titer. As shown in the box-and-whisker plot (Fig. 5), the mean IFAT titer for M10-3432 was 1.8–3 logs below the mean titers for BVDV Singer and 125c strains, respectively. The mean IFAT titer for M10-5347 was 2.6–3 logs below the titers for Singer and 125c, respectively. This data clearly indicated that the BVDV isolates were less reactive than the reference strains to the raw porcine anti-BVDV antisera, a major component of BVDV DFAT conjugate.
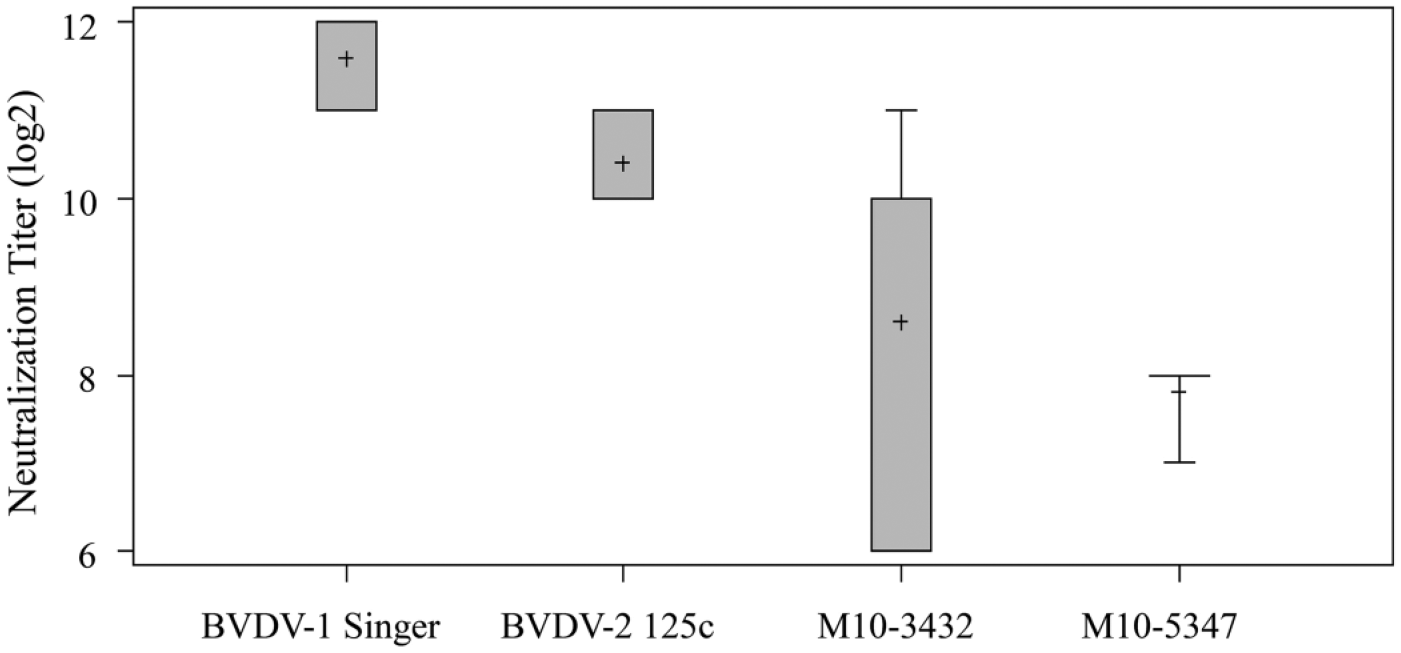
Reactivity of Bovine viral diarrhea virus (BVDV) isolates to raw porcine anti-BVDV polyclonal antiserum that was used to produce BVDV direct fluorescent antibody test conjugate. Reactivity titer was presented as the highest dilution of the serum that was able to detect BVDV antigen on infected Madin–Darby bovine kidney cells. Data from 3 independent experiments was presented in the box-and-whisker plot. The plot displays the mean, quartiles, and minimum and maximum observations for each group. The whiskers were drawn from the quartiles to the extreme values of the group.
