Abstract
Nineteen strains of infectious laryngotracheitis virus (ILTV; Gallid herpesvirus 1) were isolated from dead or diseased birds in chicken flocks from different areas of China between 2010 and 2014 and used to investigate ILTV epidemiology. These strains were characterized using polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) patterns and sequence analysis of the thymidine kinase (TK) gene. PCR-RFLP analysis showed that the TK gene generated 2 patterns when digested with restriction endonuclease enzymes. Pattern A corresponded to 2 virulent field strains, while pattern B was characteristic of 2 virulent field strains, 15 low pathogenicity field strains, and all vaccine strains. Sequence analysis of the TK gene indicated that the messenger RNA polyadenylation signals could be identified in some isolates where amino acid 252 was threonine, and in those with methionine at that position. The present study has demonstrated that most of the outbreaks of ILT in China were caused either by low virulence strains or by vaccine-related strains, and also emphasizes the importance of reinforcing ILTV surveillance in both vaccinated and nonvaccinated flocks.
Infectious laryngotracheitis (ILT) is an acute upper respiratory disease of poultry caused by a double-stranded DNA virus, Gallid herpesvirus 1 (commonly referred to as infectious laryngotracheitis virus, ILTV; order Herpesvirales, family Herpesviridae, subfamily Alphaherpesvirinae, genus Iltovirus). 5 ILT results in significant economic losses due to delayed growth, decreased production, and occasional early death.4,9 The mortality rate in birds is dependent on the virulence of the strains. 8
Two types of attenuated vaccines developed by consecutive passage of virulent virus in embryonated eggs (chicken embryo origin, CEO) or in cell cultures (tissue culture origin, TCO) are now used to control the disease.6,15 However, the vaccine strains can establish latent infections in trigeminal ganglia after the acute phase of infection, and may revert to virulence after reactivation from latency. 11 As a result, after the introduction of virus vaccines, it is difficult to determine whether the new outbreaks of ILT are caused by vaccine-related or virulent strains. Methods using sequencing and/or polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) of multiple genes were thus applied to investigate the epidemiology of ILTV.1,2,10
During the past 5 years, China has experienced many sporadic cases of ILT.13,16 To date, there have been no reports on the epidemiology of ILTV or the relationship between the field and vaccine viruses in China, to our knowledge. In this preliminary study of the epidemiology of ILTV, we characterized 19 ILTV strains isolated from outbreaks of ILT in 5 provinces of China.
Five CEO vaccines (strain K317 from China, strain Connecticut from Germany, strain T20 from France, strain CHP50 from Spain, and strain Serva from The Netherlands) and 1 TCO vaccine (strain Samberg from the United States) were obtained from their manufacturers.
