Abstract
An epidemic that mainly endangered 3–7-day-old piglets struck many farms in Shandong Province, China in 2013 and caused heavy losses. To identify the pathogenesis, the type of lesions, and the causative agent, systemic examinations were performed. Autopsy showed multiple lesions, including necrotic foci of the spleen and liver, punctate hemorrhage of the renal cortex, and interstitial pneumonia. Histological examinations showed typical nonsuppurative encephalitis, necrotic lymphocytes, and reticuloendothelial cells in lymphatic tissues, as well as eosinophilic inclusion bodies in the nuclei of reticuloendothelial cells, necrotic foci in liver cells, and hemorrhagic glomeruli. The average seroprevalence rate of field pseudorabies virus (PRV; Suid herpesvirus 1) of a representative farm tested by enzyme-linked immunosorbent assay was 46%, indicating that the PRV infectious pressure was quite severe especially among gilts, young multiparous sows, boars, and growing–finishing pigs. The glycoprotein E (gE) gene of PRV was detected in 8 of 10 clinical samples, and the virus in the positive samples induced obvious cytopathic effects. An immunoperoxidase monolayer assay showed that PRV antigens were distributed both in the nucleoli and cytoplasm of infected cells. Sequencing and phylogenetic analysis of the gE gene showed that the strain isolated herein, TaiAn SD 2013, was highly similar to previously isolated strains, especially those isolated in northern China in 2013, and was closely related to other isolates from Asia. Evidence confirmed that the variant PRV was the etiologic agent of this epidemic, suggesting that the Bartha-K61 vaccine does not provide complete protection against PRV infection. Further challenge tests are ongoing to investigate the virulence of variant PRV.
Aujeszky’s disease, more generally known as pseudorabies (PR), is caused by the pseudorabies virus (PRV; Suid herpesvirus 1; order Herpesvirales, family Herpesviridae, subfamily Alphaherpesvirinae, genus Varicellovirus), which can infect both livestock and wild animals. 9 Infected animals exhibit a range of symptoms, including fever, pruritus (except pigs), reproductive failure, and encephalomyelitis.2,12 Pigs are the natural reservoir of PRV and the only species that can survive infection with the virus. 3
Pseudorabies virus infects pigs at various production phases and causes high mortality rates and nervous system disorders in newborn piglets, respiratory disorders in older pigs, and reproductive failure in pregnant sows, resulting in significant economic losses to the swine industry.17,18 PRV continues to spread worldwide, especially among some countries in South America, Europe, and Asia, while others, including Germany, Sweden, Denmark, and England, have eradicated PR. The United States, France, Poland, and other countries have eradicated PR among domestic pig populations (Lipowski A, Pejsak Z, Antibody prevalence of pseudorabies virus in feral pigs in Poland, In: Proceedings of 17th International Pig Veterinary Society Congress, 2002;2:223). 5
Like other alphaherpesviruses, PRV can establish a lifelong infection in the peripheral nervous system of infected pigs, which can be reactivated under certain conditions, releasing viral particles into the environment. 12 In this way, neighboring pigs are at a high risk of infection, thus latent infection should be considered in eradication and control programs. 4
For ~30 years, attenuated live or inactivated vaccines have been used to control outbreaks of PR.6,16 Bartha-K61 is the most popular (>80% of pigs in China were vaccinated) attenuated strain to produce vaccines because of the natural lack of the glycoprotein E (gE) gene and the development of a gE-specific enzyme-linked immunosorbent assay (ELISA) used for the differential diagnosis of infection with field strains and vaccine strains.7,15 Thus, Bartha-K61 has played a critical role in the control and eradication of PRV since the 1990s, and vaccine immunization has been considered the most effective measure for the prevention and control of PR. 1
However, beginning in late 2011, a severe PR epidemic occurred among some large pig farms and resulted in high morbidity and mortality among pigs that were vaccinated with the Bartha-K61 vaccine.1,11 The outbreak rapidly spread throughout northern China and mostly affected piglets and growing pigs with a reported mortality rate of 10–50%, while a later epidemic in southern China resulted in relatively slight losses and seroconversion among pigs in eradicated farms.1,18 Here, we report an outbreak of variant PRV among Bartha-K61–vaccinated piglets, which devastated swine production in central Shandong Province in 2013. Approximately 200,000 pigs were affected, and the mortality rate among newborn piglets was 17–48% (according to interviews and investigations of 86 pig farms). We systemically analyzed the outbreak to identify the viral pathogenesis, as well as associated lesions and viral characteristics on a representative farm.
