Abstract
Fetal tissues and placenta from a third trimester Mediterranean miniature donkey (
Gammaherpesviruses are lymphotropic viruses that are characterized as slow-growing in vitro. There are several important human gammaherpesviruses, such as
In April 2014, a 3.9-kg, preterm, female Mediterranean miniature donkey fetus was submitted for autopsy to the Animal Disease and Food Safety Laboratory (Sparks, Nevada). The dam was reported to be current on vaccines and was 1 of 11 equids on a property that housed horses, miniature donkeys, and miniature zonkeys (zebra–miniature donkey hybrids). The dam previously had a full-term abortion 2 years prior. The previous abortion occurred on an adjacent property while the jenny was housed with a pony, now deceased. There were no other reproductive losses on either farm. The fetus was submitted along with an intact allantoamnion and allantochorion. The stage of pregnancy was estimated to be early to mid-third trimester. No gross lesions were noted on the fetus or the allantoamnion. The allantochorion was covered with multifocal, small, yellow-gray regions of altered villi with a gritty consistency; these villi grossly appeared necrotic. Formalin-fixed tissues and fresh, frozen placenta were submitted to the Washington State University, Washington Animal Disease Diagnostic Laboratory (Pullman, Washington) for additional diagnostic evaluation.
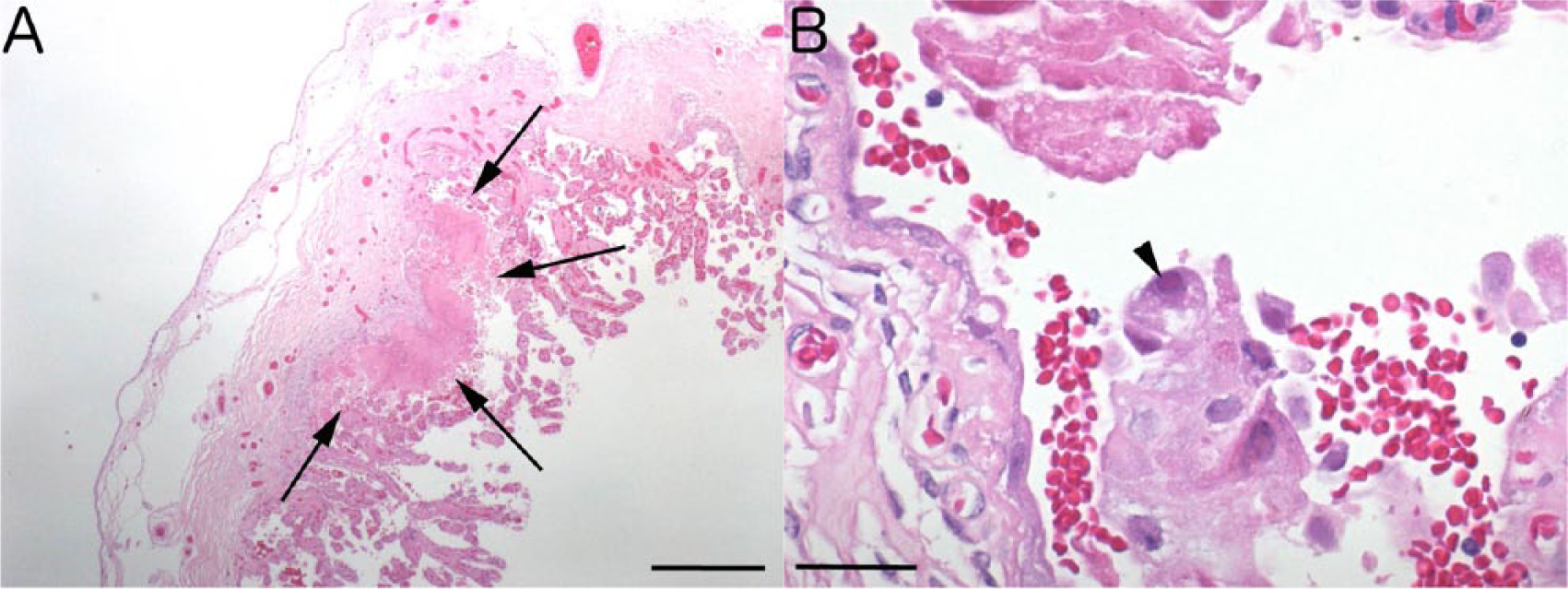
Microscopic evaluation of the placenta revealed scattered hypereosinophilic foci within the chorionic villi accompanied by loss of cellular and nuclear detail (Fig. 1). Chorionic stroma subjacent to these foci was occasionally infiltrated with small numbers of lymphocytes and macrophages. Occasional cells within and adjacent to the necrotic foci had eosinophilic, intranuclear inclusions (Fig. 1B). The fetal heart, spleen, liver, kidney, lung, thymus, ovary, and umbilicus were examined and were considered microscopically normal.