Infectious laryngotracheitis virus strains were isolated from chicken flocks in Guangdong, Guangxi, Jiangsu, Jiangxi, and Hainan provinces in China between 2010 and 2014 (Table 1). The chickens examined were ~30–70 days old, and showed typical respiratory symptoms, with a 0.5–10% mortality in isolated cases. Tracheal samples from dead or diseased chickens were homogenized, and then inoculated into 9–11-day-old specific pathogen–free (SPF) embryonated chicken eggs via the chorioallantoic membrane (CAM) route. Six days after inoculation, the positive isolates produced typical yellowish pock lesions with opaque edges in the CAM. The pathogenicity of the ILTV isolate was evaluated using the mortality index for chicken embryos (MICE). 12
Characteristics of infectious laryngotracheitis virus (ILTV) strains tested in this study.*
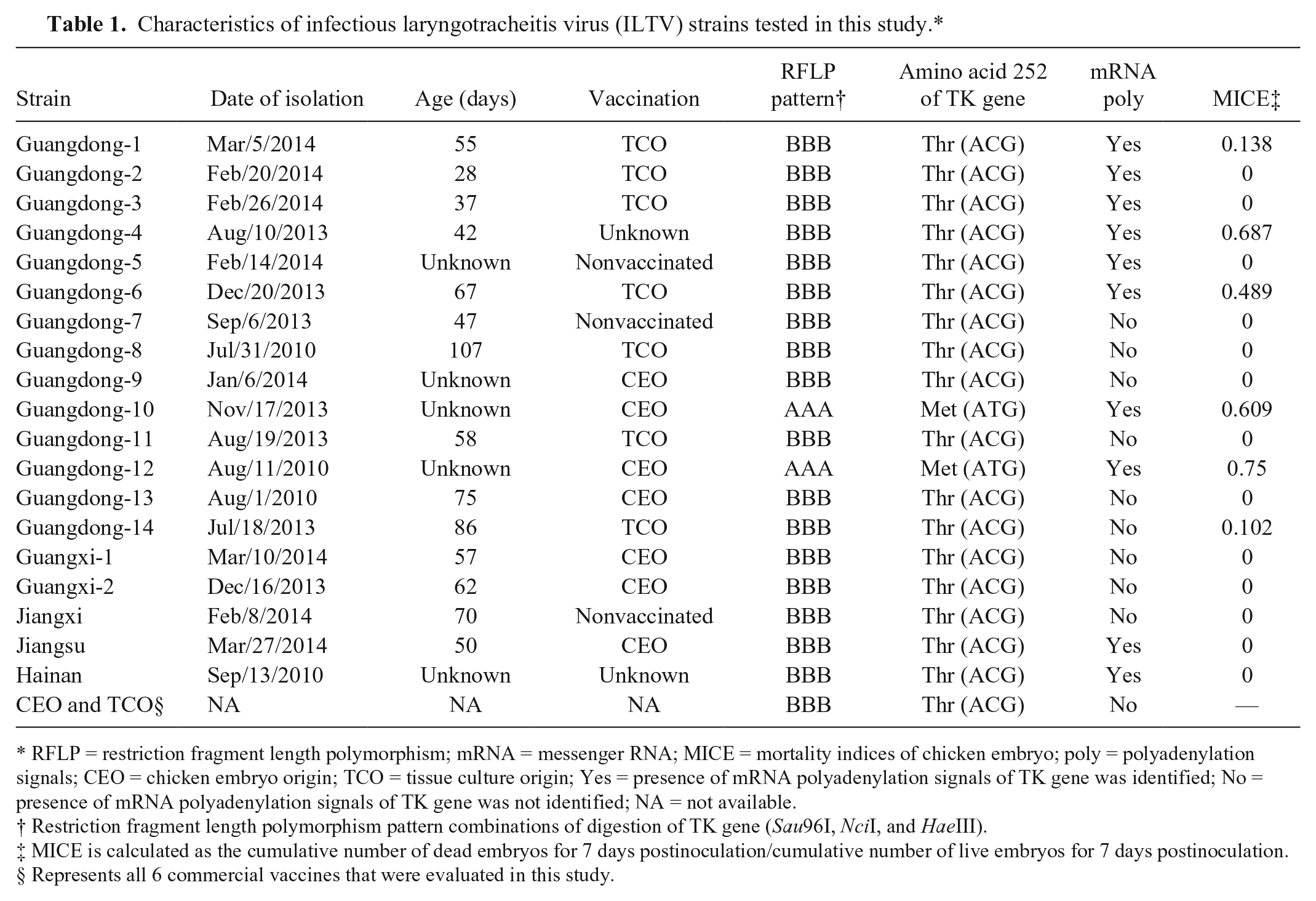
RFLP = restriction fragment length polymorphism; mRNA = messenger RNA; MICE = mortality indices of chicken embryo; poly = polyadenylation signals; CEO = chicken embryo origin; TCO = tissue culture origin; Yes = presence of mRNA polyadenylation signals of TK gene was identified; No = presence of mRNA polyadenylation signals of TK gene was not identified; NA = not available.
Restriction fragment length polymorphism pattern combinations of digestion of TK gene (Sau96I, NciI, and HaeIII).
MICE is calculated as the cumulative number of dead embryos for 7 days postinoculation/cumulative number of live embryos for 7 days postinoculation.
Represents all 6 commercial vaccines that were evaluated in this study.
DNA for PCR-RFLP analysis was extracted from the supernatants of both the homogenized CAMs and the ILTV vaccines using a viral DNA/RNA miniprep kit. a Primers used for amplification of the thymidine kinase (TK) gene were synthesized as described previously. 10 All amplifications were performed using DNA polymerase b following the manufacturer’s recommendations. Ten microliters of each PCR product was digested with 1 µL of restriction endonuclease enzymes (Sau96I, NciI, and HaeIII) c for 5 min, and then separated in 2% agarose gel.
The primer set TK-up (5′-CGGGATCCATCGTATAGGCCAGCCTT-3′) and TK-down (5′-GCTCTAGACCACGCTCTCTCGAGTAA-3′) was used to amplify a 1,296-bp region of the TK gene from all of the field and vaccine strains. The PCR products were purified using a DNA gel extraction kit, d and sent to a genomics facility e for sequencing in both directions. Assembly and analysis of the sequences were done using commercial software. f The sequences of the TK genes were published in the GenBank database with accession numbers from KM115551 to KM115573.
The MICE of the ILTV isolates was calculated using the formula shown in Table 1. The results showed that 15 isolates, each with a MICE <0.16, had a low pathogenicity for chickens. 12 The other 4 isolates (Guangdong-4, Guangdong-6, Guangdong-10, and Guangdong-12), each with a MICE >0.27, were highly pathogenic.
PCR-RFLP analysis showed that the TK gene generated 2 patterns when digested with enzymes Sau96I, NciI, and HaeIII (Fig. 1). Pattern A corresponded to 2 virulent strains (Guangdong-10 and Guangdong-12). Pattern B was characteristic of 2 virulent strains, 15 field strains of low pathogenicity, and all of the vaccine strains (Table 1). The results demonstrated that most of the outbreaks of ILT were caused by strains of low virulence or by vaccine-related strains.