On the representative farm, a total of 74 piglets, born 3–7 days prematurely, exhibited a series of symptoms, including anorexia, hydrostomia, vomiting, diarrhea, tremor, and posterior paralysis. The morbidity rate among the newborn piglets was 39% (74/190), and 23 of the 74 piglets died within 2 days, yielding a mortality rate of 31%. Other pigs did not show obvious symptoms except for spontaneous abortions of 2 sows at gestational ages of 60–70 days. Pigs (sows, gilts, boars, newborn piglets, and 10-week-old growing pigs) on the farm were vaccinated with imported PRV vaccine (Bartha-K61 strain) according to the procedure recommended by the manufacturer. The farm abandoned the eradication program in 2012 because of the high seroprevalence rate of anti-gE antibodies (Abs) of the breeding pigs (16%) and thereafter reduced infection of PRV field strains among breeding pigs through comprehensive immune procedures, quarantine, and biosecurity.
Of the dead piglets, 10 were autopsied in a biosafety laboratory, and the PRV-infected piglets had observable lesions in several organs. Yellowish white necrotic foci were scattered on the surface of livers and spleens. Lesions with necrotic foci and obvious ulcer could be seen on the tonsil surfaces. Hyperemia and edema were observed in the meninges. The renal cortex was densely covered with petechial hemorrhages and necrotic spots. The lung tissues exhibited dark red consolidation and yellowish white necrotic foci. Multiple lymph nodes were hyperemic and turgid (Supplementary Fig. 1).
Tissue samples from the brain, tonsils, spleen, lymph nodes, liver, kidney, and lung were segmented for cryopreservation and fixation by 10% neutral formalin. The fixed samples were processed and embedded in paraffin. Thin sections of the fixed tissues were stained with hematoxylin and eosin and examined microscopically. Histological examination mainly confirmed viral encephalitis and necrotic foci of many tissues. The brain tissue exhibited typical nonsuppurative encephalitis with lymphocyte infiltration around the small blood vessels and formation of glial nodules. The liver tissues contained randomly scattered lymphocytes and macrophages in necrotic foci. Sporadic necrotic foci could be seen in the splenic white and red pulp, while necrotic lymphocytes and reticuloendothelial cells were found in necrotic foci; the nuclei of necrotic cells were disrupted, concentrated, deeply stained, and eliminated. Eosinophilic inclusion bodies were observed in the nuclei of reticuloendothelial cells, which were scattered around the necrotic foci. Neutrophils and macrophages were scattered in the necrotic foci of the tonsils and lymph nodes. Thickened alveolar walls exhibited hyperemia accompanied with infiltration and proliferation of macrophages and lymphocytes, while some lymphocytes were necrotic. Renal tubular epithelial cells were swollen and degenerated, while the glomeruli showed signs of hemorrhage (Supplementary Fig. 2).
A total of 102 serum samples from different production phases were collected to detect anti-gE Abs using a commercial ELISA kit a to differentiate the vaccine strain from field strains. Serum with an S/N (the absorbance of a well with serum to that of a control well with negative control) ratio ≤0.60 was considered positive for infection by PRV field strains. The results (Table 1) showed that the average seroprevalence rate (SPR) of the pigs on the representative farm was 46% (40/87; except for the 2–8-week-old pigs, because maternal anti-gE Abs cannot be differentiated from anti-gE Abs induced by field virus). The SPRs of boars, multiparous sows, and growing pigs were 33.3% (7/21), 33.3% (12/36), and 55% (11/20), respectively, while that of the gilts reached 80% (12/15). Multiparous sows of the 1st to 2nd and 5th to 6th birth orders showed high SPRs of 54.5% (6/11) and 44.4% (4/9), respectively, and the 3 sows at 1–7 days postpartum, which were positive for anti-gE Abs from the 1st to 2nd birth orders, gave birth to infected piglets. As shown in Figure 1, the S/N ratios and SPRs of 2–8-week-old piglets gradually increased and decreased, respectively, while those of 8–20-week-old pigs gradually decreased and increased, respectively; the SPR of 20-week-old growing pigs sharply increased to 80% (4/5). The examination suggested that PRV infectious pressure was quite prominent especially among gilts, young multiparous sows, boars, and growing–finishing pigs.
Seroprevalence of anti–pseudorabies virus (PRV) glycoprotein E antibodies in the study farm by enzyme-linked immunosorbent assay.