Based on the placental lesions, virus isolation was performed on frozen, fresh placenta using standard detection techniques on multiple cell lines.
4
Processed tissue supernatant was inoculated onto confluent monolayers of rabbit kidney–derived (RK)-13B and primary equine kidney cells. Positive controls consisted of
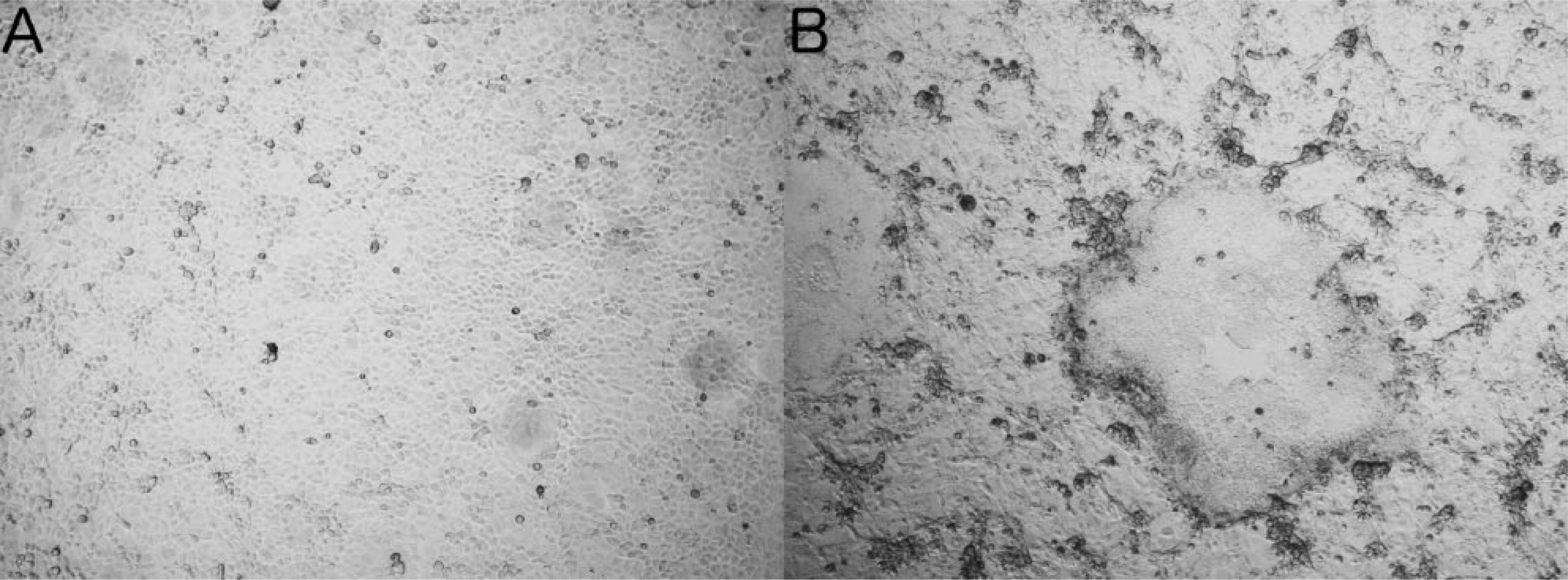
Supernatant from the virus amplified in RK-13B cells showing CPE was prepared for ultrastructural analysis using standard electron microscopic negative stain. 7 Examination of uranyl acetate–stained grids revealed roughly hexagonal nucleocapsids ~100 nm in diameter, surrounded by electron-sparse material interpreted as envelope, consistent with herpesvirus ultrastructural morphology 5 (Fig. 3).