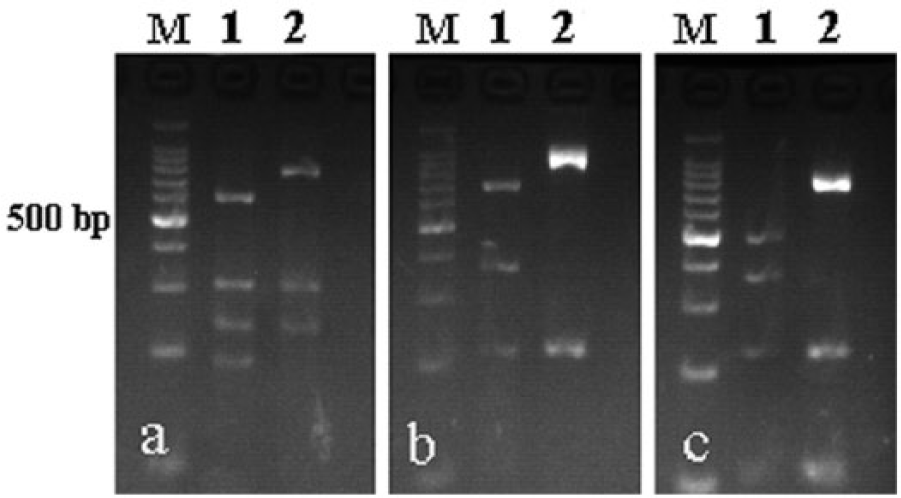
The TK gene of infectious laryngotracheitis virus (ILTV) field strains and vaccine (tissue culture origin and chicken embryo origin) strains digested with Sau96I (
The location of messenger (m)RNA polyadenylation signals (AATAAA) is important for mRNA polyadenylation. 17 For the 19 field strains, when amino acid 252 of the TK gene was methionine (ATG), the typical mRNA polyadenylation signal was consistently identified 24 bp downstream from the stop codon. When the amino acid 252 was threonine (ACG), the mRNA polyadenylation signals could still be identified in 8 field strains. However, for the 6 vaccine strains, the amino acid located at 252 was threonine (ACG), and the mRNA polyadenylation signals could not be identified (Table 1).
To assist in the control of ILTV outbreaks, methods such as RFLP, PCR-RFLP, real-time PCR, and sequencing have been utilized to differentiate vaccine and field strains.15,16 The present study utilized PCR-RFLP and sequence analysis to characterize 6 vaccine strains and 19 field strains of ILTV, collected between August 2010 and March 2014 in China. PCR-RFLP analysis of the TK gene suggested that 17 of the field strains used in our study are indistinguishable from CEO vaccine strains, suggesting a close relationship between field and vaccine strains. Notably, 2 of the 17 isolates (Guangdong-4 and Guangdong-6) were identified as virulent strains, thus providing support for the hypothesis that vaccine viruses tend to displace wild-type viruses in the field. 7 The reason for this may be that the introduction of attenuated vaccines increases the opportunity for ILTV to recombine into fitter variant, as a different pool of genes is available for recombination and selection. 3
We found that the typical mRNA polyadenylation signal (AATAAA) could also be identified in those strains that had threonine at position 252, disagreeing with the results of a previous study. 10 This situation had previously been reported for the virulent strain CSW-1, isolated from layer birds in Australia in 1970, of which strain the complete genome sequence had been determined by high-throughput sequencing. 14
Our study has demonstrated the continued circulation of ILTV strains in chickens in China between 2010 and 2014. Molecular data indicates that those viruses mainly prevalent are low virulence strains or vaccine-related strains. The detection of ILTV strains in vaccinated flocks suggests that further work on examining the efficacy and safety of conventional vaccines, as well as on the design of vaccination programs or new recombinant viral-vectored vaccines, is needed. Taken together, these results manifest the importance of continuous molecular monitoring of ILTV in order to understand the emergence of potentially pandemic strains in China.
Footnotes
Acknowledgements
We thank Guangdong Wen’s Foodstuffs Group for excellent technical assistance.
Authors’ note
Zhuanqiang Yan and Shengpeng Li contributed equally to this study.
Authors’ contributions
Z Yan contributed to design of the study; contributed to analysis of data; and drafted the manuscript. S Li contributed to conception of the study and contributed to acquisition of data. Q Xie and Y Bi contributed to conception of the study and contributed to interpretation of data. F Chen contributed to design of the study and contributed to analysis of data. All authors critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
AxyPrep body fluid viral DNA/RNA miniprep kit, Axygen Inc., Union City, CA.
b.
PrimeSTAR Max DNA polymerase, Takara Biotechnology, Dalian, China.
c.
FastDigest, Thermo Fisher Scientific, Shanghai, China.
d.
AxyPrep, Axygen Inc., Union City, CA.
e.
BGI, Guangzhou, China.
f.
Lasergene SeqMan 11.2, DNASTAR, Madison, WI.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the Natural Science Foundation of Guangdong Province (S2013030013313).