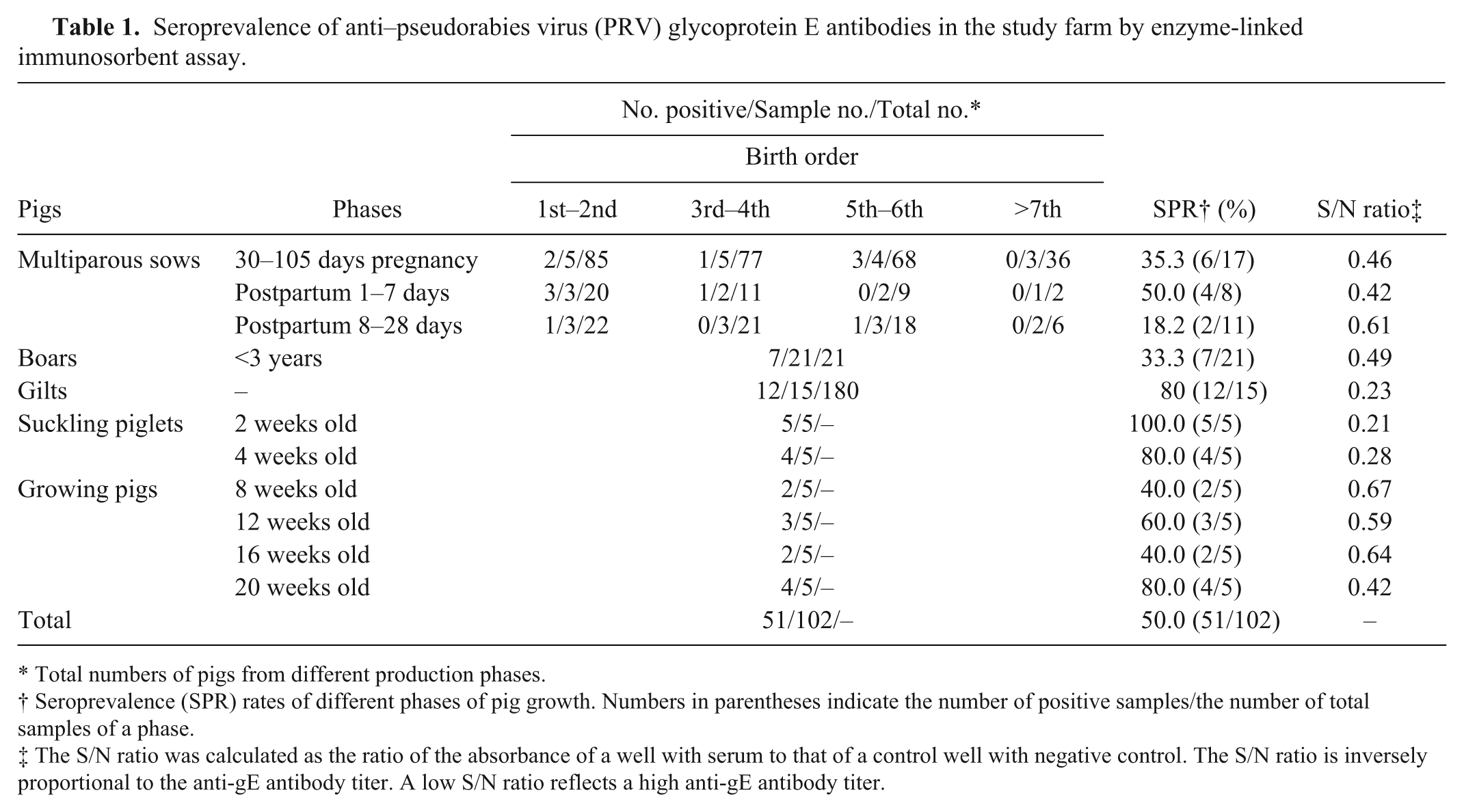
Total numbers of pigs from different production phases.
Seroprevalence (SPR) rates of different phases of pig growth. Numbers in parentheses indicate the number of positive samples/the number of total samples of a phase.
The S/N ratio was calculated as the ratio of the absorbance of a well with serum to that of a control well with negative control. The S/N ratio is inversely proportional to the anti-gE antibody titer. A low S/N ratio reflects a high anti-gE antibody titer.