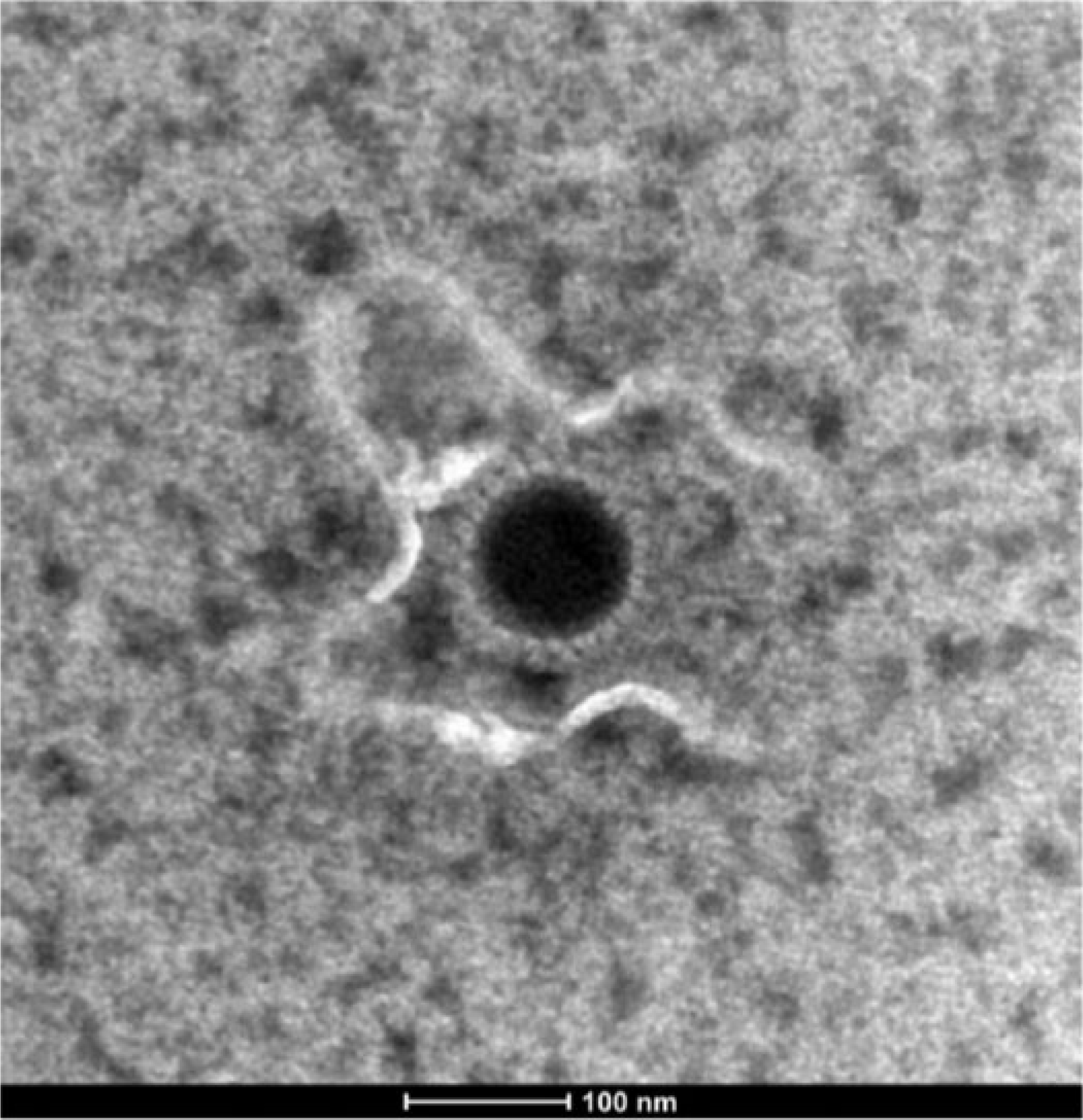
Electron photomicrograph of the virus isolated from the placenta of a miniature donkey abortion. Nucleocapsid is surrounded by electron-sparse material, consistent with a viral envelope. Uranyl acetate stain. Bar = 100 nm.