To follow up on our initial investigation, serum samples were collected from 10 animals from the breeding farm 2 weeks after the initial submission of M10-3432. Serological assays were performed to detect antibodies against BTV, IBRV, and BVDV. BTV ELISA was performed according to the protocol provided by the manufacturer. h IBRV and BVDV virus neutralization tests were performed as described above. The ages of the animals varied greatly, from 3 months to 10 years. The laboratory results showed that 8 animals (65FHC, 2A, 3A, 4A, 72, Bull, Blue15, and Jersey) were positive for BVDV with neutralization antibody titers ranging from 4 to 256. Eight animals (65FHC, 1A, 2A, 4A, 72, Bull, 5A, and Blue15) were positive for BTV (qualitative assay). With regard to IBRV, 8 animals (65FHC, 1A, 2A, 4A, 72, Bull, Blue15, and Jersey) were positive, with titers ranging from 4 to 64 (Table 4). The wide variation in antibody titers against BVDV and IBRV indicated poor herd immunity, which could have contributed to the severe clinical signs and the fatal outcome of BVDV infection.
Serological survey of farm A where case M10-3432 (adult beef breed cow) originated.*
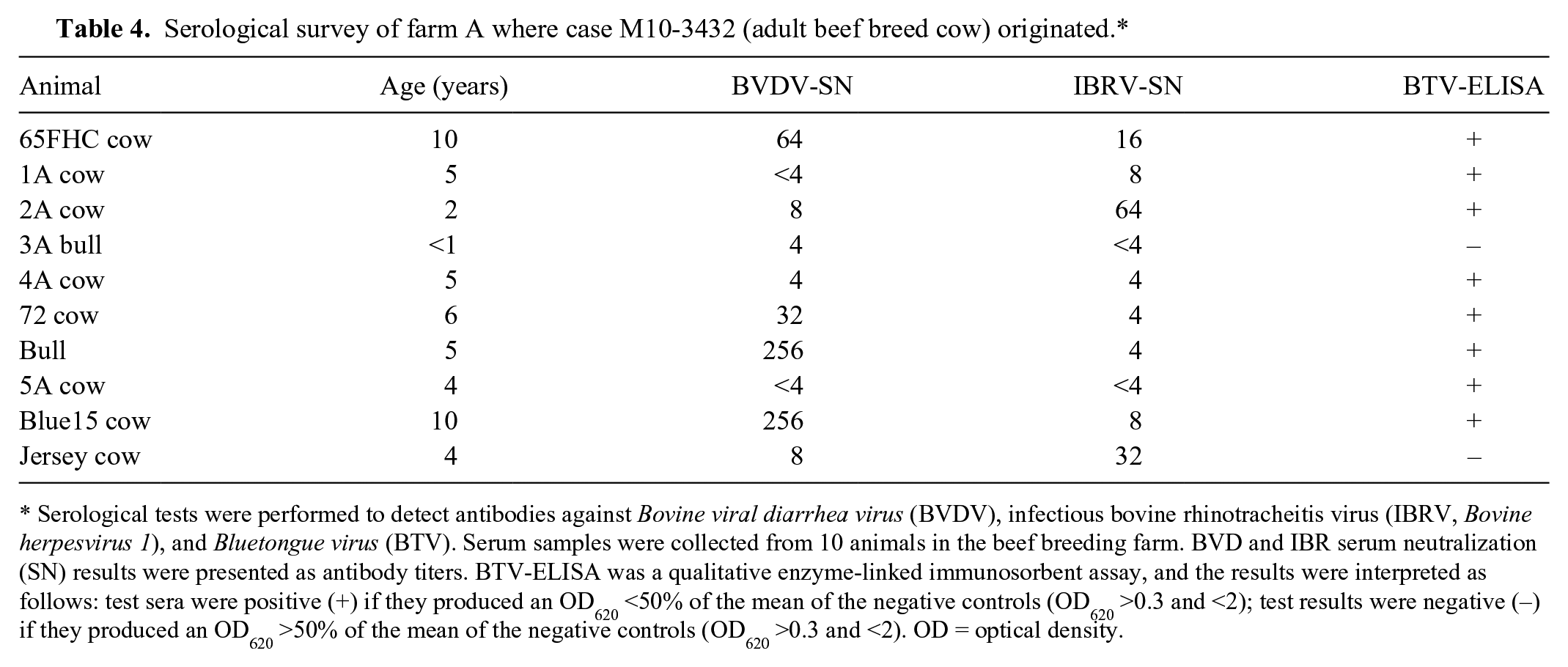
Serological tests were performed to detect antibodies against Bovine viral diarrhea virus (BVDV), infectious bovine rhinotracheitis virus (IBRV, Bovine herpesvirus 1), and Bluetongue virus (BTV). Serum samples were collected from 10 animals in the beef breeding farm. BVD and IBR serum neutralization (SN) results were presented as antibody titers. BTV-ELISA was a qualitative enzyme-linked immunosorbent assay, and the results were interpreted as follows: test sera were positive (+) if they produced an OD620 <50% of the mean of the negative controls (OD620 >0.3 and <2); test results were negative (−) if they produced an OD620 >50% of the mean of the negative controls (OD620 >0.3 and <2). OD = optical density.
In summary, we performed complete diagnostic workups on 2 cases involving a beef cow and a beef calf with severe clinical signs compatible with acute BVDV infection. DFAT, with proper positive and negative controls, failed to detect BVDV antigen. Although autofluorescence associated with lung tissue could complicate the interpretation of DFAT results, it was unlikely the cause of detection failure because DFAT did detect BVDV in any other tissues examined (Table 1). Genetic characterization of the viral isolates revealed mutations in the 5′-UTR and the E2 gene, which were associated with reduced real-time RT-PCR efficiency and amino acid alterations in the E2 glycoprotein, known as a major immunogen for pestiviruses.7,9,11 Serological characterization of the isolates revealed cross-reactivity to anti–BVDV-1 and anti–BVDV-2 reference sera and a low reactivity to commercial porcine anti-BVDV antisera. Compromised sensitivity of IHC and ELISA associated with antigenic variations have been reported.9,11 Data from the present study indicates that genetic and antigenic changes could also affect the performance of DFAT and PCR. Therefore, it is necessary to monitor genetic and antigenic changes in field isolates and evaluate the performance of existing test reagents.
Footnotes
Acknowledgements
We thank Candy Zhang, Tina Hay, Jay Kay Thornton, Latoya Sly, and Corey Williams for their technical assistance. We thank Dr. Patricia Stewart for her critical review of the manuscript.
Authors’ contributions
L Yan contributed to design of the study; contributed to acquisition and interpretation of data; and drafted the manuscript. LW Pace contributed to analysis and interpretation of data, and critically revised the manuscript. B Baughman and FD Wilson contributed to design of the study; contributed to acquisition and analysis of data; and critically revised the manuscript. S Zhang contributed to conception and design of the study; contributed to analysis of data; and critically revised the manuscript. MZ Zhang contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Veterinary Medical Research & Development, Pullman, WA.
b.
Evans Blue, Sigma-Aldrich, St. Louis, MO.
c.
Olympus EX51, Olympus Imaging America Inc., Center Valley, PA.
d.
National Veterinary Services Laboratories, Ames, IA.
e.
TOPO-TA cloning vectors, Life Technologies, Carlsbad, CA.
f.
QIAprep miniprep, Qiagen Inc., Valencia, CA.
g.
DNA sequencing, Eurofins Genomics, Huntsville, AL.
h.
Bluetongue Competition Ab Test, IDEXX Laboratories, Westbrook, ME.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, commercialization, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the Mississippi Veterinary Research & Diagnostic Laboratory, Department of Pathobiology and Population Medicine, College of Veterinary Medicine, Mississippi State University.