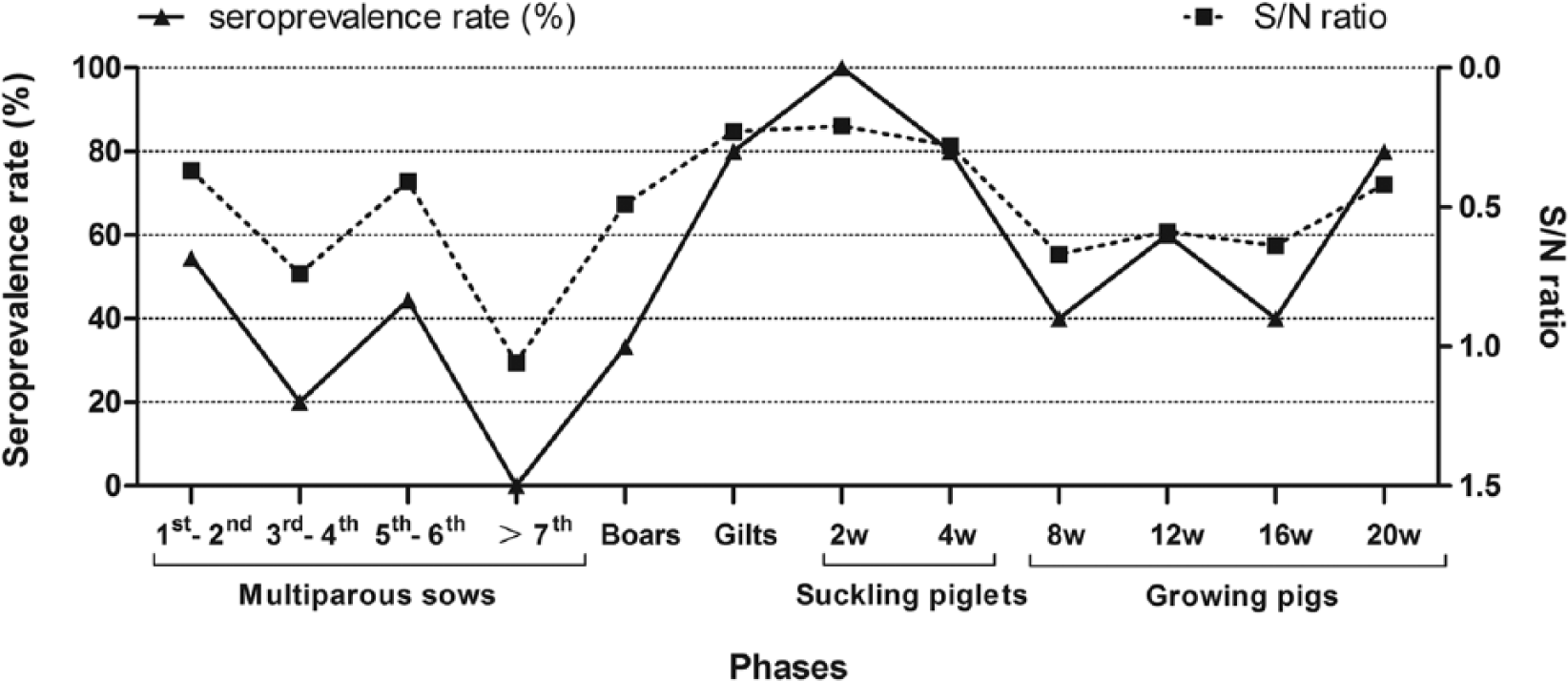
Dynamics of anti–glycoprotein E antibody seroprevalences (SPRs) and S/N ratios (n = 102). Piglets (2–8-week-old) were protected by maternal antibodies, and the SPRs decreased gradually. The growing pigs were injected with Bartha-K61 vaccine at the age of 10 weeks; the vaccine did not confer effective protection, and the majority (80%) of the 20-week-old pigs was infected by pseudorabies virus field strain.
The 10 cryopreserved tissue homogenates consisting of a mixture of brain, tonsils, spleen, and lymph nodes from each of the 10 sampled piglets were centrifuged at 10,000 × g (4°C) for 10 min. The supernatants were prepared for genome extraction. Viral DNA/RNA extraction was performed using a commercial kit b according to the manufacturer’s instructions. The extracted DNA was used for the detection of PRV (partial gE gene, forward: 5′-GATGACCTCAACGGCGAC CTC-3′ and reverse: 5′-GCGAGAAGAGCTGCGAGTG G-3′) and Porcine circovirus-2 (PCV-2; forward: 5′-CCCC AGGAGGGGGCTCAAACC-3′ and reverse: 5′-GCCTACG TGGTCTACATTCCC-3′) through polymerase chain reaction (PCR). c RNA was used the detection of Classical swine fever virus (CSFV; forward: 5′-CCAARAGAGCATGAGAAGG-3′ and reverse: 5′-TTCTCTATAGTGTTGGTCAT TCC-3′) and Porcine reproductive and respiratory syndrome virus (PRRSV; forward: 5′-CGGTTTTGATGGGCGACA-3′ and reverse: 5′-TGCAGGCGTGCGAGGTAA-3′) through reverse transcription (RT)-PCR. c The following cycling conditions were used for PCR amplification: 94°C for 5 min followed by 30 cycles of 94°C for 30 sec, 58°C for 30 sec, and 72°C for 50 sec, and a final extension at 72°C for 10 min. The following cycling conditions were used for one-step RT-PCR amplification: 45°C for 25 min, 94°C for 5 min followed by 30 cycles of 94°C for 30 sec, 56°C for 30 sec, and 72°C for 50 sec, and a final extension at 72°C for 10 min. PCR products were subjected to electrophoresis on a 1% agarose gel stained with ethidium bromide and visualized with an ultraviolet light transilluminator. d The results showed that only PRV could be amplified using primers specific to unique gene fragments in 8 (80%) of 10 clinical samples tested (not shown).