The supernatant fluids from infected RK-13B cell cultures were subjected to polymerase chain reaction (PCR) followed by sequencing. Specifically, DNA was extracted using a commercial kit following manufacturer directions. a A single-round PCR for amplification of a portion of the DNA polymerase gene was performed using herpesvirus-specific degenerate primers as previously described. 14 Positive results were obtained from the infected cell line as compared with the negative control, and a 699–base pair (bp) sequence was obtained from the infected RK-13B cell culture DNA extraction. Sequences were compared with data available in the National Center for Biotechnology Information database (GenBank). The partial sequences of the herpesvirus DNA polymerase gene clustered with equid gammaherpesviruses (Fig. 4). 6 The virus had 96.7% sequence similarity to a portion of the DNA polymerase gene from EHV-7 (GenBank accession EU165547) and 98–100% sequence similarity to short (166 bp) sequences of the DNA polymerase gene of asinine herpesvirus 4 (GenBank accession AY054991.1, AY054992.1, AY054990.1). Sequence similarity, along with moderately high (≥95%) sequence coverage, was 91% to asinine herpesvirus 5 (GenBank accession FJ98319.1), 89% to EHV-5 (GenBank accession JX125459.1), and 87% to EHV-2 (GenBank accession HQ247790.1). EHV-1 and -4 were not detected via PCR, using specific EHV-1 and EHV-4 primers on infected RK-13B cell culture supernatant. An identical sequence was obtained when the same degenerate herpesvirus primers were used for PCR on the formalin-fixed, paraffin-embedded placenta. DNA was extracted from the formalin-fixed, paraffin-embedded placenta using a commercial kit b as previously described. 16 The sequence obtained for the WADDL 2014-5403 herpesvirus was deposited into GenBank as accession KP245904.
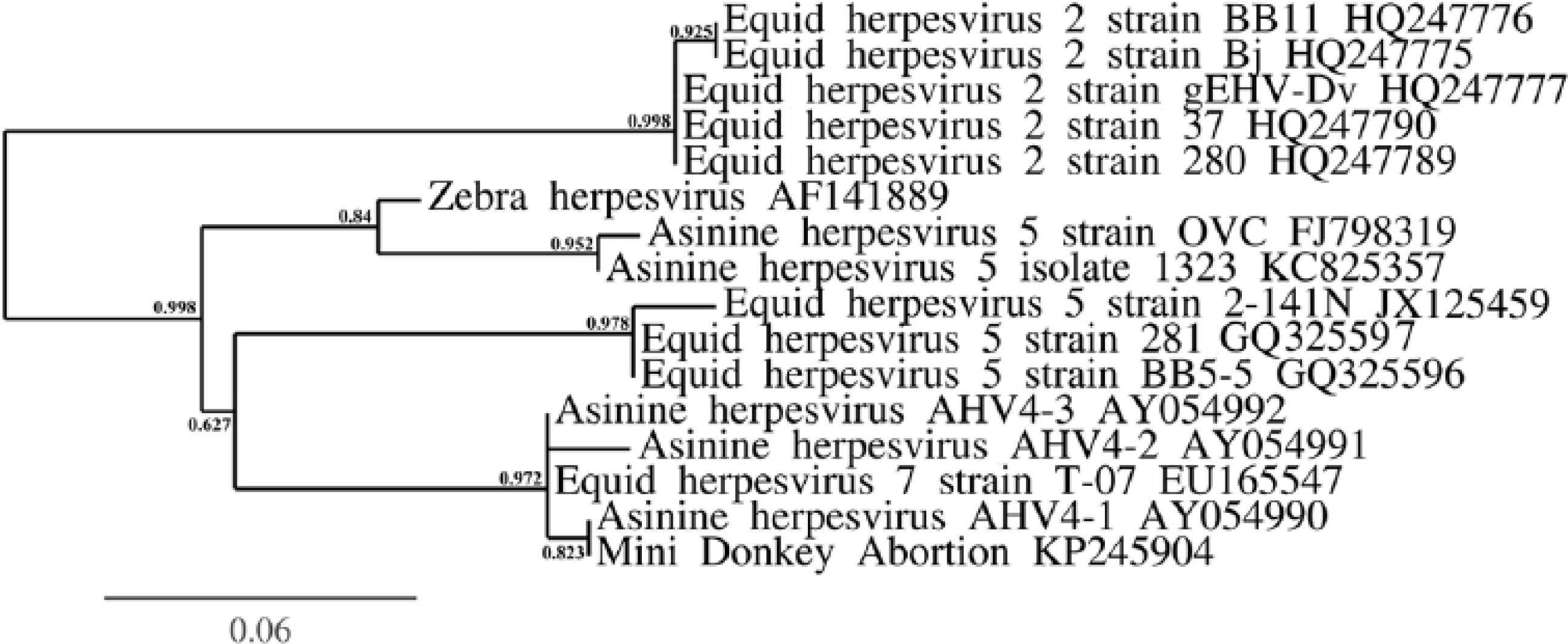
Phylogenetic relationship, using the approximate likelihood ratio test for branches, 4 between the abortion-associated WADDL 2014-5403 herpesvirus (listed as “mini donkey abortion”) and the other viruses to which it had greatest nucleotide identity (all equine gammaherpesviruses).