Pseudorabies virus–positive tissue homogenates detected by PCR were centrifuged at 800 × g (4°C) for 10 min and filtered through disposable filters e with an average pore diameter of 0.22 µm, and transferred to Vero (kidney cells of the African green monkey) cell monolayers. The growth medium consisted of Dulbecco modified Eagle medium (DMEM f ) supplemented with 10% fetal bovine serum (FBS g ). The inoculum was allowed to adsorb for 1 hr at 37°C. The monolayers were then washed once, incubated in fresh DMEM supplemented with 2% FBS at 37°C in an atmosphere of 5% CO2 in air, and examined daily for signs of cytopathic effects (CPEs). Obvious CPEs appeared in Vero cells 48 hr postincubation, which were characterized by cell contraction, congregation, and detachment (not shown). Cell syncytia were observed in all culture flasks. RNA and DNA extracted from the cell cultures and amplified by RT-PCR or PCR were PRV positive, but negative for CSFV, PRRSV, and PCV-2 (not shown).
For the immunoperoxidase monolayer assay (IPMA), cells inoculated with harvested PRV-positive cell supernatants (the 3rd passage) were washed with phosphate buffered saline (PBS) and fixed with cold methanol for 15 min. The plates were dried, rewashed with PBS, and then incubated with PRV gE monoclonal Ab (dilution 1:800 in PBS containing 1% Tween 20) for 1 hr at room temperature. After washing 3 times with PBS, anti-mouse horseradish peroxidase–conjugated antibody h was used as a secondary antibody (dilution 1:1,000 in PBS containing 1% Tween 20). PRV plaques were detected using H2O2/AEC i (3-amino-9-ethyl-carbazole). The results showed that the infected cells were stained red and that viral antigens were distributed both in the nucleoli and cytoplasm of infected cells, while supernatants of uninoculated cells returned no signal (not shown).
After the appearance of CPEs, cells were collected and stored at −20°C. The harvested cells were prepared for further gE PCR after 3 freeze–thaw cycles. A 2,060-bp target was amplified using a gene-specific primer pair (forward: 5′-GCCACCATCGCAGAGGAACAAT-3′ and reverse: 5′-TCGTTGTCGCTCTCGCTGTAGT-3′). The following cycling conditions were used for PCR amplification: 94°C for 5 min followed by 30 cycles of 94°C for 30 sec, 59°C for 30 sec, and 72°C for 2.5 min, and a final extension at 72°C for 10 min. Three positive PCR products that were amplified from different piglets were randomly selected and cloned into the pMD18-T vector. j The recombinant plasmids together with both the forward and reverse primers were sequenced. k The sequence data of the 3 products were analyzed in case of errors and compared with other PRV gE sequences available in the GenBank database (http://www.ncbi.nlm.nih.gov/genbank). Subsequently, a phylogenetic tree was generated by the MEGA6 software with the neighbor-joining method. 14 A bootstrap analysis was performed using 1,000 replicates. The isolated viruses were designated as PRV TaiAn SD 2013 (SD-2013). The complete gE gene sequence of this strain was 1,740 bp in length (accession KM983048). Analysis revealed that the nucleic acid sequence homology among the SD-2013 strain and other previous strains shared a homology of 97.4–99.7%. Overall, the gE gene sequence of strain SD-2013 was highly homologous with that of recently isolated strains, especially those isolated in northern China in 2013 (KC415027, KC415028, KF017275, and KF130887), with a homology of 99.6–99.7%, but <97.6% when compared with the western strains JF797219, AY368490, M14336, JF460026, EU502923, JF797218, JQ809328, and FJ605135. Furthermore, nucleotide substitutions were observed and especially frequent between G and A in strain SD-2013 (positions 599, 1342, 1531, 1552, and 1553; relevant amino acid substitutions are described in Table 2), while the nucleotides of other Chinese strains were relatively conserved. A sequential GAC insertion was observed at position 142–144 when compared with some western strains, such as Kaplan (JF797218), Rice (M14336), and NiA-3 (EU502923), and the insertion resulted in an Asp (D) insertion at position 48 (Table 2). Alignment of the gE protein sequences of different PRV strains showed that the SD-2013 strain contains 2 amino acid substitutions at 105 (V–A) and 518 (G–K) compared with other isolates (Table 2). We further analyzed the relationship among SD-2013 and other PRV isolates using a phylogenetic tree based on the gE gene sequence and found that the strain formed a tightly clustered branch with strains isolated in northern China in 2013 and were closely related to other isolates from Asia (Fig. 2). Based on full consideration of epidemiological history, autopsy, histologic examination, serum testing, specific PCR, viral isolation, IPMA, and phylogenetic analysis, we arrived at a definite diagnosis of a recent viral epidemic in Shandong Province caused by PRV TaiAn SD2013 in Bartha-K61–vaccinated pig populations.
Protein-coding variations unique to the pseudorabies virus (PRV) SD-2013 strain and/or recently isolated Chinese strains, in comparison to western PRV isolates.*
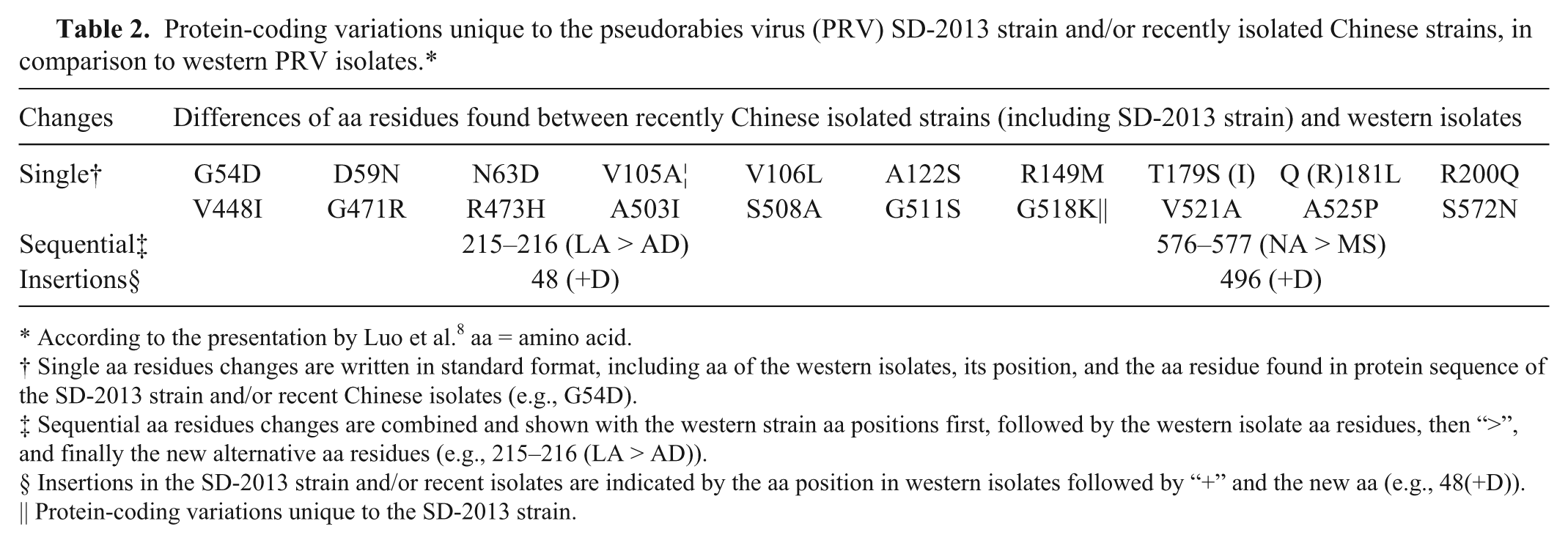
According to the presentation by Luo et al. 8 aa = amino acid.
Single aa residues changes are written in standard format, including aa of the western isolates, its position, and the aa residue found in protein sequence of the SD-2013 strain and/or recent Chinese isolates (e.g., G54D).
Sequential aa residues changes are combined and shown with the western strain aa positions first, followed by the western isolate aa residues, then “>”, and finally the new alternative aa residues (e.g., 215–216 (LA > AD)).
Insertions in the SD-2013 strain and/or recent isolates are indicated by the aa position in western isolates followed by “+” and the new aa (e.g., 48(+D)).
Protein-coding variations unique to the SD-2013 strain.