Because the sequence similarity level between the virus reported herein and EHV-7 was only 96.7%, this suggested that the virus isolated from the placenta of this miniature donkey abortion may represent a new species of equid herpesvirus, the sequence of which has not been previously reported. However, it is also known that there is sequence heterogeneity within many of the gammaherpesviruses, so the virus isolated in our study may simply be a variant of EHV-7. 13 While there was a high degree of similarity to sequences from the DNA polymerase gene of asinine herpesvirus 4, the banked sequences of AHV-4 are too short (only 166 bp) to imply that the virus described is AHV-4. In fact, in the report of EHV-7 isolated from a healthy mule, it was also noted that a portion of the DNA polymerase gene had 100% sequence identity to a short portion of AHV-4. 1
Approximately 4 months after the abortion occurred, sera were collected from the dam of the aborted fetus along with 7 herd mates. All the miniature donkeys and horses on the property were sampled (miniature donkey–zebra hybrids could not be sampled), and microtiter virus neutralization assays were performed in order to detect virus-neutralizing antibody against EHV-1 and -4, EHV-3,
Familial distribution of virus-neutralizing antibody against the abortion-associated WADDL 2014-5403 virus and unrelated
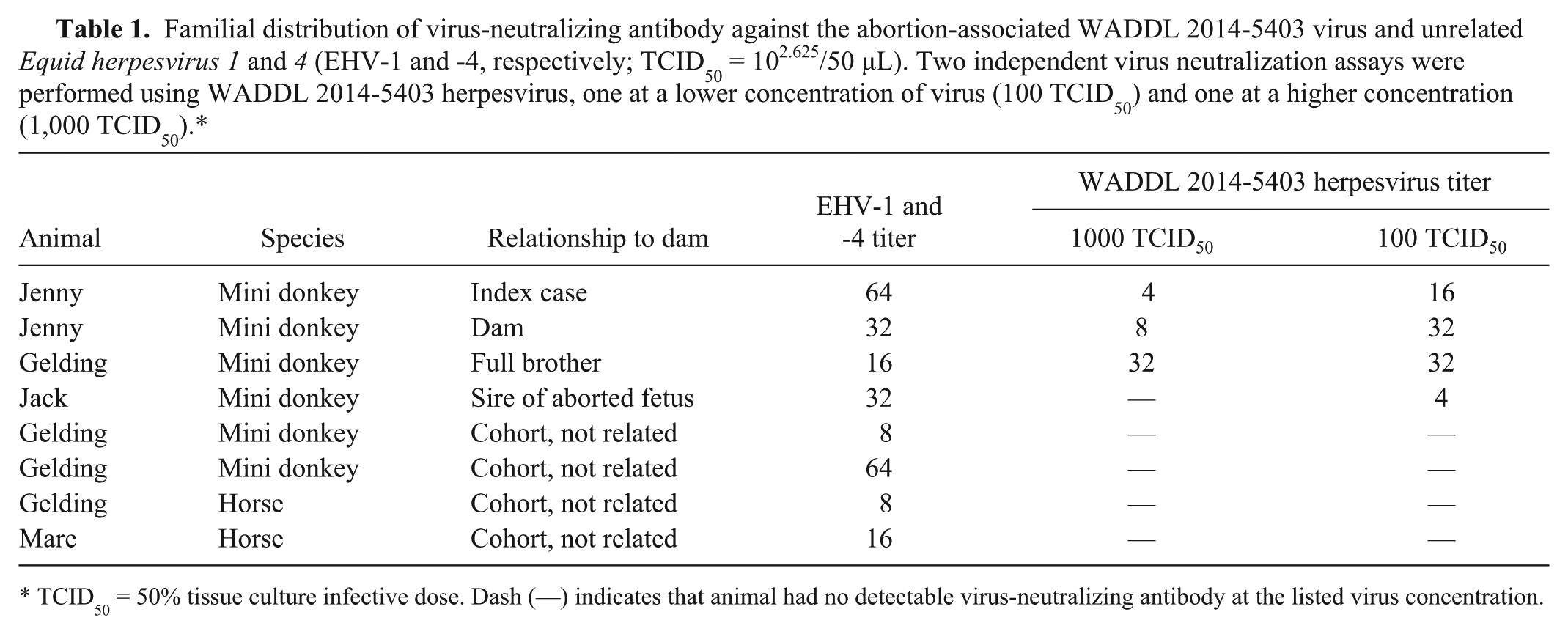
TCID50 = 50% tissue culture infective dose. Dash (—) indicates that animal had no detectable virus-neutralizing antibody at the listed virus concentration.
The microscopic lesions reported in our study are novel for herpesvirus-associated abortion in an equid. Alphaherpesviruses, such as EHV-1, are well-known abortifacient viruses in horses. Histologically, the placenta is not affected, and typical microscopic lesions include multifocal necrosis of the fetal lung, liver, spleen, and kidney, often accompanied by viral intranuclear inclusion bodies and surrounded by mononuclear inflammation. This is in sharp contrast to the placentitis observed in this abortion without any accompanying lesions in the fetal internal organs. In fact, examination of placenta was required in order to make this diagnosis, which is important to note when examining tissues from other donkey abortions submitted to veterinary diagnostic laboratories. Although the gammaherpesviruses EHV-2 and EHV-5 have been infrequently isolated from aborted equine fetal tissues (0.75% and 0.25% incidence, respectively), the role of these viruses in the pathogenesis of equid abortion remains unclear. 12