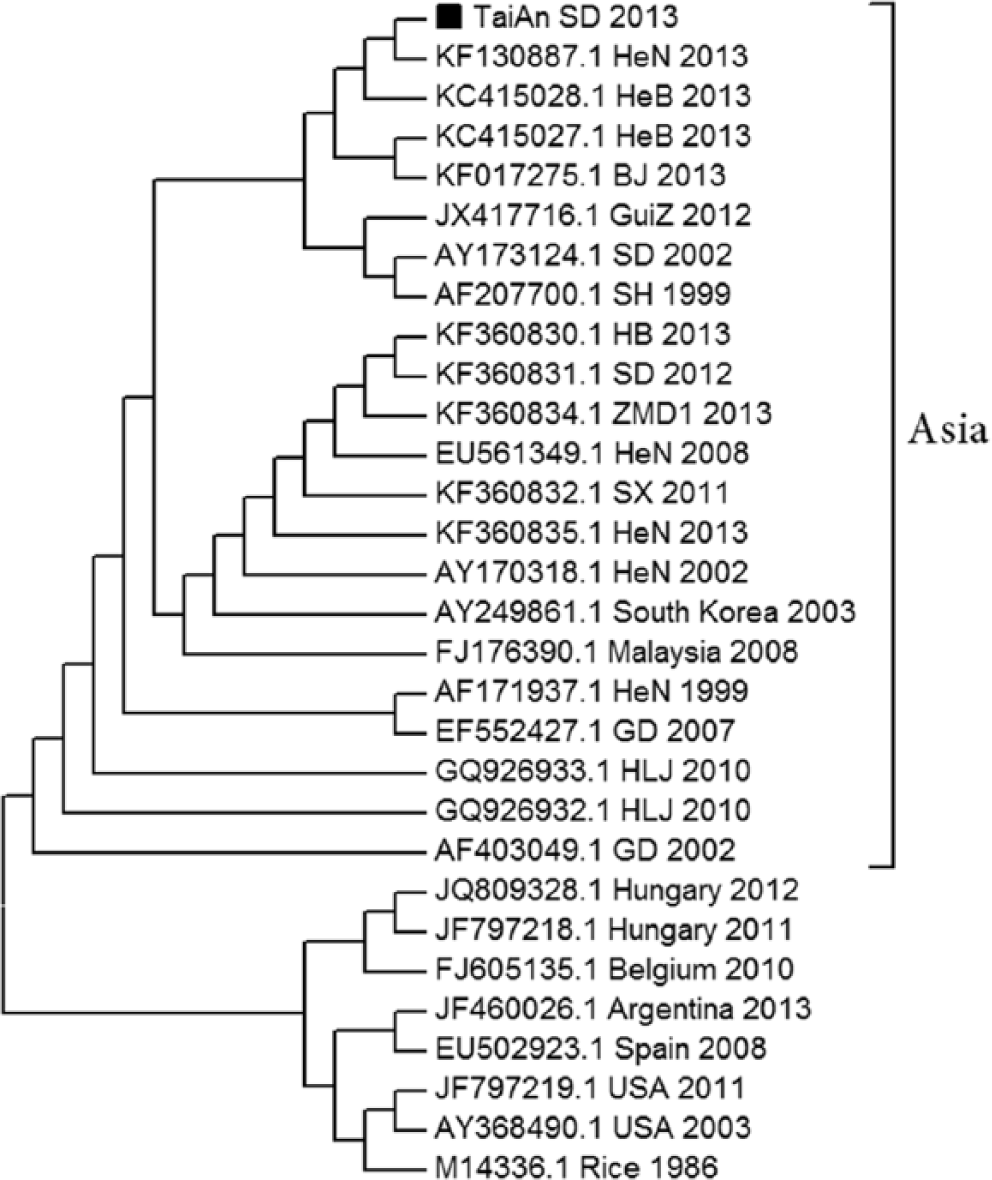

Phylogenetic analysis and comparison based on pseudorabies virus (PRV) glycoprotein E nucleotide sequences. The unrooted phylogenetic tree was generated by the distance-based, neighbor-joining method using MEGA6.0 and constructed from the aligned nucleotide sequences of 30 PRV isolates. The isolated TaiAn SD 2013 strain is marked with a black square (■).
There are a large number of pigs bred in Shandong Province. The numbers of total pigs and multiparous sows in Shandong Province in 2013 are estimated at 47,810,800 and 5,524,100, respectively, according to official statistics of the Shandong Statistical Bureau, and the number of finishing pigs for slaughter per year may reach 70,000,000, amounting to one-tenth of the yearly total of slaughtered pigs in China. Because of the lack of large-scale farms and the high breeding density in Shandong Province, the prevalence of epidemic diseases becomes more complicated with a background of ubiquitous immunosuppressive factors, such as mycotoxin and PCV-2. Swine diseases have become restrictive factors to the development of the swine industry in Shandong Province, especially the variant PRV epidemic among Bartha-K61–vaccinated piglets, which has created great challenges to the prevention and control of swine diseases.
This severe PRV outbreak occurred in several provinces (>9) and municipalities (e.g., Beijing, among others) with extensive pig farming operations in northern China, and nearly all of the affected farms used live PRV vaccines (mainly Bartha-K61 strain) to prevent PRV infection of sows and piglets. 18 The variant PRV primarily jeopardized pregnant sows and suckling piglets. Of the pregnant sows at a gestational age of 70–90 days, 5–35% experienced spontaneous abortion, 19 while the surviving piglets mainly exhibited anorexia, hydrostomia, vomiting, diarrhea, tremor, and posterior paralysis. The PRV gE gene was detected in clinical samples obtained from pigs from 15 farms in 6 provinces by specific PCR, and analysis of the gE gene sequence showed that all isolates belonged to a relatively independent cluster and contained 2 amino acid insertions. 1 The results of our study showed that the SD-2013 strain was highly homologous with recently isolated strains, especially those isolated in northern China in 2013. Nucleotide substitutions were observed in SD-2013, with G and A especially frequent, and a sequential insertion of GAC was observed at position 142–144 when compared with other western strains (i.e., Kaplan, Rice, and NiA-3). Alignment of the gE protein sequences of different PRV strains showed that the SD-2013 strain contains 2 amino acid substitutions at 105 (V–A) and 518 (G–K) compared with other isolates. The SD-2013–infected piglets exhibited severe lesions, typical nonsuppurative encephalitis, necrotizing inflammation of immune organs, necrotic foci of the liver, and diffuse hemorrhage of the renal cortex, which were more severe than lesions caused by traditional strains and strain HeN1.1,10,13
According to the gE ELISA results, 46% of the pigs were infected by PRV field strain, especially gilts, young multiparous sows, boars, and growing–finishing pigs. The sharp increase in the circulating virus infection rate was most likely due to the decreased vaccine protection against variant PRV rather than inhibition of immune responses by immunosuppressive factors, such as mycotoxin and PCV-2, as fodder was carefully selected and the farm animals were PCV-2 negative. It is well known that infected breeding pigs can spread viruses to piglets via vertical transmission, while infected growing–finishing pigs may induce persistent and circulatory infection in the herd, especially on farms with poor biosecurity measures. Therefore, PRV should be given special consideration to prevent seroconversion of the field strains on serological negative farms and limit economic losses.
Based on this epidemiological investigation, we suspected that decreased vaccine protection might be a primary risk factor for the outbreak of variant PRV among Bartha-K61–vaccinated piglets. We also noticed that some watchdogs and sheep in or around the pig farms were infected by variant PRV (PCR positive). The dogs mainly exhibited hydrostomia and tremor, while infected sheep typically exhibited hydrostomia and tended to rub their noses and bodies. Most of the affected dogs and sheep died in the acute phase of infection. Considering the high fatality rate, severe lesions, and pathogenicity to dogs and sheep, we surmise that the viral variation enhanced PRV virulence, and further challenge tests are ongoing to investigate our theory.
Footnotes
Acknowledgements
We acknowledge the help of Liang Wei and the postgraduates in the animal pathology lab of Shandong Agricultural University.
Authors’ contributions
D Hu and S Liu contributed to conception and design of the study. Y Xiao contributed to design of the study. G Wang contributed to acquisition and analysis of data. Z Zhang, Y Qu, and H Ma contributed to acquisition of data. L Lv and Y Niu contributed to analysis of data. D Hu, Z Zhang, L Lv, and H Ma drafted the manuscript. Y Qu, Y Niu, G Wang, Y Xiao, and S Liu critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
anti-gE Abs ELISA kit (lot FH060), IDEXX Laboratories Inc., Westbrook, ME.
b.
MiniBEST Viral RNA/DNA extraction kit, version 5.0, Takara Biomedical Technology, Beijing, China.
c.
2720 Thermal Cycler, Applied Biosystems, Foster City, CA.
d.
Ultraviolet light transilluminator (JY-02S), Beijing Junyi Dongfang Electrophoresis Co. Ltd., Beijing, China.
e.
Disposable filters, EMD Millipore Corp., Billerica, MA.
f.
DMEM, Life Technologies Inc., Grand Island, NY.
g.
FBS, Life Technologies Inc., Grand Island, NY.
h.
Goat anti-mouse IgG/HRP (bs-0296G-HRP), Biosynthesis Biotechnology Co. Ltd., Beijing, China.
i.
AEC kit (PW020), Sangon Biotech Co. Ltd., Shanghai, China.
j.
pMD18-T vector, Takara Biomedical Technology, Beijing, China.
k.
Sangon Biotech Co. Ltd., Shanghai, China.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