Evidence suggests that the etiologic agent associated with the miniature donkey abortion described herein is a novel gammaherpesvirus similar to EHV-7, as supported by isolation of the virus from the placenta, and histologic observation of intranuclear viral inclusion bodies, which are compatible with herpesvirus inclusions, that accompanied placental necrosis. Additional work will be required to further compare this potentially novel virus to EHV-7, and to better understand the association between this virus and fetal loss in donkeys and other equids.
Footnotes
Acknowledgements
We thank Lorraine Tanaka, Jessica Schiflett, Jennifer Baisley, and Dr. Christine Davitt for technical assistance.
Authors’ contributions
A Rink contributed to conception of the study and acquisition of data. JF Evermann and GJ Haldorson contributed to conception and design of the study. TE LeCuyer, DS Bradway, and T Baszler contributed to design of the study. TE LeCuyer, DS Bradway, JF Evermann, and GJ Haldorson contributed to acquisition, analysis, and interpretation of data. AV Nicola and T Baszler contributed to interpretation of data. TE LeCuyer drafted the manuscript. A Rink, DS Bradway, JF Evermann, AV Nicola, T Baszler, GJ Haldorson critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Ambion MagMax AM1836TM-96 viral isolation kit, Thermo Fisher Scientific Inc., Waltham, MA.
b.
QIAamp DNA minikit, Qiagen Inc., Valencia, CA.
c.
070-EDV equine viral arteritis virus, 370-EDV EVA antiserum, 040-EDV EHV-1 virus, 340-EDV EHV-1 antiserum, 042-EDV EHV-3 virus, 342-EDV EHV-3 antiserum; National Veterinary Services Laboratories, Ames, IA.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Research was funded by teaching funds provided by the Department of Veterinary Microbiology and Pathology and by the Washington Animal Disease Diagnostic Laboratory, Washington State University. Dr. Tessa LeCuyer is supported by an Achievement Rewards for College Scientists Fellowship.
